Abstract
Chinese privet (Ligustrum sinense) is a common invasive shrub in hardwood forests of the southeastern US and has been shown to negatively affect native herbaceous and woody plants. The ability to map the distribution of L. sinense on a property could help land managers plan and budget for control operations. We evaluated whether freely available moderate resolution multispectral imageries (Landsat 8 and Sentinel 2) and open-source GIS software (QGIS with the Semi-Automatic Classification Plugin) could be effective tools for this application. These tools are widely used by remote sensing and mapping professionals; however their adoption by field-level land managers appears limited, and their utility for mapping L. sinense invasions is untested. We evaluated how satellite type, image acquisition date, classification algorithm, and L. sinense cover affected detection accuracy. We found that Sentinel 2 imagery from March tended to produce good results, especially when analyzed using the maximum likelihood algorithm. Our best classifier obtained an overall accuracy of 92.3% for areas with ≥ 40% L. sinense cover. We recommend that land managers interested in applying this tool use an adaptive process for developing training polygons and test multiple images and classification algorithms in order to achieve optimal results.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Chinese privet (Ligustrum sinense) is an invasive shrub with a broad global range outside its native distribution [1]. It is particularly problematic in the southeastern US, where it and congeneric European privet (L. vulgare L.) were estimated in 2008 to cover over a million hectares [2, 3]. Ligustrum sinense can outcompete native plant species, potentially degrading wildlife habitat and limiting forest regeneration. Control costs are estimated around $216–$1820 per ha [4, 5], necessitating careful planning and budgeting on behalf of land managers who are interested in forest restoration. The objective of this study is to evaluate whether free satellite imagery and simple to use open-source software could be an effective tool for land managers who need to map L. sinense invasions to help plan hardwood forest restoration projects.
Ligustrum sinense was introduced to the southeastern US for landscaping in 1852 and has since spread throughout the region, primarily through endozoochory and hydrochory [6,7,8]. Individuals can have a single- or multi-stemmed growth form and may reach 10 m tall [6, 7]. The phenology of the plant is variable depending on the local climate and can range from evergreen to deciduous [7]. Negative correlations between L. sinense abundance and native plant abundance and diversity have been documented by many studies [e.g., 9, 10], and some authors are concerned that the lack of woody regeneration under L. sinense canopies could lead to severe forest degradation over time [10,11,12,13]. The plant has broad environmental tolerances and can be found in upland and bottomland sites [6, 14,15,16].
Public and private land managers interested in controlling L. sinense would benefit from being able to estimate the acreage requiring treatment on a particular property so that costs can be calculated and budgeted for. On large properties it would be time consuming and difficult to determine the invaded acreage based solely on field surveys. In situations where L. sinense is growing under a deciduous hardwood overstory, the phenological differences between L. sinense and the overstory can be exploited during the dormant season to map L. sinense coverage using satellite or aerial data. In a 2002 study, investigators took advantage of these phenological differences to map L. sinense based on manual interpretation of 1-m resolution color infrared or black and white aerial photographs [17]. This method seemed to be relatively successful, although they did not conduct a formal accuracy assessment. However, there are some notable downsides to their approach. Manual photo interpretation is time consuming and accuracy is highly dependent on the skill of the interpreter. Additionally, high-resolution leaf-off imagery is not always freely available, possibly requiring data to be purchased.
A more data-intensive approach for mapping L. sinense presence has also been utilized. Investigators used 1-m resolution LiDAR (light detection and ranging) data and 1-m resolution leaf-off color infrared IKONOS imagery (both resampled to 5 m) to create 80 model variables (43 canopy and 23 topographic metrics derived from LiDAR and 14 spectral metrics derived from IKONOS imagery) [18]. These variables were used in logistic regression and random forest (RF) classification models. The best performing models were RF models based on LiDAR derived metrics, which took into consideration vegetation structure, topography, and spectral characteristics. The downside to this method is that LiDAR is not always freely available and can be expensive to acquire [19]. There is also a relatively high level of technical expertise needed to process LiDAR data and run RF classifiers in a programming environment such as interactive data language (IDL). The cost and expertise required to implement this technique may serve as a barrier to its implementation by land managers.
Fortunately, there are free and easy-to-use data sources that could be used for mapping L. sinense. Moderate resolution, multispectral satellite imagery is commonly used for land cover mapping [20,21,22], including invasive plant detection [e.g., 23]. These satellite sensors measure the reflectivity of the earth’s surface at multiple wavelengths, or bands, of the electromagnetic spectrum. This includes the visible spectrum (i.e., blue, green, and red), as well as the wavelengths outside the visible spectrum such as infrared. Different land cover types reflect sunlight with varying intensities across the electromagnetic spectrum due to variation in pigmentation, texture, water content, and other factors [24, 25]. These differences in reflectivity, known as spectral signatures, can be used to distinguish among land cover types [24, 25]. Healthy vegetation is particularly easy to distinguish, versus non-vegetated areas or dormant vegetation, due to the near-infrared reflecting properties of leaf cell tissues [24]. Moderate resolution multispectral imagery is provided free to the public through the United States’ Landsat and European Space Agency’s Sentinel 2 (S2) programs [26, 27]. Landsat 8 (L8), the most recent iteration of the Landsat series, uses its onboard Operational Land Imager to collect 9 band imagery at 30-m spatial resolution (except for the 15-m panchromatic band; [27]). Sentinel 2 uses its Multispectral Instrument to collect 13 band imagery at resolutions of 10-, 20-, and 60-m [26].
A recent study tested the effectiveness of mapping L. sinense in North Carolina using Landsat 5 imagery and a RF classifier implemented in the R statistical software [28]. They tested a range of models that included various combinations of Landsat bands, vegetation indices based on the Landsat bands, and topographic indices based on digital elevation models. They found that imagery from early- to mid-March captured the greatest phenological differences between L. sinense and uninvaded deciduous forest, and thus resulted in the most accurate detection models. This study effectively demonstrated that Landsat imagery can be used to map L. sinense coverage with accuracy that is sufficient for monitoring and management purposes.
The method employed by [28] utilized free data (Landsat 5), making it more accessible than previous methods [i.e., 17, 18]. However, its reliance on the R programming language and the incorporation of vegetation and topographic indices means that it requires a level of technical skill that may still be beyond the abilities of many public and private land managers, due to a lack of relevant training. In order for a remote sensing technique to be accessible for land managers, software with a point-and-click graphical user interface (GUI) and a straight-forward, well documented workflow would be best. Fortunately, such software exists in the form of the Semi-Automatic Classification Plugin (SCP; [29]) within QGIS [30]. This software is open source (i.e., free), has a simple to use GUI, and there are excellent support materials and tutorials available online, all of which make this a seemingly ideal option for users who have limited geographic information system (GIS) experience. QGIS is commonly used by mapping, remote sensing, and land management professionals; however its utility for mapping L. sinense cover in hardwood forests is untested.
The primary objective of this study is to determine whether the SCP could be an effective tool for mapping L. sinense cover in a bottomland hardwood forest. If the SCP is determined to be an effective tool for this application, then we plan on producing a step-by-step guide for land managers interested in implementing this technique themselves. Secondary objectives are to evaluate the influence of L. sinense cover, imagery type (S2 vs. L8), classification algorithm, and imagery acquisition date on classification accuracy.
2 Methods
2.1 Study site
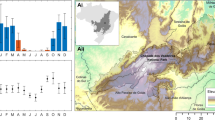
We conducted our study on a 2300 ha private property located in the floodplain of the Black Warrior River in west-central Alabama, on the border between Hale and Tuscaloosa counties (Fig. 1). The property is dominated by bottomland hardwood forests, with some interspersed loblolly pine (Pinus taeda) stands, hay fields, wildlife food plots, swamps, and oxbow lakes. Bottomland hardwood forests on the property occupied a range of geomorphic and topographic positions, with forests at various successional stages. Common forest species include cherrybark oak (Quercus pagoda), sweetgum (Liquidambar styraciflua), swamp chestnut oak (Quercus michauxii), and bitternut hickory (Carya cordiformis). Bald cypress (Taxodium distichum) and water tupelo (Nyssa aquatic) occur in forested swamps and along the edges of oxbow lakes. The bottomland hardwood forests on the property exhibit a range of L. sinense cover, including uninvaded areas and dense L. sinense monocultures. The proportion of invaded and uninvaded hardwood forests is relatively equal.
Map of our study site in the floodplain of the Black Warrior River, in western Alabama (relative location shown by yellow square on inset map, not to scale). The background imagery is an infrared false color composite Sentinel 2 image taken on 2017/03/09. The training polygons used to compute spectral signatures for each land cover class are shown in yellow, and the random points used for accuracy assessment are shown in black
2.2 Imagery acquisition
We downloaded four S2 and four L8 scenes from earthexplorer.usgs.gov. For each satellite, we chose two early- to mid-March images and two January images (Table 1). Only images from 2017 or later were considered to limit potential changes in L. sinense cover that may have occurred between the image acquisition date and our field survey. Early- to mid-March is considered late dormant season and has been identified by previous researchers as the period of maximum phenological difference between L. sinense and deciduous hardwoods [28]. January is the middle of the dormant season and provided a useful comparison to the late dormant season March imagery. Landsat 8 scenes were downloaded as Collection 1, Level 1 products [27] and Sentinel 2 scenes were downloaded as Level 1C products [26]. Atmospheric correction to surface reflectance was unnecessary, because a separate set of training signatures were calculated for each image, precluding the need for radiance values to be standardized [31]. The bands for each image were clipped to our study site and a separate band stack was created for each image. Band stacks for L8 images included bands 2–7, while band stacks for S2 images included bands 2–8, 8A, and 11–12, based on preset options in the SCP. Multiband stacks had a spatial resolution of 10-m for S2 and 30-m for L8.
2.3 Supervised classification
We implemented a supervised classification approach using SCP (version 6.2.9) in QGIS (version 3.6.2). In a supervised classification the user creates a set of training areas that are representative of the land cover classes of interest. The software then calculates the spectral signatures of all pixels within those training areas. The spectral signature of a pixel is a representation of the intensity of the light being reflected within each of the bands of the electromagnetic spectrum sampled by the satellite sensor. Once the training signatures have been created, the software sorts all the pixels in the image into the appropriate land cover classes by comparing the spectral signature of each image pixel to the training signatures and choosing the best match. There are multiple algorithms available within the SCP that sort pixels based on different definitions of “best match.” We tested three of the available options: minimum distance (MD), maximum likelihood (ML), and spectral angle mapping (SA), with no minimum thresholds [29]. We classified each of our 8 images using all 3 classification algorithms, producing a total of 24 classified maps. Classified maps are referred to in this study using the following naming convention: Satellite YYYYMMDD algorithm (e.g., S2 20170309 ML for a Sentinel 2 image acquired on March 3, 2017 classified using the maximum likelihood algorithm).
Although we were primarily interested in mapping L. sinense distribution, the classification algorithms require multiple land cover types in the analysis for comparison. We included the following land cover types: L. sinense invaded hardwoods, uninvaded hardwoods, swamp, open water, fields, and pine stands. We delineated 3 training polygons for each land cover type based on prior knowledge of the study site, visual interpretation of the satellite imagery, and (in rare cases) ground surveys (Fig. 1). Training polygons for the L. sinense invaded category were primarily in areas with significant L. sinense cover, although we did not measure cover or set specific thresholds for the training areas. We refined the training polygons by conducting a series of informal trial-and-error classifications (primarily using ML and MD algorithms) on a subset of our L8 and S2 imagery. We adjusted the training polygons—and thus the training spectral signatures—as necessary until these initial classification attempts showed an adequate level of accuracy. This adaptive approach to creating and refining the training polygons is similar to what a land manager would use when applying this technique. Once we were satisfied with the training polygons, we calculated a separate set of training signatures for each L8 and S2 scene and ran the final classification algorithms.
2.4 Accuracy assessment
We conducted an accuracy assessment with reference data based on 250 random points surveyed during late winter/spring 2019 (Fig. 1). For each random point we sampled 2 plots, one corresponding to the nearest L8 pixel and one corresponding to the nearest S2 pixel (two S2 plots were excluded, because they fell outside the property boundary). The L8 plots were 30 m in diameter and the S2 plots were 10 m in diameter, which allowed land cover to be assessed at the pixel scale for each satellite image type. We navigated to the center point of each plot via global positioning system (GPS) receiver and visualized the edges of the plot using a Nikon Forestry 550 laser range finder (Nikon Vision CO., Ltd, Tokyo, Japan). We used a Garmin 64st recreational grade GPS receiver (Garmin Ltd., Olathe, Kansas, US) for the first 74 points; however concerns over potentially low positional accuracy led us to switch to a Trimble Geo7x GPS receiver for the final 176 points (Trimble Inc., Sunnyvale, California, US). We used circular plots rather than square plots for the sake of convenience. At each plot we recorded the land cover type and visually estimated the percent L. sinense cover within the plot. An informal assessment of classification accuracy differences between plots surveyed using the two GPS receivers did not reveal a significant difference.
We were specifically interested in L. sinense classification accuracy, so we recoded the maps into a binary invaded/uninvaded scheme. We assessed how L. sinense cover affected classification accuracy by using a range of thresholds (1, 10, 20, 30, 40, 50, 60, 70, 80, and 90%) as the cut-offs for what would be classified as an invaded plot in the reference data. For the lowest threshold (1%) we classified a field plot as L. sinense invaded if it had any L. sinense plants, even a single individual. For higher thresholds (e.g., 40%) we only classified the plot as invaded in the reference data if it had L. sinense cover equal to or greater than the threshold. This helped determine how the classified maps should be interpreted (i.e., is this a map of all L. sinense on the property or a map of areas with greater than X% L. sinense cover?). We calculated overall accuracy, user’s accuracy, and producer’s accuracy for each map at each threshold, and the results were displayed using accuracy curves [32], color coded based on image month, satellite type, and classification algorithm. The purpose of these accuracy curves is to demonstrate general trends related to these categories. Overall accuracy was calculated using Eq. 1:
where TP = true positive (the map and the reference data agree that L. sinense is present), TN = true negative (the map and the reference data agree L. sinense is absent), and Total = the total number of plots [33, 34]. Producer’s accuracy was calculated using Eq. 2:
where FN = false negative (the map predicts L. sinense is absent but the reference data indicate it is present) [33, 34]. User’s accuracy was calculated based on Eq. 3:
where FP = false positive (the map says L. sinense is present but the reference data indicate it is absent) [33, 34]. Estimates of the area invaded by L. sinense were extracted from each map based on pixel counts and compared.
3 Results and discussion
We found that the various combinations of satellite type, image date, and classification algorithm tended to highlight the same general areas on the maps as invaded, although there were some variation among all maps and a few major exceptions. Classified map: S2 20170309 ML produced the highest overall accuracy of 92.3% at a L. sinense cover threshold of 40% (Fig. 2). This is on par with the overall accuracy (89.2%) achieved by the best model reported elsewhere [28], although they did not take into account cover thresholds in their presence/absence reference data and doing so may have improved their results.
Accuracy curves for the most accurate map in our study, a maximum likelihood classified Sentinel 2 image from 2017/03/09. The three curves demonstrate the trade-off between Ligustrum sinense cover threshold and the three accuracy types. Using a 40% cover threshold, the map can be interpreted as having 92.3% overall accuracy, 77.8% producer’s accuracy, and 79.5% user’s accuracy
It is worth noting that the S2 20170309 image played an important role in our adaptive training site development phase, in part because it showed the greatest visual contrast between invaded and uninvaded areas in the infrared false color composite (Fig. 1). Thus, the finding that S2 20170309 ML had the highest overall accuracy could be partially due to the fact that the training polygons were somewhat tailored to that image and classification algorithm. The fact that there was a strong visual contrast in the infrared false color composite also shows that there was a high degree of spectral separation in this image, which almost certainly played a role in the high accuracy as well.
The average estimate of invaded area across all maps was 670.91 ha (± 134.99 SD), excluding 3 of the maps that failed to produce useful estimates (Fig. 3). The estimate from the map with the highest overall accuracy (S2 20170309 ML) was 554.40 ha; however the differences in the optimal threshold levels interpreted from the accuracy curves of the different maps complicates comparisons of invaded areas.
Estimates of hectares invaded by Ligustrum sinense based on each of the classified maps. The dotted lines separate classified maps based on the same imagery. The average estimate was 670.91 ha (± 134.99 SD), excluding three maps based on maximum likelihood classification of Landsat 8 images that failed to produce useful results
3.1 Ligustrum sinense cover
By analyzing the accuracy curves for all three accuracy measures on a single graph we can evaluate the best L. sinense cover threshold for interpreting a particular map. For example, Fig. 2 shows that overall accuracy peaked at the 40% cover threshold while user’s and producer’s accuracy cross at 40% for S2 20170309 ML. This trade-off between user’s and producer’s accuracy occurred because changing the cover threshold affected the proportion of false positives and false negatives in the accuracy assessment. At low cover thresholds there were few false positives, because most of the plots where the map predicted L. sinense is present have at least some L. sinense, which is why user’s accuracy was high. However, there were more false negatives at low cover thresholds (hence the low producer’s accuracy), because at low L. sinense densities the spectral signature of the pixel is closer to that of an uninvaded site than that of a densely invaded site (which comprised most of the L. sinense invaded training polygons). As the cover threshold was increased, the number of false negatives dropped (and producer’s accuracy went up), because the software was more effective at detecting areas with higher L. sinense cover. However, false positives increased (and user’s accuracy went down) at higher thresholds, because the map predicted some areas as invaded that did not meet the L. sinense cover threshold and thus were classified as “uninvaded” in the reference data. Using Fig. 2, we can see that if we interpret S2 20170309 ML as a map of L. sinense presence/absence (based on a minimum threshold of only 1% L. sinense cover) we can only assume 66.9% overall accuracy and 34.9% producer’s accuracy, but 100% user accuracy. If we interpret the same map as a map of areas with at least 40% L. sinense cover, then we can assume an overall accuracy of 92.3%, 77.8% producer’s accuracy, and 79.5% user’s accuracy. This trade-off between the different accuracy metrics is different for each map, and we observed a wide range in the accuracy curves across our maps. For all maps, there was a significant increase in overall accuracy when moving from 1% to 10% cover threshold (Fig. 4), suggesting that this technique is not effective at detecting very low-density, incipient invasions. If detecting low-density invasions was the goal then creating training sites specifically tailored to those spectral signatures may help, but ultimately it may be necessary to use imagery with higher spatial and spectral resolution. Imagery with high spatial- and spectral resolution can improve detection of low density and/or spectrally indistinct species; however it may be less efficient at mapping high density invasions and is less practical for land managers to utilize due to high costs and technical complexities [35, 36].
3.2 Satellite type
The accuracy curves in Fig. 4 are color coded to represent the maps based on S2 and L8 imagery. We generally observed higher overall accuracy and producer’s accuracy in the S2 maps, although there were a few exceptions (Fig. 4) and the relationship for user’s accuracy was less clear. Both the higher spatial and spectral resolutions of the S2 imagery likely played a role in improving accuracy. Higher spatial resolution (i.e., smaller pixels) reduces the prevalence of mixed pixels, or pixels that represent more than one cover type on the ground. Mixed pixels may be more likely to be misclassified both by the mapping software and the ground surveyor. The increase in spectral resolution (i.e., more bands) increases the amount of information in each spectral signature, allowing better differentiation of similar land cover types. The finding that S2 performed better than L8 is similar to that of previous studies that used other classification algorithms to detect a variety of land cover types [37,38,39,40].
3.3 Classification algorithm
The accuracy curves in Fig. 5 are the same as Fig. 4, but are color coded to represent the classification algorithm used to create each map. The ML algorithm generally performed well, with a few notable exceptions. The SA and MD algorithms did not show a clear pattern of difference. Three out of the four ML based L8 maps failed to produce useable results (e.g., they predicted nearly complete coverage of water or fields) and were omitted from Figs. 4, 5, and 6. So while ML appeared to be the best option for analyzing the S2 imagery, it was not a great option for the L8 imagery. This may be because the ML algorithm requires adequate training sample sizes to calculate a covariance matrix [29], and since the L8 imagery has a coarser spatial resolution there are fewer pixels per training site and thus fewer training pixels in the training sample. However, past experience has shown that the SCP provides an explicit warning when the training sample is too small for the covariance matrix to be calculated and such a warning was not given during these classifications. Other researchers have also found the ML method to be robust to small training samples [41]. Thus, the reason for the poor ML performance on L8 imagery in our study is unknown.
Moderate density pine mixed with hardwoods was sometimes confused as L. sinense invaded hardwoods on several of the maps. The ML algorithm was less prone to making this mistake, as demonstrated in Fig. 7, although satellite type and month also appeared to play a role. Patches of native evergreen hardwoods such as American holly (Ilex opaca) were also confused as L. sinense on many of the maps; however there was not as clear of a relationship with classification algorithm as there was with the moderate density pine associated errors.
Comparison of different classification algorithms applied to a Sentinel 2 image collected on 2017/03/09. The circle on the east side of the maps shows an area of moderate density pine forest that was partially confused as Ligustrum sinense invaded forest by the minimum distance and spectral angle algorithms, but not the maximum likelihood algorithm
3.4 Imagery acquisition date
Imagery collected in March generally had higher overall and user’s accuracy, with some exceptions, and there was not a clear pattern with producer’s accuracy (Fig. 6). These findings tend to confirm those of others that found early- to mid-March imagery generally performed best due to greater phenological differences between the L. sinense and deciduous overstory [28]. At our site, we observed that much of the L. sinense had a brief leaf drop in late January and early February that was followed by a flush of fresh growth by late February and early March, during which time most of the hardwood canopy was still dormant or just beginning bud break. The exact timing of the optimal phenological differences between L. sinense and the overstory is dependent on local climate and annual weather patterns. Sentinel 2 imagery has a higher temporal resolution (i.e., shorter revisit time), meaning that it is more likely that cloud-free imagery will be available during the period of greatest phenological difference. The revisit time for the pair of S2 satellites is about 5 days, while L8 has a 16-day revisit time [26, 27].
3.5 Comparisons to previous studies
From an ease of use perspective, supervised classification of moderate resolution imagery within QGIS appears to be preferable over the previous methods developed for mapping L. sinense invasions [i.e., 17, 18, 28]. This method does not require potentially expensive high-resolution imagery or tedious manual interpretation across an entire study area. The point-and-click user interface of QGIS is also simpler than the programming-based technique implemented by other methods [18, 28]. Moderate resolution multispectral imagery such as S2 and L8 are typically cheaper and easier to use than the LiDAR data used by others [18].
Comparing accuracy among studies is more difficult. Comparisons to [17] are not possible, because that study did not include a formal accuracy assessment. The top models from other comparable studies [18, 28] had overall accuracies of 88.89% and 89.2%, respectively. These overall accuracies are similar to that of our top performing classifier (92.3% at a cover threshold of 40% for S2 20170309 ML); however it is important to consider that while these other studies [18, 28] collected data on L. sinense cover within field plots, it does not appear that they set a cover threshold when computing their accuracy statistics. This means that a more equitable comparison may be to use the 66.9% overall accuracy of S2 20170309 ML at a cover threshold of 1%. While this does not compare favorably to the accuracies from the other studies [18, 28], differences in training datasets may be partly responsible. Our training areas were mostly in areas with moderate to heavy L. sinense cover, so it is no surprise that our classifiers were more accurate at identifying areas with similar levels of infestation. If the goal were to identify areas with low L. sinense cover then including representative training sites may be helpful, although the moderate resolution of both S2 and L8 imagery will be a limiting factor for identifying very low-density invasions. Despite the limitations for identifying low-density invasions, the high accuracy achieved at moderate to high densities show that this is a useful method for land managers planning control measures in such conditions.
4 Conclusions
Our results show that free and simple remote sensing tools can be used to effectively map relatively dense L. sinense stands in deciduous hardwood forests, although the method is likely inadequate for low-density incipient invasions. We also discovered that there can be significant variation in accuracy results based on the type of satellite imagery, the date of the image acquisition, and the classification algorithm, even when the same training sites are used. Choosing appropriate training sites also has a large impact on accuracy, although that was not formally assessed in this study. We tended to find that S2 imagery acquired in March and processed using the ML algorithm performed well, although these patterns may not hold true for all situations. We recommend that land managers interested in deploying this method use an adaptive process for map development that includes testing at least a few variations of training sites, images, and classification algorithms to find what works best on a particular site, using our results as a guide. A “multiple classifier system” approach that combines the results of multiple classifications could be a useful way to handle uncertainty in choosing the best map [42], but further evaluation is needed to determine whether such a technique could be easily implemented in QGIS. The results of our study have more general implications for land management and remote sensing as well, in that they show that sometimes simple, open-source methods can be used in place of more complicated or expensive methods, which opens the door for land managers with limited budgets or remote sensing experience to increase their use of remote sensing for informing management decisions. Following this research, we worked with the Alabama Cooperative Extension System to publish a step-by-step introduction to remote sensing that is targeted toward land managers and uses L. sinense mapping as a primary example [43].
References
CABI (2018) Ligustrum sinense [original text by D Shaw]. In: Invasive species compendium. CAB International, Wallngford. https://www.cabi.org/isc/datasheet/30763. Accessed 1 Mar 2019
Miller JH, Chambliss EB, Bargeron CT (2004) Invasive plants of the thirteen southern states. https://www.invasive.org/south/seweeds.cfm?sort=3. Accessed 1 Mar 2019
Miller JH, Chambliss EB (2008) Estimates of acres covered of nonnative invasive plants in southern forests. Auburn, AL: US Department of Agriculture, Forest Service, Southern Research Station. https://www.invasive.org/fiamaps/summary.pdf. Accessed 1 Mar 2019
Klepac J, Rummer RB, Hanula JL, Horn S (2007) Mechanical removal of Chinese privet. U.S. Department of Agriculture, Forest Service, Southern Research Station SRS-43, Asheville
Benez-Secanho FJ, Grebner DL, Ezell AW, Grala RK (2018) Financial trade-offs associated with controlling Chinese privet (Ligustrum sinense Lour.) in forestlands in the southern USA. J Forest 116(3):236–244
Miller JH, Miller KA (2005) Forest plants of the Southeast and their wildlife uses, Revised edn. University of Georgia Press, Athens, p 360
Maddox V, Byrd J Jr, Serviss B (2010) Identification and control of invasive privets (Ligustrum spp.) in the middle southern United States. Invas Plant Sci Mana 3(4):482–488
Foard M (2014) Causes and Consequences of Chinese privet (Ligustrum sinense Lour.) invasion in hydrologically altered forested wetlands. Master’s thesis. Arkansas State University, Jonesboro
Wilcox J, Beck CW (2007) Effects of Ligustrum sinense Lour. (Chinese privet) on abundance and diversity of songbirds and native plants in a southeastern nature preserve. Southeast Nat 6(3):535–550
Green BT, Blossey B (2012) Lost in the weeds: Ligustrum sinense reduces native plant growth and survival. Biol Invasions 14:139–150
Merriam RW, Feil E (2002) The potential impact of an introduced shrub on native plant diversity and forest regeneration. Biol Invasions 4:369–373
Lowenstein NJ, Lowenstein EF (2005) Non-native plants in the understory of riparian forests across a land use gradient in the Southeast. Urban Ecosyst 8:79–91
Hart JL, Holmes BN (2013) Relationships between Ligustrum sinense invasion, biodiversity, and development in a mixed bottomland forest. Invas Plant Sci Mana 6(1):175–186
Grove E, Clarkson BD (2005) An ecological study of Chinese privet (Ligustrum sinense Lour.) in the Waikato region. Centre for Biodiversity and Ecology Research Contract Report No. 41, Hamilton
Cofer MS, Walck JL, Hidayati SN (2008) Species richness and exotic species invasion in middle Tennessee cedar glades in relation to abiotic and biotic factors. J Torrey Bot Soc 135(4):540–553
Pokswinki SM (2009) Invasive characteristics of Chinese privet (Ligustrum sinense Lour.) in a bay swamp in the fall line hills of east-central Alabama. Master’s thesis, Auburn University, Auburn
Ward RW (2002) Extent and dispersal rates of Chinese privet (Ligustrum sinense) invasion in the upper Oconee River floodplain, north Georgia. Southeast Geogr 42(1):29–48
Singh KK, Davis AJ, Meentemeyer RK (2015) Detecting understory plant invasion in urban forests using LiDAR. Int J Appl Earth Obs 38:267–279
Kelly M, Tommaso SD (2015) Mapping forests with lidar provides flexible, accurate data with many uses. Calif AGR 69(1):14–20
Xie Y, Sha Z, Yu M (2008) Remote sensing imagery in vegetation mapping: a review. J Plant Ecol 1(1):9–23
Gómez C, White JC, Wulder MA (2016) Optical remotely sensed time series data for land cover classification: a review. ISPRS J Photogramm 116:55–72
Phiri D, Morgenroth J (2017) Development in Landsat land cover classification methods: a review. Remote Sens 9:967–992
Resasco J, Hale AN, Henry MC, Gorchov DL (2007) Detecting an invasive shrub in a deciduous forest understory using late-fall Landsat sensor imagery. Int J Remote Sens 28(16):3739–3745
Knipling EB (1970) Physical and physiological basis for the reflectance of visible and near-infrared radiation from vegetation. Remote Sens Environ 1:155–159
National Aeronautics and Space Administration (NASA) (1999) Land cover classification: how scientists differentiate between land cover types. https://earthobservatory.nasa.gov/features/LandCover/land_cover_2.php. Accessed 1 Mar 2019
European Space Agency (ESA) (2015) Sentinel-2 user handbook. European Space Agency, Paris, p 64
US Geological Survey (USGS) (2019) Landsat 8 (L8) data user’s handbook. 4th ed. US Geological Survey, Earth Resources Observation and Science Center, LSDS-1574, Sioux Falls
Singh KK, Chen Y, Smart L, Gray J, Meentemeyer RK (2018) Intra-annual phenology for detecting understory plant invasion in urban forests. ISPRS J Photogramm 142:151–161
Congedo L (2019) Semi-automatic plugin documentation. Release 6.2.0.1. https://fromgistors.blogspot.com/p/user-manual.html. Accessed 1 Mar 2019
QGIS Development Team (2019) QGIS geographic information system. Open Source Geospatial Foundation Project. http://qgis.osgeo.org/. Accessed 1 Mar 2019
Song C, Woodcock CE, Seto KC, Lenney MP, Macomber SA (2001) Classification and change detection using Landsat TM data: when and how to correct atmospheric effects. Remote Sens Environ 75:230–244
Morisette JT, Khorram S (2000) Accuracy assessment curves for satellite-based change detection. Photogramm Eng Remote Sens 66(7):875–880
Congalton RG (1991) A review of assessing the accuracy of classifications of remotely sensed data. Remote Sens Environ 37:35–46
Sanchez-Hernadez C, Boyd DS, Foody GM (2007) One-class classification for mapping a specific land-cover class: SVDD classification of fenland. IEEE Trans Geosci Remote 45(4):1061–1073
Underwood EC, Ustin SL, Ramirez CM (2007) A comparison of spatial and spectral image resolution for mapping invasive plants in coastal California. Environ Manag 39(1):63–83
He KS, Rocchini D, Neteler M, Nagendra H (2011) Benefits of hyperspectral remote sensing for tracing plant invasions. Divers Distrib 17:381–392
Pesaresi M, Corbane C, Julea A, Florczyk AJ, Syrris V, Soille P (2016) Assessment of the added-value of Sentinel-2 for detecting built-up areas. Remote Sens 8:299. https://doi.org/10.3390/rs8040299
Sibanda M, Mutanga O, Rouget M (2016) Discriminating rangeland management practices using simulated HyspIRI, Landsat 8 OLI, Sentinel 2 MSI, and VENµS spectral data. IEEE J Sel Top Appl 9(9):3957–3969
Forkuor G, Dimobe K, Serme I, Tondoh JE (2017) Landsat-8 vs. Sentinel 2: examining the added value of Sentinel-2’s red edge bands to land-use and land-cover mapping in Burkina Faso. GISci Remote Sens 55(3):331–354
Sothe C, Almeida CMD, Liesenberg V, Schimalski MB (2017) Evaluating Sentinel-2 and Landsat-8 data to map successional forest stages in a subtropical forest in southern Brazil. Remote Sens 9:838. https://doi.org/10.3390/rs9080838
Li C, Wang J, Wang L, Hu L, Gong P (2014) Comparison of classification algorithms and training sample sizes in urban land classification with Landsat thematic mapper imagery. Remote Sens 6:964–983
Du P, Xia J, Zhang W, Tan K, Liu Y, Liu S (2012) Multiple classifier system for remote sensing image classification: a review. Sensors 12:4764–4792
Cash J, Loewenstein N, Anderson C, Brodbeck A (2020) An introduction to satellite imagery analysis for land managers. Alabama Cooperative Extension System FOR-2072. 21 p
Funding
Research assistantship funding for JSC was provided by the Harry Murphy Dean’s Enhancement Fund for Excellence in The School of Forestry and Wildlife Sciences at Auburn University. Project funding was provided by EBSCO Industries, Inc. Additional funding was provided by the Birmingham Audubon Society through the Walter F. Coxe Research Fund, the Alabama Invasive Plant Council, and The Curtis & Edith Munson Foundation.
Author information
Authors and Affiliations
Contributions
JSC collected data and lead writing process. CJA and LM assisted with study design and manuscript edits.
Corresponding author
Ethics declarations
Conflict of interest
The author’s declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cash, J.S., Anderson, C.J. & Marzen, L. Evaluating free and simple remote sensing methods for mapping Chinese privet (Ligustrum sinense) invasions in hardwood forests. SN Appl. Sci. 2, 789 (2020). https://doi.org/10.1007/s42452-020-2596-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-020-2596-4