Abstract
Background
The induction of neural regeneration is vital to the repair of spinal cord injury (SCI). While compared with peripheral nervous system (PNS), the regenerative capacity of the central nervous system (CNS) is extremely limited. This indicates that modulating the molecular pathways underlying PNS repair may lead to the discovery of potential treatment for CNS injury.
Methods
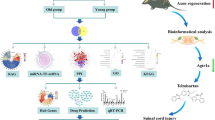
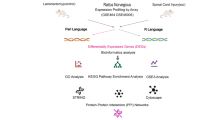
Based on the gene expression profiles of dorsal root ganglion (DRG) after a sciatic nerve injury, we utilized network guided forest (NGF) to rank genes in terms of their capacity of distinguishing injured DRG from sham-operated controls. Gene importance scores deriving from NGF were used as initial heat in a heat diffusion model (HotNet2) to infer the subnetworks underlying neural regeneration in the DRG. After potential regulators of the subnetworks were found through Connectivity Map (cMap), candidate compounds were experimentally evaluated for their capacity to regenerate the damaged neurons.
Results
Gene ontology analysis of the subnetworks revealed ubiquinone biosynthetic process is crucial for neural regeneration. Moreover, almost half of the genes in these subnetworks are found to be related to neural regeneration via text mining. After screening compounds that are likely to modulate gene expressions of the subnetworks, three compounds were selected for the experiment. Of them, trichostatin A, a histone deacetylase inhibitor, was validated to enhance neurite outgrowth in vivo via an optic nerve crush mouse model.
Conclusions
Our study identified subnetworks underlying neural regeneration, and validated a compound can promote neurite outgrowth by modulating these subnetworks. This work also suggests an alternative approach for drug repositioning that can be easily extended to other disease phenotypes.

Article PDF
Similar content being viewed by others
References
Hulsebosch, C. E. (2002) Recent advances in pathophysiology and treatment of spinal cord injury. Adv. Physiol. Educ., 26, 238–255
Smith, D. S. and Pate Skene, J. H. (1997) A transcription-dependent switch controls competence of adult neurons for distinct modes of axon growth. J. Neurosci., 17, 646–658
Silver, J. and Miller, J. H. (2004) Regeneration beyond the glial scar. Nat. Rev. Neurosci., 5, 146–156
McKerracher, L., David, S., Jackson, D. L., Kottis, V., Dunn, R. J. and Braun, P. E. (1994) Identification of myelin-associated glycoprotein as a major myelin-derived inhibitor of neurite growth. Neuron, 13, 805–811
Fitch, M. T. and Silver, J. (2008) CNS injury, glial scars, and inflammation: inhibitory extracellular matrices and regeneration failure. Exp. Neurol., 209, 294–301
Filbin, M. T. (2003) Myelin-associated inhibitors of axonal regeneration in the adult mammalian CNS. Nat. Rev. Neurosci., 4, 703–713
Wang, K. C., Koprivica, V., Kim, J. A., Sivasankaran, R., Guo, Y., Neve, R. L. and He, Z. (2002) Oligodendrocyte-myelin glycoprotein is a Nogo receptor ligand that inhibits neurite outgrowth. Nature, 417, 941–944
Schmitt, A. B., Breuer, S., Liman, J., Buss, A., Schlangen, C., Pech, K., Hol, E. M., Brook, G. A., Noth, J. and Schwaiger, F.-W. (2003) Identification of regeneration-associated genes after central and peripheral nerve injury in the adult rat. BMC Neurosci., 4, 8
Giger, R. J., Hollis, E. R. 2nd and Tuszynski, M. H. (2010) Guidance molecules in axon regeneration. Cold Spring Harb. Perspect. Biol., 2, a001867
Zuo, J., Neubauer, D., Dyess, K., Ferguson, T. A. and Muir, D. (1998) Degradation of chondroitin sulfate proteoglycan enhances the neurite-promoting potential of spinal cord tissue. Exp. Neurol., 154, 654–662
Chen, M. S., Huber, A. B., van der Haar, M. E., Frank, M., Schnell, L., Spillmann, A. A., Christ, F. and Schwab, M. E. (2000) Nogo-A is a myelin-associated neurite outgrowth inhibitor and an antigen for monoclonal antibody IN-1. Nature, 403, 434–439
Rubin, B. P., Dusart, I. and Schwab, M. E. (1994) A monoclonal antibody (IN-1) which neutralizes neurite growth inhibitory proteins in the rat CNS recognizes antigens localized in CNS myelin. J. Neurocytol., 23, 209–217
Michaelevski, I., Segal-Ruder, Y., Rozenbaum, M., Medzihradszky, K. F., Shalem, O., Coppola, G., Horn-Saban, S., Ben-Yaakov, K., Dagan, S. Y., Rishal, I., et al. (2010) Signaling to transcription networks in the neuronal retrograde injury response. Sci. Signal., 3, ra53
Nix, P., Hisamoto, N., Matsumoto, K. and Bastiani, M. (2011) Axon regeneration requires coordinate activation of p38 and JNK MAPK pathways. Proc. Natl. Acad. Sci. USA, 108, 10738–10743
Yiu, G. and He, Z. (2006) Glial inhibition of CNS axon regeneration. Nat. Rev. Neurosci., 7, 617–627
Horner, P. J. and Gage, F. H. (2000) Regenerating the damaged central nervous system. Nature, 407, 963–970
Mattson, M. P. (1989) Acetylcholine potentiates glutamateinduced neurodegeneration in cultured hippocampal neurons. Brain Res., 497, 402–406
Connor, B. and Dragunow, M. (1998) The role of neuronal growth factors in neurodegenerative disorders of the human brain. Brain Res. Rev., 27, 1–39
Kamei, N., Tanaka, N., Oishi, Y., Hamasaki, T., Nakanishi, K., Sakai, N. and Ochi, M. (2007) BDNF, NT-3, and NGF released from transplanted neural progenitor cells promote corticospinal axon growth in organotypic cocultures. Spine, 32, 1272–1278
Ziegner, U. H., Kobayashi, R. H., Cunningham-Rundles, C., Español, T., Fasth, A., Huttenlocher, A., Krogstad, P., Marthinsen, L., Notarangelo, L. D., Pasic, S., et al. (2002) Progressive neurodegeneration in patients with primary immunodeficiency disease on IVIG treatment. Clin. Immunol., 102, 19–24
Blesch, A., Lu, P. and Tuszynski, M. H. (2002) Neurotrophic factors, gene therapy, and neural stem cells for spinal cord repair. Brain Res. Bull., 57, 833–838
Huang, D. W., McKerracher, L., Braun, P. E. and David, S. (1999) A therapeutic vaccine approach to stimulate axon regeneration in the adult mammalian spinal cord. Neuron, 24, 639–647
Sicotte, M., Tsatas, O., Jeong, S. Y., Cai, C.-Q., He, Z. and David, S. (2003) Immunization with myelin or recombinant Nogo-66/MAG in alum promotes axon regeneration and sprouting after corticospinal tract lesions in the spinal cord. Mol. Cell. Neurosci., 23, 251–263
Chandran, V., Coppola, G., Nawabi, H., Omura, T., Versano, R., Huebner, E. A., Zhang, A., Costigan, M., Yekkirala, A., Barrett, L., et al. (2016) A systems-level analysis of the peripheral nerve intrinsic axonal growth program. Neuron, 89, 956–970
Dutkowski, J. and Ideker, T. (2011) Protein networks as logic functions in development and cancer. PLoS Comput. Biol., 7, e1002180
Dong, X., Jiang, Z., Peng, Y. L. and Zhang, Z. (2015) Revealing shared and distinct gene network organization in Arabidopsis immune responses by integrative analysis. Plant Physiol., 167, 1186–1203
Leiserson, M. D. M., Vandin, F., Wu, H. T., Dobson, J. R., Eldridge, J. V., Thomas, J. L., Papoutsaki, A., Kim, Y., Niu, B., McLellan, M., et al. (2015) Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat. Genet., 47, 106–114
Lamb, J., Crawford, E. D., Peck, D., Modell, J. W., Blat, I. C., Wrobel, M. J., Lerner, J., Brunet, J. P., Subramanian, A., Ross, K. N., et al. (2006) The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science, 313, 1929–1935
Johnson, W. E., Li, C. and Rabinovic, A. (2007) Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics, 8, 118–127
Kim, J., So, S., Lee, H. J., Park, J. C., Kim, J. J. and Lee, H. (2013) DigSee: disease gene search engine with evidence sentences (version cancer). Nucleic Acids Res., 41, W510–W517
Fleming, C. E., Saraiva, M. J. and Sousa, M. M. (2007) Transthyretin enhances nerve regeneration. J. Neurochem., 103, 831–839
Egan, M. F., Goldberg, T. E., Kolachana, B. S., Callicott, J. H., Mazzanti, C. M., Straub, R. E., Goldman, D. and Weinberger, D. R. (2001) Effect of COMT Val108/158Met genotype on frontal lobe function and risk for schizophrenia. Proc. Natl. Acad. Sci. USA, 98, 6917–6922
Chen, W., Chen, C., Xia, M., Wu, K., Chen, C., He, Q., Xue, G., Wang,W., He, Y. and Dong, Q. (2016) Interaction effects of BDNF and COMT genes on resting-state brain activity and working memory. Front. Hum. Neurosci., 10, 540
Lewin, S. L., Utley, D. S., Cheng, E. T., Verity, A. N. and Terris, D. J. (1997) Simultaneous treatment with BDNF and CNTF after peripheral nerve transection and repair enhances rate of functional recovery compared with BDNF treatment alone. Laryngoscope, 107, 992–999
Kuleshov, M. V., Jones, M. R., Rouillard, A. D., Fernandez, N. F., Duan, Q., Wang, Z., Koplev, S., Jenkins, S. L., Jagodnik, K. M., Lachmann, A., et al. (2016) Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res., 44, W90–W97
Namm, A., Arend, A. and Aunapuu, M. (2013) Pax proteins in embryogenesis and their role in nervous system development. Pap. Anthropol., 22, 133–142
Burrill, J. D., Moran, L., Goulding, M. D. and Saueressig, H. (1997) PAX2 is expressed in multiple spinal cord interneurons, including a population of EN1+ interneurons that require PAX6 for their development. Development, 124, 4493–4503
Ziman, M. R., Rodger, J., Chen, P., Papadimitriou, J. M., Dunlop, S. A. and Beazley, L. D. (2001) Pax genes in development and maturation of the vertebrate visual system: implications for optic nerve regeneration. Histol. Histopathol., 16, 239–249
Raivich, G., Bohatschek, M., Da Costa, C., Iwata, O., Galiano, M., Hristova, M., Nateri, A. S., Makwana, M., Riera-Sans, L., Wolfer, D. P., et al. (2004) The AP-1 transcription factor c-Jun is required for efficient axonal regeneration. Neuron, 43, 57–67
Iorio, F., Bosotti, R., Scacheri, E., Belcastro, V., Mithbaokar, P., Ferriero, R., Murino, L., Tagliaferri, R., Brunetti-Pierri, N., Isacchi, A., et al. (2010) Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc. Natl. Acad. Sci. USA, 107, 14621–14626
Sakaue, Y., Sanada, M., Sasaki, T., Kashiwagi, A. and Yasuda, H. (2003) Amelioration of retarded neurite outgrowth of dorsal root ganglion neurons by overexpression of PKCd in diabetic rats. Neuroreport, 14, 431–436
Duan, Q., Reid, S. P., Clark, N. R., Wang, Z., Fernandez, N. F., Rouillard, A. D., Readhead, B., Tritsch, S. R., Hodos, R., Hafner, M., et al. (2016) L1000CDS2: LINCS L1000 characteristic direction signatures search engine. NPJ Syst. Biol. Appl., 2, 16015
Sun, F., Park, K. K., Belin, S., Wang, D., Lu, T., Chen, G., Zhang, K., Yeung, C., Feng, G., Yankner, B. A., et al. (2011) Sustained axon regeneration induced by co-deletion of PTEN and SOCS3. Nature, 480, 372–375
Agudelo, M., Gandhi, N., Saiyed, Z., Pichili, V., Thangavel, S., Khatavkar, P., Yndart-Arias, A. and Nair, M. (2011) Effects of alcohol on histone deacetylase 2 (HDAC2) and the neuroprotective role of trichostatin A (TSA). Alcohol. Clin. Exp. Res., 35, 1550–1556
Bolstad, B. M., Collin, F., Simpson, K. M., Irizarry, R. A. and Speed, T. P. (2004) Experimental design and low-level analysis of microarray data. Int. Rev. Neurobiol., 60, 25–58
Szklarczyk, D., Morris, J. H., Cook, H., Kuhn, M., Wyder, S., Simonovic, M., Santos, A., Doncheva, N. T., Roth, A., Bork, P., et al. (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res., 45, D362–D368
Breiman, L. I., Friedman, J. H., Olshen, R. A. and Stone, C. J. (1984) Classification and Regression Trees (CART). 1 Ed., Chapman and Hall/CRC
Chung, F. (2007) The heat kernel as the pagerank of a graph. Proc. Natl. Acad. Sci. USA, 104, 19735–19740
Vandin, F., Upfal, E. and Raphael, B. J. (2011) Algorithms for detecting significantly mutated pathways in cancer. J. Comput. Biol., 18, 507–522
Vandin, F., Clay, P., Upfal, E. and Raphael, B. J. (2012) Discovery of mutated subnetworks associated with clinical data in cancer. In Proceedings of the Pacific Symposium of Biocomputing 2012, pp. 55–66. World Scientific
Huang, W., Sherman, B. T. and Lempicki, R. A. (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res., 37, 1–13
Huang, W., Sherman, B. T. and Lempicki, R. A. (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc., 4, 44–57
Mi, H., Huang, X., Muruganujan, A., Tang, H., Mills, C., Kang, D. and Thomas, P. D. (2017) PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res., 45, D183–D189
Templeton, J. P. and Geisert, E. E. (2012) A practical approach to optic nerve crush in the mouse. Mol. Vis., 18, 2147–2152
Acknowledgements
This work was supported by the Fundamental Research Funds for the Central Universities (No. 2662017PY115)
Author information
Authors and Affiliations
Corresponding authors
Additional information
Author summary: Compared with peripheral nervous system, the regenerative capacity of the central nervous system is extremely limited. Thus, it may lead to the discovery of treatment for central nerve injury by modulating gene networks underlying peripheral nerve injury. Based on the gene expression profiles of dorsal root ganglion after a sciatic nerve injury, we defined gene networks underlying peripheral nerve injury. After screening compounds that were likely to modulate gene expressions of the networks, trichostatin A, a histone deacetylase inhibitor, was validated to enhance neurite outgrowth in vivo via an optic nerve crush mouse model.
Rights and permissions
About this article
Cite this article
Wang, H., Wang, G., Zhu, LD. et al. Subnetwork identification and chemical modulation for neural regeneration: A study combining network guided forest and heat diffusion model. Quant Biol 6, 321–333 (2018). https://doi.org/10.1007/s40484-018-0159-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-018-0159-0