Abstract
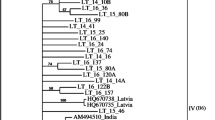
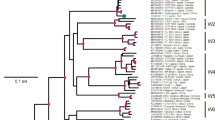
Apple chlorotic leaf spot virus (ACLSV) infects temperate rosaceous fruit trees worldwide and causes a wide range of diseases that are economically highly damaging. This study was carried out to analyze genetic variability of regional ACLSV isolates and compare it with ACLSV isolates from other parts of the world. Nineteen amplicons of ACLSV, corresponding to the coat protein (CP) gene of isolates from apple, plum, and nectarine, from two states in southern Brazil, have been analyzed for genetic variation and compared phylogenetically among themselves and with other sequences available in GenBank. Sequences identities among complete CP genes of these isolates ranged from 87.5 to 100% and 92.2 to 100% at the nucleotide (nt) and the deduced amino acid (daa) levels, respectively. In the comparison with isolates from Asia, Europe and North America, identities were 68.4 to 100% and 72.5 to 100% at nt and daa levels, respectively. Phylogenetic trees based on nucleotide sequences showed that these isolates grouped into two clusters, cluster 1 containing apple isolates and cluster 2 comprising apple, plum and nectarine isolates. Most Brazilian isolates showed conserved signatures (Ser40, Leu59, Tyr75, Thr130 and Leu184) in their CPs, which place them with type B6 isolates. However, some Brazilian isolates were found to be variants of type B6. These analyzes indicated that Brazilian isolates had lower genetic variability compared to isolates from China, India and Japan and that the CP genes were under negative selection. The greatest diversity of nucleotides was observed in the central portion of the CP gene, represented predominantly by synonymous substitutions. One natural recombinant was detected among ACLSV isolates from Brazil.



Similar content being viewed by others
References
Al Rwahnih M, Turturo C, Minafra A, Saldarelli P, Myrta A, Pallás V, Savino V (2004) Molecular variability of Apple chlorotic leaf spot virus in different hosts and geographical regions. Journal of Plant Pathology 86:117–122
Altschuh D, Lesk AM, Bloomer AC, Klug A (1987) Correlation of co-ordinated amino acid substitutions with function in viruses related to tobacco mosaic virus. Journal of Molecular Biology 193:693–707
Bonnal S, Schaeffer C, Creancier L, Clamens S, Moine H, Prats AC, Vagner S (2003) A single internal ribosome entry site containing a G quartet RNA structure drives fibroblast growth factor 2 gene expression at four alternative translation initiation codons. Journal of Biological Chemistry 278:39330–39336
Cembali T, Folwell RJ, Wandschneider P, Eastwell KC, Howell WE (2003) Economic implications of a virus prevention program in deciduous tree fruits in the USA. Crop Protection 22:1149–1156
Chare ER, Holmes EC (2004) Selection pressures in the capsid genes of plant RNA viruses reflect mode of transmission. Journal of General Virology 85:3149–3157
Chen S, Zhou Y, Ye T, Hao L, Guo L, Fan Z, Li S, Zhou T (2013) Genetic variation analysis of apple chlorotic leaf spot virus coat protein reveals a new phylogenetic type and two recombinants in China. Archives of Virology 159:1431–1438
Fajardo TVM, Silva FN, Eiras M, Nickel O (2017) High-throughput sequencing applied for the identification of viruses infecting grapevines in Brazil and genetic variability analysis. Tropical Plant Pathology 42:250–260
Gadiou S, Kundu JK, Paunovic S, Garcia-Diez P, Komorowska B, Gospodaryk A, Handa A, Massart S, Birisik N, Takur PD, Polischuk V (2010) Genetic diversity of flexiviruses infecting pome fruit trees. Journal of Plant Pathology 92:685–691
Garcia-Arenal F, Fraile A, Malpica JM (2001) Variability and genetic structure of plant virus populations. Annual Review of Phytopathology 39:157–186
German S, Delbos RP, Candresse T, Lanneau M, Dunez J (1990) Complete nucleotide sequence of the genome of a severe cherry isolate of Apple chlorotic leaf spot trichovirus (ACLSV). Archives of Virology 142:833–841
German-Retana S, Bergey B, Delbos RP, Candresse T, Dunez J (1997) Complete nucleotide sequence of the genome of a severe cherry isolate of apple chlorotic leaf spot trichovirus (ACLSV). Archives of Virology 142:833–841
Guo W, Zheng W, Wang M, Li X, Ma Y, Dai H (2016) Genome sequences of three apple chlorotic leaf spot virus isolates from hawthorns in China. PLoS One 11:e0161099
Hadidi A, Barba M (2011) Economic impact of pome fruit and stone fruit viruses and viroids. In: Hadidi A, Barba M, Candresse T, Jelkmann W (eds) Virus and virus-like diseases of pome and stone fruits. APS Press, St. Paul, pp 1–7
Hong JJ, TY W, Chang TY, Chen CY (2013) Viral IRES prediction system - a web server for prediction of the IRES secondary structure in silico. PLoS One 8:e79288
Ivanov PA, Karpova OV, Skulachev MV, Tomashevskaya OL, Rodionova NP, Dorokhov YL, Atabekov JG (1997) A tobamovirus genome that contains an internal ribosome entry site functional in vitro. Virology 232:32–43
Jelkmann W (1996) The nucleotide sequence of a strain of Apple chlorotic leaf spot virus (ACLSV) responsible for plum pseudo-pox and its relation to an apple and plum bark split strain. Phytopathology 86:101
Jelkmann W, Kunze L (1995) Plum pseudopox in German prune after infection with an isolate of Apple chlorotic leafspot virus causing plum line pattern. Acta Horticulturae 386:122–125
Lima ATM, Sobrinho RR, Gonzalez-Aguilera J, Rocha CS, Silva SJC, Xavier CAD, Silva FN, Duffy S, Zerbini FM (2013) Synonymous site variation due to recombination explains higher genetic variability in begomovirus populations infecting non-cultivated hosts. Journal of General Virology 94:418–431
Lima ATM, Silva JCF, Silva FN, Urquiza GPC, Silva FF, Seah YM, Mizubuti ESG, Duffy S, Zerbini FM (2017) The diversification of begomovirus populations is predominantly driven by mutational dynamics. Virus Evolution 3:vex005
Mackenzie DJ, Maclean MA, Ukerji S, Green M (1997) Improved RNA extraction from woody plants for the detection of viral pathogens by reverse transcription-polymerase chain reaction. Plant Disease 81:222–226
Marini DB, Gibson PG, Scott SW (2008) The complete nucleotide sequence of an isolate of apple chlorotic leaf spot virus from peach [Prunus persica (L.) batch]. Archives of Virology 153:1003–1005
Martin DP, Murrell B, Golden M, Khoosal A, Muhire B (2015) RDP4: Detection and analysis of recombination patterns in virus genomes. Virus Evolution 1:vev003
Martínez-Salas E, Pacheco A, Serrano P, Fernandez N (2008) New insights into internal ribosome entry site elements relevant for viral gene expression. Journal of General Virology 89:611–626
Moura CJM, Fajardo TVM, Eiras M, Silva FN, Nickel O (2018) Molecular characterization of GSyV-1 and GLRaV-3 and prevalence of grapevine viruses in a grape-growing area. Scientia Agricola 75(1):43–51
Németh M (1986) Virus, mycoplasma and rickettsia diseases of fruit trees. Martinus Nijhoff Publ, Dordrecht
Nickel O, Fajardo TVM, Jelkmann W, Kuhn GB (2001) Sequence analysis of the capsid protein gene of an isolate of Apple stem grooving virus, and its survey in southern Brazil. Fitopatologia Brasileira 26:655–659
Niu F, Pan S, Wu Z, Jiang D, Li S (2012) Complete nucleotide sequences of the genomes of two isolates of apple chlorotic leaf spot virus from peach (Prunus persica) in China. Archives of Virology 157:783–786
Rana T, Chandel V, Kumar Y, Ram R, Hallan V, Zaidi AA (2010) Molecular variability analyses of Apple chlorotic leaf spot virus capsid protein. Journal of Biosciences 35:605–615
Rocha CS, Castillo-Urquiza GP, Lima ATM, Silva FN, Xavier CAD, Hora-Junior BT, Beserra-Junior JEA, Malta AWO, Martin DP, Varsani A, Alfenas-Zerbini P, Mizubuti ESG, Zerbini FM (2013) Brazilian begomovirus populations are highly recombinant, rapidly evolving, and segregated based on geographical location. Journal of Virology 87:5784–5799
Rott ME, Jelkmann W (2001) Characterization and detection of several filamentous viruses of cherry: adaptation of an alternative cloning method (DOP-PCR), and modification of an RNA extraction protocol. European Journal of Plant Pathology 107:411–420
Rozas J, Sanchez-Delbarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Russel DW, Sambrook J (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Sanger F, Coulson AR (1975) A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. Journal of Molecular Biology 94:441–448
Sato K, Yoshikawa N, Takahashi T (1993) Complete nucleotide sequence of the genome of an apple isolate of Apple chlorotic leaf spot virus. Journal of General Virology 74:1927–1931
Silva FN, Nickel O, Fajardo TVM, Bogo A (2008) Indexação biológica múltipla e RT-PCR para detecção de vírus latentes em macieiras. Tropical Plant Pathology 33:157–161
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 30: 2725–2729
Yaegashi H, Isogai M, Tajima H, Sano T, Yoshikawa N (2007) Combinations of two amino acids (Ala40 and Phe75 or Ser40 and Tyr75) in the coat protein of apple chlorotic leaf spot virus are crucial for infectivity. Journal of General Virology 88:2611–2618
Yaegashi H, Yoshikawa N, Candresse T (2011) Apple chlorotic leaf spot virus in pome fruits. In: Hadidi A, Barba M, Candresse T, Jelkmann W (eds) Virus and virus-like diseases of pome and stone fruits. APS Press, St. Paul, pp 17–21
Yanase H, Mink GI, Sawamura K, Yamaguchi A (1990) Apple topworking disease. In: Jones AL, Aldwinckle HS (eds) Compendium of apple and pear diseases. APS Press, St. Paul, pp 74–75
Yoshikawa N (2001) Apple chlorotic leaf spot virus. Commonwealth Mycological Institute (CMI)/Association of Applied Biologists (AAB) Descriptions of Plant Viruses 386, no. 30 revised
Zanardo LG, Silva FN, Lima AT, Milanesi DF, Castilho-Urquiza GP, Almeida AM, Zerbini FM, Carvalho CM (2014) Molecular variability of cowpea mild mottle virus infecting soybean in Brazil. Archives of Virology 159:727–737
Acknowledgements
This work was financed by Embrapa project nr. 03.13.05.007. Authors thank Francisco J.L. Aragão (Embrapa Recursos Genéticos e Biotecnologia, Brasília, DF) for sequencing, Marcos F. Vanni for technical support, and CNPq for financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Jorge Rezende
Electronic supplementary material
Supplementary Table S1
(DOCX 30.4 kb)
Supplementary Table S2
(DOCX 18.4 kb)
Supplementary Table S3
(DOCX 19.8 kb)
Rights and permissions
About this article
Cite this article
Nickel, O., Silva, F.N., Fajardo, T.V.M. et al. Characterization and genetic variability of coat protein genes of Apple chlorotic leaf spot virus isolates from southern Brazil. Trop. plant pathol. 43, 109–116 (2018). https://doi.org/10.1007/s40858-017-0197-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-017-0197-6