Abstract
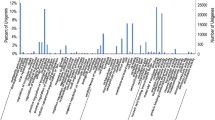
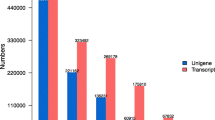
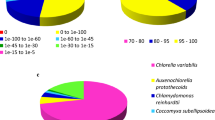
Nitrogen is a critical macronutrient for plant growth and development, and Dunaliella salina has various physiological responses to nitrogen deprivation, such as accumulating large amounts of carotenoids in its chloroplasts. However, little is known about the underlying global molecular response of D. salina to nitrogen deprivation. Herein, we used the Illumina platform to interrogate which genes of D. salina respond to nitrogen deprivation. The RNA-seq libraries of D. salina TG cultured under nitrogen-replete and -depleted conditions for 7 days generated 87,160,704 valid reads with an average length of 124.94 bp. De novo assembly produced 73,329 transcripts with an average length of 801 bp and 47,283 unigenes with an average length of 675 bp. 5803 unigenes were assigned to 50 gene ontology (GO) terms belonging to three main categories. A total of 6016 unigenes with 671 enzyme commission numbers were assigned to 251 predicted KEGG metabolic pathways, and 6296 unigenes were categorized into 25 KOG classifications. A total of 27,306 CDS were predicted from the 47,283 unigenes. Analysis of transcription levels revealed that 2380 genes were upregulated and 747 were downregulated during nitrogen deprivation. These nitrogen-deprivation responsive genes and differentially expressed transcriptional regulator genes were identified by GO classification. The transcriptome data may serve as a reference for further analysis of gene expression and functional genomics studies, and will facilitate the study of carotenoid biosynthesis and nitrogen metabolism in chlorophytes at the molecular level.







Similar content being viewed by others
References
Bairoch A, Boeckmann B (1992) The SWISS-PROT protein sequence data bank. Nucleic Acids Res 20(Suppl):2019–2022
Blaby IK, Glaesener AG, Mettler T, Fitz-Gibbon ST, Gallaher SD, Liu B, Boyle NR, Kropat J, Stitt M, Johnson S, Benning C, Pellegrini M, Casero D, Merchant SS (2013) Systems-level analysis of nitrogen starvation-induced modifications of carbon metabolism in a Chlamydomonas reinhardtii starchless mutant. Plant Cell 25:4305–4323
Burge C, Karlin S (1997) Prediction of complete gene structures in human genomic DNA. J Mol Biol 268:78–94
Cai H, Lu Y, Xie W, Zhu T, Lian X (2012) Transcriptome response to nitrogen starvation in rice. J Biosciences 37:731–747
Coesel SN, Baumgartner AC, Teles LM, Ramos AA, Henriques NM, Cancela L, Varela JC (2008) Nutrient limitation is the main regulatory factor for carotenoid accumulation and for Psy and Pds steady state transcript levels in Dunaliella salina Chlorophyta exposed to high light and salt stress. Mar Biotechnol 10(5):602–611
Consortium GO (2006) The gene ontology GO Project in 2006. Nucleic Acids Res 34:D322
de Lomana ALG, Schäuble S, Valenzuela J, Imam S, Carter W, Bilgin DD, Yohn CB, Turkarslan S, Reiss DJ, Orellana MV (2015) Transcriptional program for nitrogen starvation-induced lipid accumulation in Chlamydomonas reinhardtii. Biotechnol Biofuels 8:207
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Emanuelsson O, Nielsen H, Von HG (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8:978
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) The Pfam protein families database. Nucleic Acids Res 42:D222–D230
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, di Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480–D484
Krapp A, Daniel-Vedele F (2011) Arabidopsis roots and shoots show distinct temporal adaptation patterns toward nitrogen starvation. Plant Physiol 157:1255–1282
Lamers PP, Janssen M, De Vos RC, Bino RJ, Wijffels RH (2008) Exploring and exploiting carotenoid accumulation in Dunaliella salina for cell-factory applications. Trends Biotechnol 26:631–638
Lamers PP, Janssen M, De Vos RC, Bino RJ, Wijffels RH (2012) Carotenoid and fatty acid metabolism in nitrogen-starved Dunaliella salina, a unicellular green microalga. J Biotechnol 162:21–27
Lang JD, Hendricks WPD (2018) Identification of driver mutations in rare cancers: the role of SMARCA4 in small cell carcinoma of the ovary, hypercalcemic type (SCCOHT). Methods Mol Biol 1706:367–379
Lers A, Biener Y, Zamir A (1990) Photoinduction of massive β-carotene accumulation by the alga Dunaliella bardawil kinetics and dependence on gene activation. Plant Physiol 93:389–395
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform 12:323
Liang C, Cao S, Zhang X, Zhu B, Su Z, Dong X, Guang X, Ye N (2013) De novo sequencing and global transcriptome analysis of Nannochloropsis sp. Eustigmatophyceae following nitrogen starvation. Bioenerg Res 6:494–505
Lv H, Qu G, Qi X, Lu L, Tian C, Ma Y (2013) Transcriptome analysis of Chlamydomonas reinhardtii during the process of lipid accumulation. Genomics 101(4):229–237
Lv H, Cui X, Wahid F, Xia F, Zhong C, Jia S (2016a) Analysis of the physiological and molecular responses of Dunaliella salina to macronutrient deprivation. PLoS ONE 11:e0152226
Lv H, Cui X, Wang S, Jia S (2016b) Metabolic profiling of Dunaliella salina shifting cultivation conditions to nitrogen deprivation. Metab Open Acess 6:2153
Lv H, Cui X, Tan Z, Jia S (2017) Analysis of metabolic responses of Dunaliella salina to phosphorus deprivation. J Appl Phycol 29:1251–1260
Mamedov TG, Moellering ER, Chollet R (2005) Identification and expression analysis of two inorganic C- and N-responsive genes encoding novel and distinct molecular forms of eukaryotic phosphoenolpyruvate carboxylase in the green microalga Chlamydomonas reinhardtii. Plant J 42:832–843
Miller R, Wu G, Deshpande RR, Vieler A, Gärtner K, Li X, Moellering ER, Zäuner S, Cornish AJ, Liu B, Bullard B, Sears BB, Kuo MH, Hegg EL, Shachar-Hill Y, Shiu SH, Benning C (2010) Changes in transcript abundance in Chlamydomonas reinhardtii following nitrogen deprivation predict diversion of metabolism. Plant Physiol 154:1737–1752
Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Oren A (2005) A hundred years of Dunaliella research: 1905–2005. Saline syst 1:1–14
Polle JEW, Barry K, Cushman J, Schmutz J, Tran D, Hathwaik LT, Yim WC, Jenkins J, McKie-Krisberg Z, Prochnik S, Lindquist E, Dockter RB, Adam C, Molina H, Bunkenborg J, Jin E, Buchheim M, Magnuson J (2017) Draft nuclear genome sequence of the halophilic and beta-carotene-accumulating green alga. Genome Announc. 5(43):e01105–e01117
Pruitt KD, Tatiana T, Maglott DR (2005) NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 33:D501–D504
Ramos A, Coesel S, Marques A, Rodrigues M, Baumgartner A, Noronha J, Rauter A, Brenig B, Varela J (2008) Isolation and characterization of a stress-inducible Dunaliella salina Lcy-beta gene encoding a functional lycopene beta-cyclase. Appl Microbiol Biot 79:819–828
Ramos AA, Marques AR, Rodrigues M, Henriques N, Baumgartner A, Castilho R, Brenig B, Varela JC (2009) Molecular and functional characterization of a cDNA encoding 4-hydroxy-3-methylbut-2-enyl diphosphate reductase from Dunaliella salina. J Plant Physiol 166:968–977
Ramos AA, Polle J, Tran D, Cushman JC, Jin E, Varela JC (2011) The unicellular green alga Dunaliella salina Teod. as a model for abiotic stress tolerance: genetic advances and future perspectives. Algae 26:3–20
Salguero A, de la Morena B, Vigara J, Vega JM, Vilchez C, León R (2003) Carotenoids as protective response against oxidative damage in Dunaliella bardawil. Biomol Eng 20:249–253
Seo J, Gordishdressman H, Hoffman EP (2006) An interactive power analysis tool for microarray hypothesis testing and generation. Bioinformatics 22:808
Simionato D, Block MA, La Rocca N, Jouhet J, Maréchal E, Finazzi G, Morosinotto T (2013) The response of Nannochloropsis gaditana to nitrogen starvation includes de novo biosynthesis of triacylglycerols, a decrease of chloroplast galactolipids, and reorganization of the photosynthetic apparatus. Eukaryot Cell 12:665–676
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28:33
von Wirén N, Lauter FR, Ninnemann O, Gillissen B, Walch-Liu P, Engels C, Jost W, Frommer WB (2000) Differential regulation of three functional ammonium transporter genes by nitrogen in root hairs and by light in leaves of tomato. Plant J. 21(2):167–175
Weng L-C, Pasaribu B, Lin I-P, Tsai C-H, Chen C-S, Jiang P-L (2014) Nitrogen deprivation induces lipid droplet accumulation and alters fatty acid metabolism in symbiotic dinoflagellates isolated from Aiptasia pulchella. Sci Rep 4:5777
Wickham H (2009) ggplot2: Elegant Graphics for Data Analysis. Springer Publishing Company, Incorporated. http://dl.acm.org/citation.cfm?id=SERIES?13087.1795559
Marchler-Bauer A, Bo Y, Han L, He J, Lanczycki CJ, Lu S, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Geer LY, Bryant SH (2016) CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45:D200–D203
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293
Zhang X-N, Qu Z-C, Wan Y-Z, Zhang H-W, Shen D-L (2002) Application of suppression subtractive hybridization (SSH) to cloning differentially expressed cDNA in Dunaliella salina (chlorophyta) under hyperosmotic shock. Plant Mol Biol Rep 20:49–57
Zschiedrich CP, Keidel V, Szurmant H (2016) Molecular mechanisms of two-component signal transduction. J Mol Biol 428:3752–3775
Acknowledgements
This work is supported by the Open Research Fund Program of Beijing Key Lab of Plant Resource Research and Development, Beijing Technology and Business University and the National Natural Science Foundation of China (No. 31401029).
Author information
Authors and Affiliations
Contributions
SJ and HL conceived and designed the project. HL analyzed the data and wrote the paper. BQ performed the cultures materials preparation and qPCR analysis. QW revised the paper and JH provided bioinformatic analysis tools. All authors have read and approved the final manuscript.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 2 (XLSX 359 kb) Supplement Table 2. KEGG Pathway Classifications of Unigenes.
40626_2019_138_MOESM3_ESM.xlsx
Supplementary material 3 (XLSX 218 kb) Supplement Table 3. Differential Expressions of Unigenes of Cells under CM and -N.
40626_2019_138_MOESM5_ESM.xlsx
Supplementary material 5 (XLSX 19 kb) Supplement Table 5. Pathway Classifications of DEGs by KEGG. S gene number: Numbers of significantly changed genes in a given KEGG pathway. B gene number: Numbers of annotated genes in a given KEGG pathway.
Rights and permissions
About this article
Cite this article
Lv, H., Wang, Qe., Qi, B. et al. RNA-Seq and transcriptome analysis of nitrogen-deprivation responsive genes in Dunaliella salina TG strain. Theor. Exp. Plant Physiol. 31, 139–155 (2019). https://doi.org/10.1007/s40626-019-00138-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40626-019-00138-w