Abstract
Background
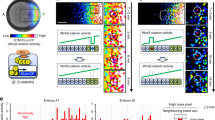
Developmental patterning is highly reproducible and accurate at the single-cell level during fly embryogenesis despite the gene expression noise and external perturbations such as the variation of the embryo length, temperature and genes. To reveal the underlying mechanism, it is very important to characterize the noise transmission during the dynamic pattern formation. Two hypotheses have been proposed. The “channel” scenario requires a highly reproducible input and an accurate interpretation by downstream genes. In contrast, the “filter” scenario proposes a noisy input and a noise filter via the cross-regulation of the downstream network. It has been under great debates which scenario the fly embryogenesis follows.
Results
The first 3-h developmental patterning of fly embryos is orchestrated by a hierarchical segmentation gene network, which rewires upon the maternal to zygotic transition. Starting from the highly reproducible maternal gradients, the positional information is refined to the single-cell precision through the highly dynamical evolved zygotic gene expression profiles. Thus the fly embryo development might strictly fit into neither the originally proposed “filter” nor “channel” scenario. The controversy that which scenario the fly embryogenesis follows could be further clarified by combining quantitative measurements and modeling.
Conclusions
Fly embryos have become one of the perfect model systems for quantitative systems biology studies. The underlying mechanism discovered from fly embryogenesis will deepen our understanding of the noise control of the gene network, facilitate searching for more efficient and safer methods for cell programming and reprogramming, and have the great potential for tissue engineering and regenerative medicine.

Article PDF
Similar content being viewed by others
References
Gregor, T., Tank, D. W., Wieschaus, E. F. and Bialek, W. (2007) Probing the limits to positional information. Cell, 130, 153–164
Gregor, T., Wieschaus, E. F., McGregor, A. P., Bialek, W. and Tank, D. W. (2007) Stability and nuclear dynamics of the bicoid morphogen gradient. Cell, 130, 141–152
Namba, R., Pazdera, T. M., Cerrone, R. L. and Minden, J. S. (1997) Drosophila embryonic pattern repair: how embryos respond to bicoid dosage alteration. Development, 124, 1393–1403
Vincent, A., Blankenship, J. T. and Wieschaus, E. (1997) Integration of the head and trunk segmentation systems controls cephalic furrow formation in Drosophila. Development, 124, 3747–3754
Liu, F., Morrison, A. H. and Gregor, T. (2013) Dynamic interpretation of maternal inputs by the Drosophila segmentation gene network. Proc. Natl. Acad. Sci. USA, 110, 6724–6729
Eldar, A. and Elowitz, M. B. (2010) Functional roles for noise in genetic circuits. Nature, 467, 167–173
Elowitz, M. B., Levine, A. J., Siggia, E. D. and Swain, P. S. (2002) Stochastic gene expression in a single cell. Science, 297, 1183–1186
Ghodsi, Z., Hassani, H. and McGhee, K. (2015) Mathematical approaches in studying bicoid gene. Quant. Biol., 3, 182–192.
Gregor, T., Garcia, H. G. and Little, S. C. (2014) The embryo as a laboratory: quantifying transcription in Drosophila. Trends Genet., 30, 364–375
Rogers, K.W. and Schier, A. F. (2011) Morphogen gradients: from generation to interpretation. Annu. Rev. Cell Dev. Biol., 27, 377–407
Porcher, A. and Dostatni, N. (2010) The bicoid morphogen system. Curr. Biol., 20, R249–R254
Jaeger, J. (2011) The gap gene network. Cell. Mol. Life Sci., 68, 243–274
Jaeger, J. (2009) Modelling the Drosophila embryo. Mol. Biosyst., 5, 1549–1568
Farrell, J. A. and O’Farrell, P. H. (2014) From egg to gastrula: how the cell cycle is remodeled during the Drosophila mid-blastula transition. Annu. Rev. Genet., 48, 269–294
Driever, W. and Nüsslein-Volhard, C. (1988) A gradient of bicoid protein in Drosophila embryos. Cell, 54, 83–93
Driever, W. and Nüsslein-Volhard, C. (1988) The bicoid protein determines position in the Drosophila embryo in a concentrationdependent manner. Cell, 54, 95–104
Irish, V., Lehmann, R. and Akam, M. (1989) The Drosophila posterior-group gene nanos functions by repressing hunchback activity. Nature, 338, 646–648
Dubuis, J. O., Samanta, R. and Gregor, T. (2013) Accurate measurements of dynamics and reproducibility in small genetic networks. Mol. Syst. Biol., 9, 639
Surkova, S., Myasnikova, E., Kozlov, K. N., Pisarev, A., Reinitz, J. and Samsonova, M. (2013) Quantitative imaging of gene expression in Drosophila embryos. Cold Spring Harb. Protoc., 2013, 488–497
Little, S. C., Tkacik, G., Kneeland, T. B., Wieschaus, E. F. and Gregor, T. (2011) The formation of the bicoid morphogen gradient requires protein movement from anteriorly localized mRNA. PLoS Biol., 9, e1000596
Grimm, O., Coppey, M. and Wieschaus, E. (2010) Modelling the bicoid gradient. Development, 137, 2253–2264
Berezhkovskii, A. M., Sample, C. and Shvartsman, S. Y. (2010) How long does it take to establish a morphogen gradient? Biophys. J., 99, L59–L61
Drocco, J. A., Grimm, O., Tank, D. W. and Wieschaus, E. (2011) Measurement and perturbation of morphogen lifetime: effects on gradient shape. Biophys. J., 101, 1807–1815
Liu, J., He, F. and Ma, J. (2011) Morphogen gradient formation and action: insights from studying bicoid protein degradation. Fly (Austin), 5, 242–246
Abu-Arish, A., Porcher, A., Czerwonka, A., Dostatni, N. and Fradin, C. (2010) High mobility of bicoid captured by fluorescence correlation spectroscopy: implication for the rapid establishment of its gradient. Biophys. J., 99, L33–L35
Liu, J. and Ma, J. (2013) Uncovering a dynamic feature of the transcriptional regulatory network for anterior–posterior patterning in the Drosophila embryo. PLoS One, 8, e62641
Margolis, J. S., Borowsky, M. L., Steingrímsson, E., Shim, C. W., Lengyel, J. A. and Posakony, J. W. (1995) Posterior stripe expression of hunchback is driven from two promoters by a common enhancer element. Development, 121, 3067–3077
Lopes, F. J., Vieira, F. M., Holloway, D. M., Bisch, P. M. and Spirov, A. V. (2008) Spatial bistability generates hunchback expression sharpness in the Drosophila embryo. PLoS Comput. Biol., 4, e1000184
Hülskamp, M., Lukowitz, W., Beermann, A., Glaser, G. and Tautz, D. (1994) Differential regulation of target genes by different alleles of the segmentation gene hunchback in Drosophila. Genetics, 138, 125–134
Schröder, C., Tautz, D., Seifert, E. and Jäckle, H. (1988) Differential regulation of the two transcripts from the Drosophila gap segmentation gene hunchback. EMBO J., 7, 2881–2887
Perry, M. W., Bothma, J. P., Luu, R. D. and Levine, M. (2012) Precision of hunchback expression in the Drosophila embryo. Curr. Biol., 22, 2247–2252
Surkova, S., Golubkova, E., Manu, L., Panok, L., Mamon, J., Reinitz, M. and Samsonova (2013) Quantitative dynamics and increased variability of segmentation gene expression in the Drosophila Krüppel and knirps mutants. Dev. Biol., 376, 99–112
Perry, M. W., Boettiger, A. N. and Levine, M. (2011) Multiple enhancers ensure precision of gap gene-expression patterns in the Drosophila embryo. Proc. Natl. Acad. Sci. USA, 108, 13570–13575
Lehmann, R. and Nüsslein-Volhard, C. (1987) hunchback, a gene required for segmentation of an anterior and posterior region of the Drosophila embryo. Dev. Biol., 119, 402–417
Petkova, M. D., Tkacik, G., Bialek, W., Wieschaus, E. F. and Gregor, T. (2016) Optimal decoding of information from a genetic network. arXiv:161208084
Petkova, M. D., Little, S. C., Liu, F. and Gregor, T. (2014) Maternal origins of developmental reproducibility. Curr. Biol., 24, 1283–1288
Wolpert, L. (1969) Positional information and the spatial pattern of cellular differentiation. J. Theor. Biol., 25, 1–47
He, F., Wen, Y., Deng, J., Lin, X., Lu, L. J., Jiao, R. and Ma, J. (2008) Probing intrinsic properties of a robust morphogen gradient in Drosophila. Dev. Cell, 15, 558–567
Houchmandzadeh, B., Wieschaus, E. and Leibler, S. (2002) Establishment of developmental precision and proportions in the early Drosophila embryo. Nature, 415, 798–802
Jaeger, J., Surkova, S., Blagov, M., Janssens, H., Kosman, D., Kozlov, K. N., Manu, E., Myasnikova, C. E., Vanario-Alonso, M., Samsonova, M, et al. (2004) Dynamic control of positional information in the early Drosophila embryo. Nature, 430, 368–371
Manu, Surkova, S., Spirov, A. V., Gursky, V. V., Janssens, H., Kim, A. R., Radulescu, O., Vanario-Alonso, C. E., Sharp, D. H., Samsonova, M., et al. (2009) Canalization of gene expression in the Drosophila blastoderm by gap gene cross regulation. PLoS Biol., 7, e1000049
Green, J. B. and Sharpe, J. (2015) Positional information and reaction-diffusion: two big ideas in developmental biology combine. Development, 142, 1203–1211
Turing, A. M. (1952) The chemical basis of morphogenesis. Philosoph. Trans. Royal Soc. London, 237, 37–72
Gregor, T., Bialek, W., de Ruyter van Steveninck, R. R., Tank, D. W. and Wieschaus, E. F. (2005) Diffusion and scaling during early embryonic pattern formation. Proc. Natl. Acad. Sci. USA, 102, 18403–18407
Grimm, O. and Wieschaus, E. (2010) The bicoid gradient is shaped independently of nuclei. Development, 137, 2857–2862
Cheung, D., Miles, C., Kreitman, M. and Ma, J. (2011) Scaling of the bicoid morphogen gradient by a volume-dependent production rate. Development, 138, 2741–2749
He, F., Wei, C., Wu, H., Cheung, D., Jiao, R. and Ma, J. (2015) Fundamental origins and limits for scaling a maternal morphogen gradient. Nat. Commun., 6, 6679
de Lachapelle, A. M. and Bergmann, S. (2010) Precision and scaling in morphogen gradient read-out. Mol. Syst. Biol., 6, 351
Bergmann, S., Sandler, O., Sberro, H., Shnider, S., Schejter, E., Shilo, B.-Z. and Barkai, N. (2007) Pre-steady-state decoding of the bicoid morphogen gradient. PLoS Biol., 5, e46
Jaeger, J. (2010) A matter of timing and precision. Mol. Syst. Biol., 6, 427
Houchmandzadeh, B., Wieschaus, E. and Leibler, S. (2005) Precise domain specification in the developing Drosophila embryo. Phys. Rev. E Stat. Nonlin. Soft Matter Phys., 72, 061920
Howard, M. and ten Wolde, P. R. (2005) Finding the center reliably: robust patterns of developmental gene expression. Phys. Rev. Lett., 95, 208103
Cheung, D. and Ma, J. (2015) Probing the impact of temperature on molecular events in a developmental system. Sci. Rep., 5, 13124
Kuntz, S. G. and Eisen, M. B. (2014) Drosophila embryogenesis scales uniformly across temperature in developmentally diverse species. PLoS Genet., 10, e1004293
Lucchetta, E. M., Lee, J. H., Fu, L. A., Patel, N. H. and Ismagilov, R. F. (2005) Dynamics of Drosophila embryonic patterning network perturbed in space and time using microfluidics. Nature, 434, 1134–1138
Lucchetta, E. M., Vincent, M. E. and Ismagilov, R. F. (2008) A precise bicoid gradient is nonessential during cycles 11–13 for precise patterning in the Drosophila blastoderm. PLoS One, 3, e3651
Lucchetta, E. M., Carthew, R. W. and Ismagilov, R. F. (2009) The endo-siRNA pathway is essential for robust development of the Drosophila embryo. PLoS One, 4, e7576
Crauk, O. and Dostatni, N. (2005) bicoid determines sharp and precise target gene expression in the Drosophila embryo. Curr. Biol., 15, 1888–1898
Kugler, J.-M. and Lasko, P. (2009) Localization, anchoring and translational control of oskar, gurken, bicoid and nanos mRNA during Drosophila oogenesis. Fly (Austin), 3, 15–28
Aegerter-Wilmsen, T., Aegerter, C. M. and Bisseling, T. (2005) Model for the robust establishment of precise proportions in the early Drosophila embryo. J. Theor. Biol., 234, 13–19
Garcia, H. G., Tikhonov, M., Lin, A. and Gregor, T. (2013) Quantitative imaging of transcription in living Drosophila embryos links polymerase activity to patterning. Curr. Biol., 23, 2140–2145
Lucas, T., Ferraro, T., Roelens, B., De Las Heras Chanes, J., Walczak, A. M., Coppey, M. and Dostatni, N. (2013) Live imaging of bicoid-dependent transcription in Drosophila embryos. Curr. Biol., 23, 2135–2139
Xu, H., Sepúlveda, L. A., Figard, L., Sokac, A. M. and Golding, I. (2015) Combining protein and mRNA quantification to decipher transcriptional regulation. Nat. Methods, 12, 739–742
Little, S. C., Tikhonov, M. and Gregor, T. (2013) Precise developmental gene expression arises from globally stochastic transcriptional activity. Cell, 154, 789–800
Garcia, H. G., Tikhonov, M., Lin, A. and Gregor, T. (2013) Quantitative imaging of transcription in living Drosophila embryos links polymerase activity to patterning. Curr. Biol., 23, 2140–2145
Golding, I., Paulsson, J., Zawilski, S. M. and Cox, E. C. (2005) Real-time kinetics of gene activity in individual bacteria. Cell, 123, 1025–1036
Reeves, G. T., Trisnadi, N., Truong, T. V., Nahmad, M., Katz, S. and Stathopoulos, A. (2012) Dorsal-ventral gene expression in the Drosophila embryo reflects the dynamics and precision of the dorsal nuclear gradient. Dev. Cell, 22, 544–557
Giepmans, B. N., Adams, S. R., Ellisman, M. H., Tsien, R. Y (2006) The fluorescent toolbox for assessing protein location and function. Science, 312, 217–224
Myasnikova, E., Samsonova, A., Kozlov, K., Samsonova, M. and Reinitz, J. (2001) Registration of the expression patterns of Drosophila segmentation genes by two independent methods. Bioinformatics, 17, 3–12
Blythe, S. A. and Wieschaus, E. F. (2016) Establishment and maintenance of heritable chromatin structure during early Drosophila embryogenesis. eLife, 5, 5
Fowlkes, C. C., Hendriks, C. L. L., Keränen, S. V., Weber, G. H., Rübel, O., Huang, M.-Y., Chatoor, S., DePace, A. H., Simirenko, L., Henriquez, C., et al. (2008) A quantitative spatiotemporal atlas of gene expression in the Drosophila blastoderm. Cell, 133, 364–374
Surkova, S., Kosman, D., Kozlov, K., Manu, E., Myasnikova, A. A., Samsonova, A., Spirov, C. E., Vanario-Alonso, M., Samsonova, J. and Reinitz (2008) Characterization of the Drosophila segment determination morphome. Dev. Biol., 313, 844–862
Barrangou, R. (2014) RNA events. Cas9 targeting and the CRISPR revolution. Science, 344, 707–708
Bassett, A. R. and Liu, J.-L. (2014) CRISPR/Cas9 and genome editing in Drosophila. J. Genet. Genomics, 41, 7–19
Papatsenko, D. and Levine, M. (2011) The Drosophila gap gene network is composed of two parallel toggle switches. PLoS One, 6, e21145
Bertrand, E., Chartrand, P., Schaefer, M., Shenoy, S. M., Singer, R. H. and Long, R. M. (1998) Localization of ASH1 mRNA particles in living yeast. Mol. Cell, 2, 437–445
Bothma, J. P., Garcia, H. G., Esposito, E., Schlissel, G., Gregor, T. and Levine, M. (2014) Dynamic regulation of eve stripe 2 expression reveals transcriptional bursts in living Drosophila embryos. Proc. Natl. Acad. Sci. USA, 111, 10598–10603
Keller, P. J. (2013) Imaging morphogenesis: technological advances and biological insights. Science, 340, 1234168
Stegmaier, J., Amat, F., Lemon, W. C., McDole, K., Wan, Y., Teodoro, G., Mikut, R. and Keller, P. J. (2016) Real-time threedimensional cell segmentation in large-scale microscopy data of developing embryos. Dev. Cell, 36, 225–240
Ji, N., Milkie, D. E. and Betzig, E. (2010) Adaptive optics via pupil segmentation for high-resolution imaging in biological tissues. Nat. Methods, 7, 141–147
Huang, A., Amourda, C., Zhang, S., Tolwinski, N. S., and Saunders, T. E. (2017) Decoding temporal interpretation of the morphogen bicoid in the early Drosophila embryo. eLife, 6, e26258
Perry, M.W., Boettiger, A. N., Bothma, J. P. and Levine, M. (2010) Shadow enhancers foster robustness of Drosophila gastrulation. Curr. Biol., 20, 1562–1567
El-Sherif, E. and Levine, M. (2016) Shadow enhancers mediate dynamic shifts of gap gene expression in the Drosophila embryo. Curr. Biol., 26, 1164–1169
Lagha, M., Bothma, J. P., Esposito, E., Ng, S., Stefanik, L., Tsui, C., Johnston, J., Chen, K., Gilmour, D. S., Zeitlinger, J., et al. (2013) Paused Pol II coordinates tissue morphogenesis in the Drosophila embryo. Cell, 153, 976–987
Liu, J. and Ma, J. (2011) Fates-shifted is an F-box protein that targets bicoid for degradation and regulates developmental fate determination in Drosophila embryos. Nat. Cell Biol., 13, 22–29
Liu, J., Xiao, Y., Zhang, T. and Ma, J. (2016) Time to move on: modeling transcription dynamics during an embryonic transition away from maternal control. Fly (Austin), 10, 101–107
Liu, J. and Ma, J. (2015) Modulation of temporal dynamics of gene transcription by activator potency in the Drosophila embryo. Development, 142, 3781–3790
Phillips, R. (2015) Theory in biology: Figure 1 or Figure 7? Trends Cell Biol., 25, 723–729
Estrada, J., Wong, F., DePace, A. and Gunawardena, J. (2016) Information integration and energy expenditure in gene regulation. Cell, 166, 234–244
Acknowledgments
The authors are grateful for the two reviewers for helpful suggestions. We apologize to our colleagues whose work could not be cited due to the page limits. This project is supported by the National Natural Science Foundation of China (No.31670852) and 100-talent plan of Peking University. The Drosophila lab used in this project is supported by Peking-Tsinghua Center for Life Sciences.
Author information
Authors and Affiliations
Corresponding author
Additional information
Author summary: It is intriguing how the development of organisms is highly reproducible and accurate despite the external and internal noise. The key to solve this mystery is to explore the noise transmission during development and discover the underlying mechanism. The developmental system could generate precise inputs and outputs in each step, or gradually generate a final precise output by filtering a noisy input according to the “channel” or “filter” scenario, respectively. The quantitative studies on the early development of the fruit fly’s embryos show that the developmental process is highly dynamic and its noise transmission may employ a hybridized strategy.
Rights and permissions
About this article
Cite this article
Yang, Z., Wu, X., Yang, N. et al. Noise transmission during the dynamic pattern formation in fly embryos. Quant Biol 6, 15–29 (2018). https://doi.org/10.1007/s40484-018-0135-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-018-0135-8