Abstract
In the present paper we describe synthesis and molecular docking simulation of substituted phenothiazines. Halonitrobenzenes on reaction with thiols give 2-amino-2′-nitrodiphenylsulphides which on formylation with 90% formic acid yield 2-formamido-2′-nitrodiphenyl sulfides. This compound undergoes Smiles rearrangement to provide phenothiazines. To evaluate the antipsychotic property of the synthesized phenothiazines derivatives (5a–d and 7), molecular docking simulation study were performed into the binding pocket of D4 dopamine receptor. These compounds have shown acceptable binding affinity with the binding pocket of the D4 dopamine receptor similar to known inhibitors. Whereas, compound 7 has displayed higher affinity for D4 dopamine receptor compared to D3 and D2 receptors. All compounds interacted with critical residues for the inhibition of D4 dopamine receptor and hence can be developed as potent antipsychotic drugs.
Similar content being viewed by others
1 Introduction
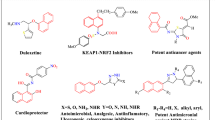
Phenothiazines are well known for their pharmacological and biological activities and their several derivatives are in clinical use [1, 2]. An important effort has been done on the bioactivity of phenothiazine and its derivatives. These are also used as analgesics [3], antihistamines [4], diuretics [5], antimicrobial [6], neuroleptics [7], antitumor activity [8,9,10]. A slight modification in the substitution pattern in phenothiazine nucleus causes a marked difference in biological activities, therefore, it is considered worthwhile to synthesize hitherto unknown phenothiazines in order to make them available for biomedical screening in search of better medicinal agents. Kumar et al. have reported optical property of chitosan-phenothiazine derivative by microwave assisted synthesis [11].
Phenothiazines are dopamine antagonists and can be utilized for calmness, tranquility, or to diminish stress or anxiety [12]. A group of phenothiazine derivatives have been utilized for the treatment of schizophrenia and related psychotic disorders [13, 14]. A kind of literatures suggests that antischizophrenic property of phenothiazine drugs attributed by blocking the dopamine-1 (D1) and dopamine-2 (D2) receptors within the mesolimbic dopamine pathway [15]. Except clozapine, most antipsychotic drugs inhibit D2 receptors. Studies revealed that in schizophrenia brain, density of D2 and D3 receptors is elevated by 10% [16], whereas density of dopamine D4 receptors is elevated by six fold [17]. Additionally, D4 receptors are predominant localized in the mesolimbic/mesocortical area suggesting that the D4 receptors are potential target for the development of newer antipsychotics [18]. Chlorpromazine is a well-known typical phenothiazine antipsychotic drug used for the treatment of manic-depression and schizophrenia in adults [19]. The clinical trial studies suggest that at varying dose ranges, and higher doses, chlorpromazine can cause more extrapyramidal adverse effects [13]. Hence, synthesis of new derivatives of phenothiazines that can inhibit the D4 dopamine receptor and act as antipsychotic drugs is greatly needed for the treatment of psychotic disorders. Compound 7 exhibited higher D4 selectivity over D2 and D3 receptors as compared to all synthesized compounds along with reference molecules. These new drugs are expected to have higher specificity causing less side effects as compared with existing compounds.
Therefore, in the present study we have developed a precise strategy to synthesize derivatives of phenothiazines that can bind to D4 receptors and act as antipsychotic drugs.
2 Materials and methods
2.1 Chemicals and reagents
2-aminobenzenethiols, 2-halonitrobenzene, 4-chloro-3,5-dinitro-benzoic acid, ethanol, anhydrous sodium acetate, potassium hydroxide, formic acid, benzene and acetone were purchased from Merck Life Science Pvt. Ltd. (India). Other chemicals used in this study were purchased from Merck Life Science Pvt. Ltd. (India). Ultrapure water was obtained by a Milli-Q® water purification system (Merck Life Science Private Limited, India) and used for all preparations.
2.2 Characterization
All the melting points of synthesized compounds are uncorrected. The purity of the synthesized compounds has been checked by thin layer chromatography using silica gel with different non-aqueous solvent systems. Characterization of the synthesized compounds was carried out by their spectral data. The compounds were initially measured by Fourier transform infrared (FT-IR) spectra (Perkin-Elmer 550 FTIR spectrophotometer) using KBr pellet. The 1H NMR has been recorded on 90 MHz on Jeol FX 9OQ FT NMR spectrometer using TMS as an internal standard in DMSO-d6. Mass spectra were recorded on a Jeol JMSD-300 Mass spectrometer at 70 eV with 100 µ amp ionizing current.
2.3 Synthesis
2.3.1 Synthesis of 2-amino-2′-nitrodiphenyl sulfides (3a–d)
In a 50 mL round bottom flask 0.01 mol of 2-aminobenzenethiols (1) dissolved in 20 mL ethanol was taken and anhydrous sodium acetate (0.01 mol) dissolved in 5 mL ethanol was added to it. To this 0.01 mol alcoholic solution (10 mL) of 2-halonitrobenzene was also added on magnetic stirring. The whole mixture was refluxed for 3–4 h. After the reaction was over the mixture was concentrated and cooled overnight in an ice chamber. The solid product was separated out and filtered, washed with 30% ethanol and crystallized from methanol.
2.3.2 Synthesis of substituted 2-formamido-2′-nitrodiphenyl sulfides (4a–d)
A mixture of 0.01 mol of diphenyl sulfides (3a–d) and 20 mL of 90% formic acid was refluxed for 4 h. After that, the contents were poured over crushed ice. The solid product separated out was filtered, washed with water till neutralization and crystallized from benzene.
2.3.3 Synthesis of substituted phenothiazines (5a–d)
In a 50 mL round bottom flask 0.01 mol of formyl derivatives (4a–d) dissolved in 15 mL acetone was taken. To this an alcoholic solution of potassium hydroxide (0.2 g dissolved in 5 mL of ethanol) was also added. The contents were refluxed for 30 min. To this refluxing solution, a second lot of potassium hydroxide (0.2 g dissolved in 5 mL ethanol) was added and again refluxed for 2 h. After the reaction was over the contents were poured over crushed ice. The solid separated out was filtered, washed with cold water and finally with 30% ethanol and recrystallized from benzene.
2.3.4 Synthesis of 1-nitro-10H-phenothiazines-3-carboxylic acid (7)
A mixture of 4-chloro-3,5-dinitro-benzoic acid (6, 0.01 mol), 2-aminobenzenethiol (1, 0.01 mol), sodium hydroxide (0.01 mol) and absolute ethyl alcohol (20 mL) was refluxed for 2 h. The reaction mixture was concentrated on rotavapour, cooled and filtered. The precipitate was washed well with hot water and finally with 30% ethanol and recrystallized from acetone/benzene.
2.4 Prediction of molecular drug properties
The synthesized phenothiazines derivatives were examined for molecular drug properties including hydrogen bond donor (HBD), hydrogen bond acceptor (HBA), polar surface area (PSA), Molar refractivity, Partition coefficient (logP), human intestinal absorption (HIA) and blood–brain barrier (BBB) penetration ability. The Molar refractivity, HBD, HBA, PSA and logP were calculated by Chemicalize-Instant Cheminformatics Solutions (https://chemicalize.com/). The Lipinski’s Rule of Five distinguishes between the drug like and non-drug like molecules. It predicts high probability of failure of molecules due to non-drug-likeness for the molecules not complying with 2 or more of the following rules: (1) molecular wt. < 500; (2) logP < 5; (3) H-bond donors < 5; and (4) H-bond acceptors < 5 [20]. The PreADMET server examined the Human intestinal absorption (HIA) property of compounds [21]. The HIA is one of the most important absorption, distribution, metabolism and excretion (ADME) properties of candidate compounds which predict the oral bioavailability of the drug [22]. The research findings revealed that > 98% of drug candidates are unable to penetrate the BBB, that is a major roadblock in the formulation of an effective neurotherapeutic drug. Hence prediction of BBB penetration ability of drug candidates is necessary during targeting it to the brain [23]. The analysis for blood brain barrier (BBB) penetration ability of phenothiazines derivatives was facilitated by online BBB prediction server with the AdaBoost algorithm and PubChem fingerprint (http://www.cbligand.org/BBB/).
2.5 Docking preparation and simulation
Atomic coordinates for the crystal structure of human D4 Dopamine receptor in complex with Nemonapride (PDB ID: 5WIU) were downloaded from RCSB-Protein Data Bank (PDB). The structure of D4 receptor (chain A) was determined by X-ray diffraction at 1.962 Å resolutions. The three-dimensional structures of synthesized phenothiazines derivatives were drawn by using MarvinSketch (version 6.2.2, calculation module developed by ChemAxon, http://www.chemaxon.com/products/marvin/marvinsketch/, 2014. The 3D structures of two atypical antipsychotics (Nemonapride and clozapine) and a typical antipsychotic (chlorpromazine) were retrieved from NCBI PubChem database. The energy minimization of D4 receptor, phenothiazines derivatives (5a–d and 7) and antipsychotics were performed by UCSF Chimera according to Kumar et al. [24]. The docking simulation was performed according to Kumar et al. [25, 26]. Briefly, a grid box, which comprises the entire binding pocket of D4 receptor including orthosteric binding pocket (OBP) and extended binding pocket (EBP) was generated that provides enough space for rotational and translational walk to the ligands. The values for center grid box were kept − 16.685, 9.323 and − 19.581 for X, Y, and Z-center with 0.375 Å spacing and number of points 76, 100 and 88, in X, Y, and Z dimension. The docking was performed by using Lamarckian genetic algorithm (LGA) with 30 independent runs. All compounds were also docked with D2 dopamine receptor (PDB ID: 6CM4) and D3 dopamine receptor (PDB ID: 3PBL) to evaluate its D4 receptor (PDB ID: 5WIU) selectivity by keeping constant grid box’s number of points 76, 100 and 88, in X, Y, and Z dimension.
Consequently, poseview (https://poseview.zbh.uni-hamburg.de/) was used for visualization of the interaction pattern between the D4 receptor-ligand complexes.
3 Results and discussions
3.1 Characterization of substituted phenothiazines
The overall syntheses of substituted phenothiazines are illustrated in Schemes 1 and 2. The physical property, FTIR and 1HNMR data of the synthesized compounds are represented in Tables 1, 2 and 3. The reaction of 2-aminobenzenethiol (1) with 2-halonitrobenzenes (2) containing a nitro group ortho position to the halogen atom gives rise to diphenylsulfides (3a–d). The latter on formylation yield 2-formamido-2′-nitrodiphenyl sulfides (4a–d). In the presence of alkali, the imido nitrogen donates its lone pair of electrons for intramolecular nucleophilic attack on the carbonium ion. As results, sulfur loses its electron pair and the positively charged nitrogen provides the proton which attaches to sulfide (S−) yielding mercaptodiphenylamine. The latter undergoes cyclization with the loss of nitrous acid accompanied by the simultaneous hydrolysis of formyl group on nitrogen yielding phenothiazines (5a–d) Scheme 1. The condensation of 2-aminobenzenethiol with 4-chloro-3,5-dinitro-benzoic acid containing a nitro group at both ortho position to the halogen atom in which the increased resonance effect of both nitro groups, due to their ortho position and combined resonance and inductive effect reinforced by the same nitro groups, activates the Smiles rearrangement [27] as well as the ring closure to such an extent that the processes take place instantaneously and in situ yielding 1-nitro-10H-phenothiazines-3-carboxylic acid (7) Scheme 2.
3.2 Molecular drug properties
The molecular drug properties of synthesized substituted phenothiazines (5a–d) and 1-nitro-10H-phenothiazines-3-carboxylic acid (7) are listed in the Table 4. All compounds obey Lipinski rule of 5 (Ro5) because they have molecular mass ≤ 500 Dalton; logP ≤ 5; HBD ≤ 5; HBA ≤ 10 and Molar refractivity in between 40 and 130 [28]. All compounds were also predicted as BBB permeable and their HIA ranges 80.76–96.78.
3.3 Binding mode analysis
All ligands including antipsychotic drugs (nemonapride, clozapine and chlorpromazine) and phenothiazines derivatives (5a–d and 7) were docked at the binding pocket of D4 dopamine receptor to predict the possible binding interactions between them. Table 5 summarizes the docking results including inhibition constant (Ki), hydrogen bonding and hydrophobic interaction displayed at lowest binding energy conformation. Among antipsychotic drugs, nemonapride has exhibited higher binding affinity toward the binding pocket of D4 receptor compared to chlorpromazine and Clozapine. Whereas, among synthesized phenothiazines derivatives, 7 has bound to the D4 receptor with lower binding energy compared to 5a, 5b, 5c and 5d. However, both 7 and 5c has shown higher affinity in terms of formation of hydrogen bonding and hydrophobic interaction with binding site of D4 receptor.
According to Wang et al. [29], the binding site of D4 dopamine receptor consist orthosteric binding pocket (OBP) and extended binding pocket (EBP). The binding pocket of D4 dopamine receptor comprises Val 193, Leu187, Asp115, Cys185, His414, Ser197, Ser196, Ser200, Thr120, Tyr438 and Phe91. The Nemonapride formed hydrogen bonds with Asp115, Ser196 and water molecules in the binding pocket. Its benzamide ring binds within a conserved OBP and unsubstituted benzyl group binds to the EBP. Wang et al. suggested that the interaction of drug molecule with EBP binding pocket might confer selectivity for the D4 dopamine receptor [29]. The Nemonapride interacts with the conserved Asp115 and nonconserved residues i.e. Leu111 and Phe91. In present study, nemonapride re-docked to the binding pocket of D4 receptor to validate the docking accuracy. We found that nemonapride formed hydrogen bonds with Asp 115 and Leu187 as well as established hydrophobic contacts with His414, Leu187, Val87, Leu111, Leu 90, and Phe91. The nemonapride bound to OBP and EBP binding pocket of D4 dopamine receptor similarly as suggested by Wang et al. (Figure 1) that validated the docking accuracy of the present study [29]. Clozapine, an atypical antipsychotic has ten fold higher affinity for D4 compared to D2 and D3 receptors [30]. The clozapine when docked with D4 receptor, displayed lower affinity than nemonapride and bound only into the OBP binding pocket (Fig. 2). In the lowest binding energy conformation clozapine formed two H-bonds (Leu187, His414gj) and established four hydrophobic interactions (Val 193, Phe410, Leu187, His414) with D4 OBP binding pocket (Fig. 2). Similarly, chlorpromazine a typical antipsychotic drug, has also showed lower affinity than nemonapride and clozapine as well as bound to EBP binding pocket in its lowest binding conformation (Fig. 4). It formed only one H-bond with Asp115 residue but has seven hydrophobic contacts (Val116, Met112, Asp115, Cys 119, Phe411, His414, Leu187) in the EBP and OBP binding pocket of D4 receptor.
Interaction of nemonapride with D4 dopamine receptor. The nemonapride binds in the OBP and EBP binding pockets (left; surface structure, colored by heteroatoms). Hydrogen bonding and hydrophobic interaction of nemonapride shown in 2D plot (right). Green line represents hydrophobic interaction and hydrogen bonding presented in dotted lines
Binding pattern of clozapine and 5a with OBP binding pocket of D4 dopamine receptor (left; surface structure, colored by heteroatoms). Hydrogen bonding and hydrophobic interaction of clozapine and 5a shown in 2D plot (right). Green line represents hydrophobic interaction and hydrogen bonding presented in dotted lines
All phenothiazines derivatives (5a–d and 7) bound to the D4 receptor with modest binding energy that displayed lower binding affinity compared to nemonapride, clozapine and chlorpromazine. Out of five synthesized compounds, 5c and 7 have higher binding affinity toward D4 receptor compared to 5a, 5b and 5d. The 5a molecule only occupy the OBP pocket of the receptor similar to binding patten of clozapine antipsychotic. The compound 5a formed a hydrogen bond with Asp115 residue and established hydrophobic contacts with Val116 and His414 OBP pocket of the receptor (Fig. 2). The compounds 5b, 5c and 5d have shown similar binding pose as displayed by nemonapride by occupying the EBP and OBP binding pocket of the D4 dopamine receptor (Fig. 3). Compound 5b interacted with Leu187 via hydrogen bonding and has Leu187, Val87, Leu111 residues in its hydrophobic contacts in the lowest binding conformation (Fig. 3). The 5c has formed two hydrogen bonds with His414 and Leu187 residues of OBP pocket and established hydrophobic interactions with Val87 and Leu111 residues of EBP pocket (Fig. 3). Similarly, 5d molecule binds in between the OBP and EBP pockets and formed a hydrogen bonds with Cys185 residue as well as it established hydrogen bonding with both pocket’s residues i.e. Met112 (OBP) and Leu111 (EBP) (Fig. 3). Whereas, compound 7 has only bound into the EBP pocket and established two hydrogen bonds with Cys185 and Arg186 residues as well as has hydrophobic contacts with Val87, Leu111 and Phe91 residues (Fig. 4).
Briefly, compounds 5b, 5c and 5d have binding pattern similar to nemonapride and compound 5a has similar binding conformation as displayed by clozapine and chlorpromazine. Whereas, molecule 7 showed distinct binding pattern to all compound i.e. it bound to EBP binding pocket of the D4 receptor as displayed chlorpromazine with higher affinity compared to compounds 5a–d. The selectivity study was also carried out to analyze the D4 selectivity of all compounds over D2 and D3 receptors (Table 6). All synthesized phenothiazines derivatives (5a–d and 7) along with nemonapride, clozapine and chlorpromazine were docked with D2, D3 and D4 dopamine receptors and Ki values were compared for the selectivity analysis. It is demonstrated that nemonapride is selective inhibitor of D4 receptor [29]. In the present docking study we also found that nemonapride exhibits twofold selectivity for D4 over D2 receptor and fourfold selectivity for D4 over D3 receptor. Similarly, compound 7 has shown about sixfold higher affinity for D4 as compared to D3 receptor and 11-fold higher affinity for D4 as compared to D2 receptor {selectivity is presented as the ratios of Ki (D3 or D2) to Ki (D4)}. The compound 7 binds D4 receptor with lowest binding energy of − 7.88 kcal/mol and 1.68 µM Ki. Whereas, the lowest binding energy and Ki value of compound 7 for D3 and D2 receptors were − 6.80 kcal/mol and 10.35 µM as well as − 6.42 kcal/mol and 19.52 µM respectively.
Despite that the synthesized phenothiazines derivatives (5a–d and 7) have shown lower binding affinity for D4 dopamine receptor as compared to nemonapride and clozapine, all interacted with critical residues of D4 dopamine receptor’s binding pocket. Moreover, compound 7, due to its ability to interact only with the EBP binding pocket, show higher affinity for D4 as compared to D3 and D2 receptors and may be the basis for the synthesis of new compounds with improved binding properties.
4 Conclusions
We synthesized substituted phenothiazines followed by formylation and one pot reaction of 1-nitro-10H-phenothiazines-3-carboxylic acid via Smiles rearrangement. The synthesized phenothiazines derivatives 5a, 5b, 5c and 5d have shown acceptable binding affinity with the binding pocket of the D4 dopamine receptor similar to known inhibitors. Whereas, compound 7 has displayed entirely different binding pattern with the receptor’s binding pocket. These all compounds interacted with critical residues for mediating inhibiton hence can be developed as potent inhibitor of D4 dopamine receptor. Further, in vitro and in vivo studies are being carried out to evaluate their D4 receptor inhibition capacity and antipsychotic property.
References
Zwanenburg B (2010) Phenothiazines and 1,4-benzothiazines. Chemical and Biomedical aspects, R. R. Gupta. Elsevier, Amsterdam, 1988, XXII + 992 pp. US $284.25/Dfl. 540,00. ISBN 0-444-42967-0. Recl des Trav Chim des Pays-Bas
Varga B, Csonka Á, Csonka A, Molnár J, Amaral L, Spengler G (2017) Possible biological and clinical applications of phenothiazines. Anticancer Res 37:5983–5993
Chen YW et al (2010) Phenothiazine-type antipsychotics elicit cutaneous analgesia in rats. Acta Anaesthesiol Taiwan 48:3–7
Maurer H, Pfleger K (1988) Identification of phenothiazine antihistamines and their metabolites in urine. Arch Toxicol 62:185–191
Jarchovsky J, Zamir D, Plavnik L (1991) Torsade de pointes and use of phenothiazine and diuretic. Harefuah 121:435–436
Singh G, Kumar N, Yadav AK, Mishra AK (2003) Potential antimicrobial agents: Trifluoromethyl-10H-phenothiazines and ribofuranosides. Heteroat Chem 14:481–486
Kiselyuk A et al (2010) Phenothiazine neuroleptics signal to the human insulin promoter as revealed by a novel high-throughput screen. J Biomol Screen 15:663–670
Motohashi N, Kawase M, Saito S, Sakagami H (2005) Antitumor potential and possible targets of phenothiazine-related compounds. Curr Drug Targets 1:237–246
Motohashi N et al (1999) Antitumor activity of benzo[a]phenothiazines. Anticancer Res 19:1837–1842
Motohashi N, Gollapudi SR, Emrani J, Bhattiprolu KR (1991) Antitumor properties of phenothiazines. Cancer Invest 9:305–319
Kumar S, Dutta J, Dutta PK (2009) Preparation, characterization and optical property of chitosan- phenothiazine derivative by microwave assisted synthesis. J Macromol Sci Part A Pure Appl Chem 46:1095–1102
Suckow M, Stevens K, Wilson R (2012) The laboratory rabbit, guinea pig, hamster, and other rodents. Academic Press, Cambridge
Ramachandraiah CT, Subramaniam N, Tancer M (2009) The story of antipsychotics: past and present. Indian J Psychiat 51:324
Wu CH et al (2016) Pharmacological exploitation of the phenothiazine antipsychotics to develop novel antitumor agents-A drug repurposing strategy. Sci Rep 6:2740
Horn AS, Snyder SH (1971) Chlorpromazine and dopamine: conformational similarities that correlate with the antischizophrenic activity of phenothiazine drugs. Proc Natl Acad Sci USA 68:2325–2328
Seeman P, Van Tol HHM (1994) Dopamine receptor pharmacology. Trends Pharmacol Sci 15:264–270
Seeman P, Van Tol HHM (1993) Dopamine receptor pharmacology. Curr Opin Neurol Neurosurg 15:264–270
Kulkarni SK, Ninan I (2000) Dopamine D4 receptors and development of newer antipsychotic drugs. Fundam Clin Pharmacol 14:529–539
Saha KB, Bo L, Zhao S, Xia J, Sampson S, Zaman RU (2016) Chlorpromazine versus atypical antipsychotic drugs for schizophrenia. Cochrane Database Syst Rev 4:1–253
Benet LZ, Hosey CM, Ursu O, Oprea TI (2016) BDDCS, the Rule of 5 and drugability. Adv Drug Deliv Rev 101:88–98
Lee SK, Lee IH, Kim HJ, Chang GS, Chung JE, No KT (2003) The PreADME approach: web-based program for rapid prediction of physico-chemical, drug absorption and drug-like properties. Blackwell Publishing, Massachusetts, pp 418–420
Yan A, Wang Z, Cai Z (2008) Prediction of human intestinal absorption by GA feature selection and support vector machine regression. Int J Mol Sci 9:1961–1976
Mukherjee S, Kumar G, Patnaik R (2017) Identification of potential inhibitors of PARP-1, a regulator of caspase-independent cell death pathway, from Withania somnifera phytochemicals for combating neurotoxicity: a structure-based in silico study. J Theor Comput Chem 16(7):1750062
Kumar S, Kumar G, Tripathi AK, Seena S, Koh J (2018) Enhanced fluorescence norfloxacin substituted naphthalimide derivatives: molecular docking and antibacterial activity. J Mol Struct 1157:292–299
Kumar G, Paliwal P, Patnaik N, Patnaik R (2017) Withania somnifera phytochemicals confer neuroprotection by selective inhibition of nNos: an in silico study to search potent and selective inhibitors for human nNOS. J Theor Comput Chem 16(5):1750042
Kumar G, Paliwal P, Patnaik R (2017) Withania somnifera phytochemicals confer neuroprotection by inhibition of the catalytic domain of human matrix metalloproteinase-9. Lett Drug Des Discov 14(6):718–726
McLafferty FW (1959) Mass spectrometric analysis: molecular rearrangements. Anal Chem 32:82–87
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2012) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23:3–25
Wang S et al (2017) D4 dopamine receptor high-resolution structures enable the discovery of selective agonists. Science 358:381–386
Sanyal S, Van Tol HHM (1997) Review the role of dopamine D4 receptors in schizophrenia and antipsychotic action. J Psychiat Res 31:219–232
Acknowledgements
The authors are thankful to University Grant Commission, New Delhi, for financial assistance in the form of a major research project.
Funding
This study was funded by UGC, New Delhi.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kumar, S., Kumar, G. & Shukla, I.C. Substituted phenothiazines: synthesis and in silico evaluation of D4 dopamine receptor inhibition. SN Appl. Sci. 2, 1241 (2020). https://doi.org/10.1007/s42452-020-3067-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-020-3067-7