Abstract
In this exciting era of “next-gen cytogenetics”, the use of novel molecular methods such as comparative genome hybridization and whole genome and whole exome sequencing becomes more and more common in clinics. This results in generation of large amounts of high-resolution patient-specific data and challenges the development of new approaches for interpretation of obtained information. Usually, interpretation of chromosomal rearrangements is focused on alterations of linear genome sequence, underestimating the role of spatial chromatin organization. In this article, we describe the main features of 3-dimentional genome organization, emphasizing their role in normal and pathological development. We highlight some tips to help physicians estimating the impact of chromosomal rearrangements on the patient phenotype. A separate section describes available tools that can be used to visualize and analyze human genome architecture.
Similar content being viewed by others
Abbreviations
- 3C:
-
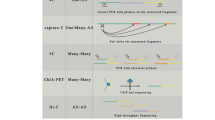
chromosome conformation capture (technology)
- ChIP:
-
chromatin immunoprecipitation
- DSBs:
-
double-strand DNA breaks
- Hi-C method:
-
high-throughput extension of 3C technology
- IGH:
-
immunoglobulin heavy chain
- TADs:
-
topologically associating domains
References
Richmond, T. J., and Davey, C. A. (2003) The structure of DNA in the nucleosome core, Nature, 423, 145–150.
Mirny, L. A. (2011) The fractal globule as a model of chromatin architecture in the cell, Chromosome Res., 19, 37–51.
Merkenschlager, M., and Nora, E. P. (2016) CTCF and cohesin in genome folding and transcriptional gene regulation, Annu. Rev. Genom. Human Genet., 17, 17–43.
Bannister, A. J., and Kouzarides, T. (2011) Regulation of chromatin by histone modifications, Cell Res., 21, 381–395.
Dixon, J. R., Jung, I., Selvaraj, S., Shen, Y., Antosiewicz-Bourget, J. E., Lee, A. Y., Ye, Z., Kim, A., Rajagopal, N., Xie, W., Diao, Y., Liang, J., Zhao, H., Lobanenkov, V. V., Ecker, J. R., Thomson, J. A., and Ren, B. (2015) Chromatin architecture reorganization during stem cell differentiation, Nature, 518, 331–336.
Dixon, J. R., Selvaraj, S., Yue, F., Kim, A., Li, Y., Shen, Y., Hu, M., Liu, J. S., and Ren, B. (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions, Nature, 485, 376–380.
Rao, S. S. P., Huntley, M. H., Durand, N. C., Stamenova, E. K., Bochkov, I. D., Robinson, J. T., Sanborn, A. L., Machol, I., Omer, A. D., Lander, E. S., and Aiden, E. L. (2014) A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping, Cell, 159, 1665–1680.
Cremer, C., and Cremer, T. (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells, Nat. Rev. Genet., 2, 292–301.
Dekker, J., Rippe, K., Dekker, M., and Kleckner, N. (2002) Capturing chromosome conformation, Science, 295, 1306–1311.
De Wit, E., and De Laat, W. (2012) A decade of 3C technologies-insights into nuclear organization, Genes Dev., 26, 11–24.
Lieberman-Aiden, E., Van Berkum, N. L., Williams, L., Imakaev, M., Ragoczy, T., Telling, A., Amit, I., Lajoie, B. R., Sabo, P. J., Dorschner, M. O., Sandstrom, R., Bernstein, B., Bender, M. A., Groudine, M., Gnirke, A., Stamatoyannopoulos, J., Mirny, L. A., Lander, E. S., and Dekker, J. (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome, Science, 326, 289–293.
Imakaev, M., Fudenberg, G., McCord, R. P., Naumova, N., Goloborodko, A., Lajoie, B. R., Dekker, J., and Mirny, L. A. (2012) Iterative correction of Hi-C data reveals hall-marks of chromosome organization, Nat. Methods, 9, 999–1003.
Talbert, P. B., and Henikoff, S. (2006) Spreading of silent chromatin: inaction at a distance, Nat. Rev. Genet., 7, 793–803.
Lupianez, D. G., Kraft, K., Heinrich, V., Krawitz, P., Brancati, F., Klopocki, E., Horn, D., Kayserili, H., Opitz, J. M., Laxova, R., Santos-Simarro, F., Gilbert-Dussardier, B., Wittler, L., Borschiwer, M., Haas, S. A., Osterwalder, M., Franke, M., Timmermann, B., Hecht, J., Spielmann, M., Visel, A., and Mundlos, S. (2015) Disruptions of topological chromatin domains cause pathogenic rewiring of gene–enhancer interactions, Cell, 161, 1–14.
Franke, M., Ibrahim, D. M., Andrey, G., Schwarzer, W., Heinrich, V., Schopflin, R., Kraft, K., Kempfer, R., Jerkovic, I., Chan, W.-L., Spielmann, M., Timmermann, B., Wittler, L., Kurth, I., Cambiaso, P., Zuffardi, O., Houge, G., Lambie, L., Brancati, F., Pombo, A., Vingron, M., Spitz, F., and Mundlos, S. (2016) Formation of new chromatin domains determines pathogenicity of genomic duplications, Nature, 538, 265–269.
Dekker, J., and Heard, E. (2015) Structural and functional diversity of topologically associating domains, FEBS Lett., 589, 2877–2884.
Valton, A. L., and Dekker, J. (2016) TAD disruption as oncogenic driver, Curr. Opin. Genet. Dev., 36, 34–40.
Holwerda, S., and De Laat, W. (2012) Chromatin loops, gene positioning, and gene expression, Front. Genet., 3.
Tang, Z., Luo, O. J., Li, X., Zheng, M., Zhu, J. J., Szalaj, P., Trzaskoma, P., Magalska, A., Wlodarczyk, J., Ruszczycki, B., Michalski, P., Piecuch, E., Wang, P., Wang, D., Tian, S. Z., Penrad-Mobayed, M., Sachs, L. M., Ruan, X., Wei, C. L., Liu, E. T., Wilczynski, G. M., Plewczynski, D., Li, G., and Ruan, Y. (2015) CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription, Cell, 163, 1611–1627.
Flyamer, I. M., Gassler, J., Imakaev, M., Brandao, H. B., Ulianov, S. V., Abdennur, N., Razin, S. V., Mirny, L. A., and Tachibana-Konwalski, K. (2017) Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition, Nature, 544, 110–114.
Ulianov, S. V., Khrameeva, E. E., Gavrilov, A. A., Flyamer, I. M., Kos, P., Mikhaleva, E. A., Penin, A. A., Logacheva, M. D., Imakaev, M. V., Chertovich, A., Gelfand, M. S., Shevelyov, Y. Y., and Razin, S. V. (2016) Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains, Genome Res., 26, 70–84.
Weinreb, C., and Raphael, B. J. (2016) Identification of hierarchical chromatin domains, Bioinformatics, 32, 1601–1609.
Denker, A., and de Laat, W. (2016) The second decade of 3C technologies: detailed insights into nuclear organization, Genes Dev., 30, 1357–1382.
Vicente-Garcia, C., Villarejo-Balcells, B., Irastorza-Azcarate, I., Naranjo, S., Acemel, R. D., Tena, J. J., Rigby, P. W. J., Devos, D. P., Gomez-Skarmeta, J. L., and Carvajal, J. J. (2017) Regulatory landscape fusion in rhabdomyosarcoma through interactions between the PAX3 promoter and FOXO1 regulatory elements, Genome Biol., 18, 106.
Ordulu, Z., Kammin, T., Brand, H., Pillalamarri, V., Redin, C. E., Collins, R. L., Blumenthal, I., Hanscom, C., Pereira, S., Crandall, B. F., Gerrol, P., Hayden, M. A., Hussain, N., Kanengisser-Pines, B., Kantarci, S., Levy, B., Macera, M. J., Quintero-Rivera, F., Spiegel, E., Stevens, B., Ulm, J. E., Warburton, D., Wilkins-Haug, L. E., Yachelevich, N., Gusella, J. F., Talkowski, M. E., and Morton, C. C. (2016) Structural chromosomal rearrangements require nucleotide-level resolution: lessons from next-generation sequencing in prenatal diagnosis, Am. J. Hum. Genet., 99, 1–19.
Battulin, N., Fishman, V. S., Mazur, A. M., Pomaznoy, M., Khabarova, A. A., Afonnikov, D. A., Prokhortchouk, E. B., and Serov, O. L. (2015) Comparison of the 3D organization of sperm and fibroblast genomes using the Hi-C approach, Genome Biol., 16, 77.
Kerpedjiev, P., Abdennur, N., Lekschas, F., McCallum, C., Dinkla, K., Strobelt, H., Luber, J. M., Ouellette, S. B., Ahzir, A., Kumar, N., Hwang, J., Alver, B. H., Pfister, H., Mirny, L. A., Park, P. J., and Gehlenborg, N. (2017) HiGlass: web-based visual comparison and exploration of genome interaction maps, bioRxiv, 1–7.
Durand, N. C., Robinson, J. T., Shamim, M. S., Machol, I., Mesirov, J. P., Lander, E. S., and Aiden, E. L. (2016) Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom, Cell Systems, 3, 99–101.
Phanstiel, D. H., Van Bortle, K., Spacek, D., Hess, G. T., Shamim, M. S., Machol, I., Love, M. I., Aiden, E. L., Bassik, M. C., and Snyder, M. P. (2017) Static and dynamic DNA loops form AP-1-bound activation hubs during macrophage development, Mol. Cell, 67, 1037–1048.
Mullighan, C. G., Goorha, S., Radtke, I., Miller, C. B., Coustan-Smith, E., Dalton, J. D., Girtman, K., Mathew, S., Ma, J., Pounds, S. B., Su, X., Pui, C.-H., Relling, M. V., Evans, W. E., Shurtleff, S. A., and Downing, J. R. (2007) Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia, Nature, 446, 758–764.
Hnisz, D., Weintraub, A. S., Day, D. S., Valton, A., Bak, R. O., Li, C. H., Goldmann, J., Lajoie, B. R., Fan, Z. P., Sigova, A., Reddy, J., Borges-Rivera, D., Lee, T. I., Jaenisch, R., Porteus, M. H., Dekker, J., and Young, R. (2016) Activation of proto-oncogenes by disruption of chromosome neighborhoods, Science, 351, 1454–1458.
Li, R., Liu, Y., Li, T., and Li, C. (2016) 3Disease browser: a web server for integrating 3D genome and disease-associated chromosome rearrangement data, Sci. Rep., 6, 34651.
Engreitz, J. M., Agarwala, V., and Mirny, L. A. (2012) Three-dimensional genome architecture influences partner selection for chromosomal translocations in human disease, PLoS One, 7, e44196.
Zhang, Y., McCord, R. P., Ho, Y.-J., Lajoie, B. R., Hildebrand, D. G., Simon, A. C., Becker, M. S., Alt, F. W., and Dekker, J. (2012) Chromosomal translocations are guided by the spatial organization of the genome, Cell, 148, 908–921.
Lieber, M. R. (2016) Mechanisms of human lymphoid chromosomal translocations, Nat. Rev. Cancer, 16, 387–398.
Aten, J. A., Stap, J., Krawczyk, P. M., Van Oven, C. H., Hoebe, R. A., Essers, J., and Kanaar, R. (2004) Dynamics of DNA double-strand breaks revealed by clustering of damaged chromosome domains, Science, 303, 92–95.
Iarovaia, O. V., Rubtsov, M. A., Ioudinkova, E., Tsfasman, T., Razin, S. V., and Vassetzky, Y. S. (2014) Dynamics of double strand breaks and chromosomal translocations, Mol. Cancer, 13, 249.
Grogg, K. L., Miller, R. F., and Dogan, A. (2006) HIV infection and lymphoma, J. Clin. Pathol., 60, 1365–1372.
Osborne, C. S., Chakalova, L., Mitchell, J. A., Horton, A., Wood, A. L., Bolland, D. J., Corcoran, A. E., and Fraser, P. (2007) Myc dynamically and preferentially relocates to a transcription factory occupied by Igh, PLoS Biol., 5, 1763–1772.
Musinova, Y. R., Sheval, E. V., Dib, C., Germini, D., and Vassetzky, Y. S. (2016) Functional roles of HIV-1 Tat protein in the nucleus, Cell. Mol. Life Sci., 73, 589–601.
Parada, L. A., McQueen, P. G., and Misteli, T. (2004) Tissue-specific spatial organization of genomes, Genome Biol., 5, R44.
Whalen, S., Truty, R. M., and Pollard, K. S. (2016) Enhancer–promoter interactions are encoded by complex genomic signatures on looping chromatin, Nat. Genet., 48, 488–496.
Di Pierro, M., Cheng, R. R., Lieberman Aiden, E., Wolynes, P. G., and Onuchic, J. N. (2017) De novo prediction of human chromosome structures: epigenetic marking patterns encode genome architecture, Proc. Natl. Acad Sci. USA, 114, 12126–12131.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fishman, V.S., Salnikov, P.A. & Battulin, N.R. Interpreting Chromosomal Rearrangements in the Context of 3-Dimentional Genome Organization: A Practical Guide for Medical Genetics. Biochemistry Moscow 83, 393–401 (2018). https://doi.org/10.1134/S0006297918040107
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297918040107