Abstract
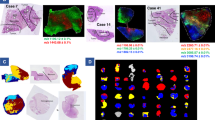
Using the sectional analysis of two-dimensional electrophoretic gels with liquid chromatography-mass spectrometry, proteoform profiles for individual genes expressed in cancer (glioblastoma) and normal (FLEH) cells were obtained. Profiles of more than 5000 genes were analyzed. It turned out that many genes encoding potential biomarkers of glioblastoma are characterized by sets of proteoforms that are different in normal and cancer cells. These proteoforms could be sources of highly specific markers and targets for therapy. Using a section analysis of two-dimensional electrophoretic gels with liquid chromatography by mass spectrometry, proteoform profiles were obtained for individual genes expressed in cancer (glioblastoma) and normal (FLEH) cells. Profiles of more than 5000 genes were analyzed. It turned out that many genes encoding potential biomarkers of glioblastoma are characterized by sets of proteoforms, which are different in normal and cancer cells. These proteoforms could be sources of highly specific markers and targets for therapy.


Similar content being viewed by others
REFERENCES
Bradford M. M., A rapid and sensitive method for the quantitation of microgram quantities utilizing the principle of protein–dye binding, Anal. Biochem., 1976, vol. 72, pp. 248–254.
Cohen, A.L. and Colman, H., Glioma biology and molecular markers, Cancer Treatment Res., 2015, vol. 163, pp. 15–30.
Fiorentino, A., Caivano, R., Pedicini, P., and Fusco, V., Clinical target volume definition for glioblastoma radiotherapy planning: magnetic resonance imaging and computed tomography, Clinical Translat. Oncol., 2013, vol. 15, pp. 754–758.
Furnari, F.B., Fenton, T., Bachoo, R.M., et al., Malignant astrocytic glioma: genetics, biology, and paths to treatment, Genes Dev., 2007, vol. 21, pp. 2683–710.
Ishihama, Y., Oda, Y., Tabata, T., Sato, T., Nagasu, T., Rappsilber, J., and Mann, M., Exponentially modified protein abundance index (empai) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein, Mol. Cell. Proteomics, 2005, vol. 4, pp. 1265–1272.
Kalinina, J., Peng, J., Ritchie, J.C., and Van Meir, E.G., Proteomics of gliomas: initial biomarker discovery and evolution of technology, Neuro Oncol., 2011, vol. 13, pp. 926–942.
Louis, D.N, Ohgaki, H., Wiestler, O.D., Cavenee, W.K., Burger, P.C., Jouvet, A., Scheithauer, B.W., and Kleihues, P., The 2007 WHO classification of tumours of the central nervous system, Acta Neuropathol., 2007, vol. 114, pp. 97–109.
Ludwig, K. and Kornblum, H.I., Molecular markers in glioma, J. NeuroOncol., 2017, vol. 134, pp. 505–512.
Naryzhny, S.N., Blue dry western: simple, economic, informative, and fast way of immunodetection, Anal. Biochem., 2009, vol. 392, no.1, pp. 90–95.
Naryzhny, S.N., Ronzhina, N.L., Mainskova, M.A., Belyakova, N.V., Pantina, R.A., and Filatov, M.V., Development of barcode and proteome profiling of glioblastoma, Biochemistry (Moscow) Suppl. Ser. B: Biomed. Chem., 2014a, vol. 8, no. 3, pp. 243–251.
Naryzhny, S.N., Lisitsa, A.V., Zgoda, V.G., Ponomarenko, E.A., and Archakov, A.I., 2DE-Based approach for estimation of number of protein species in a cell, Electrophoresis, 2014b, vol. 35, pp. 895–900.
Naryzhny, S.N., Maynskova, M.A., Zgoda, V.G., Ronzhina, N.L., Kleyst, O.A., Vakhrushev, I.V., and Archakov, A.I., Virtual-experimental 2DE approach in chromosome-centric human proteome project, J. Proteome Res., 2016a, vol. 15, pp. 525–3010.
Naryzhny, S.N., Maynskova, M.A., Zgoda, V.G., Ronzhina, N.L., Novikova, S.E., Belyakova, N.V., Kleyst, O.A., Legina, O.K., Pantina, R.A., and Filatov, M.V., Proteomic profiling of high-grade glioblastoma using virtual-experimental 2DE, J. Proteom. Bioinform., 2016b, vol. 9, pp. 158–165.
Naryzhny, S.N., Zgoda, V.G., Maynskova, M.A., Novikova, S.E., Ronzhina, N.L., Vakhrushev, I.V., Khryapova, E.V., and Archakov, A.I., Combination of virtual and experimental 2DE together with esi lc-ms/ms gives a clearer view about proteomes of human cells and plasma, Electrophoresis, 2016c, vol. 37, pp. 302–309.
Naryzhny, S., Maynskova, M., Zgoda, V., and Archakov, A., Dataset of protein species from human liver, Data Brief, 2017, vol. 12, pp. 584–588.
Zhang, R., Tremblay, T.-L., Mcdermid, A., Thibault, P., and Stanimirovic, D., Identification of differentially expressed proteins in human glioblastoma cell lines and tumors, Glia, 2003, vol. 42, pp. 194–208.
ACKNOWLEDGMENTS
This work was performed with support from the Russian Foundation for Basic Research of State Academies of Sciences for 2013–2020. Mass-spectrometry measurements were performed using the equipment of “Human Proteome” Core Facilities of the Institute of Biomedical Chemistry (Russia) which is supported by Ministry of Education and Science of the Russian Federation (agreement 14.621.21.0017, unique project ID RFMEFI62117X0017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Сonflict of interests. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. This article does not contain any studies involving animals performed by any of the authors.
Additional information
Translated by P. Kuchina
Abbreviations: IEF—isoelectric focusing, FLEH—fibroblasts of lungs of human embryos, 2DE—two-dimensional electrophoresis, ESI LC-MS/MS—liquid chromatography–mass spectrometry.
Rights and permissions
About this article
Cite this article
Petrenko, E.S., Kopylov, A.T., Kleist, O.A. et al. Searching for Specific Markers of Glioblastoma: Analysis of Glioblastoma Cell Proteoforms. Cell Tiss. Biol. 12, 455–459 (2018). https://doi.org/10.1134/S1990519X18060068
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1990519X18060068