Abstract
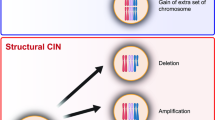
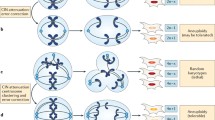
Genetic instability is a defining property of cancer cells and is the basis of various lesions including point mutations, copy number alterations and translocations. Chromosomal instability (CIN) is part of the genetic instability of cancer and consists of copy number alterations in whole or parts of cancer cell chromosomes. CIN is observed in differing degrees in most cancers. In breast cancer, CIN is commonly part of the genomic landscape of the disease and has a higher incidence in aggressive sub-types. Tumor suppressors that are commonly mutated or disabled in cancer, such as p53 and pRB, play roles in protection against CIN, and as a result, their dysfunction contributes to the establishment or tolerance of CIN. Several structural and regulatory proteins of the centromeres and kinetochore, the complex structure that is responsible for the correct distribution of genetic material in the daughter cells during mitosis, are direct or, mostly, indirect transcription targets of p53 and pRB. Thus, despite the absence of structural defects in genes encoding for centromere and kinetochore components, dysfunction of these tumor suppressors may have profound implications for the correct function of the mitotic apparatus contributing to CIN. CIN and its prognostic and therapeutic implications in breast cancer are discussed in this article.

Similar content being viewed by others
References
McGranahan N, Burrell RA, Endesfelder D, Novelli MR, Swanton C. Cancer chromosomal instability: therapeutic and diagnostic challenges. EMBO Rep. 2012;13:528–38.
Musacchio A, Desai A. A molecular view of kinetochore assembly and function. Biol. 2017;6:5.
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, et al. DNA breaks and chromosome pulverization from errors in mitosis. Nature. 2012;482:53–8.
Bakhoum SF, Ngo B, Laughney AM, Cavallo J-A, Murphy CJ, Ly P, et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature. 2018;553:467–72.
Bakhoum F, Silkworth WT, Nardi IK, Nicholson JM, Compton DA, Cimini D. The mitotic origin of chromosomal instability. Curr Biol. 2014;24:R148–9.
Duijf PHG, Nanayakkara D, Nones K, Srihari S, Kalimutho M, Khanna KK. Mechanisms of genomic instability in breast cancer. Trends Mol Med. 2019; S1471-4914(19)30090-5 [Epub ahead of print].
Danielsen HE, Pradhan M, Novelli M. Revisiting tumour aneuploidy—the place of ploidy assessment in the molecular era. Nat Rev Clin Oncol. 2016;13:291–304.
Pinto AE, André S, Pereira T, Silva G, Soares J. DNA flow cytometry but not telomerase activity as predictor of disease-free survival in pT1-2/N0/G2 breast cancer. Pathobiol. 2006;73:63–70.
Moureau-Zabotto L, Bouchet C, Cesari D, Uzan S, Lefranc J-P, Antoine M, et al. Combined flow cytometry determination of S-phase fraction and DNA ploidy is an independent prognostic factor in node-negative invasive breast carcinoma: analysis of a series of 271 patients with stage I and II breast cancer. Breast Cancer Res Treat. 2005;91:61–71.
Li L, Mu K, Zhou G, Lan L, Auer G, Zetterberg A. Genomic instability and proliferative activity as risk factors for distant metastases in breast cancer. Br J Cancer. 2008;99:513–9.
Habermann JK, Doering J, Hautaniemi S, Roblick UJ, Bündgen NK, Nicorici D, et al. The gene expression signature of genomic instability in breast cancer is an independent predictor of clinical outcome. Int J Cancer. 2009;124:1552–64.
Pinto AE, Pereira T, Santos M, Branco M, Dias A, Silva GL, et al. DNA ploidy is an independent predictor of survival in breast invasive ductal carcinoma: a long-term multivariate analysis of 393 patients. Ann Surg Oncol. 2013;20:1530–7.
Carter SL, Eklund AC, Kohane IS, Harris LN, Szallasi Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet. 2006;38:1043–8.
Szász AM, Li Q, Eklund AC, Sztupinszki Z, Rowan A, Tökés A-M, et al. The CIN4 chromosomal instability qPCR classifier defines tumor aneuploidy and stratifies outcome in grade 2 breast cancer. PLoS One. 2013;8:e56707.
Spears M, Yousif F, Lyttle N, Boutros PC, Munro AF, Twelves C, et al. A four gene signature predicts benefit from anthracyclines: evidence from the BR9601 and MA.5 clinical trials. Oncotarget. 2015;6:31693–31701.
Spears M, Lyttle N, D’Costa A, Chen BE, Yao CQ, Boutros PC, et al. A four gene signature of chromosome instability (CIN4) predicts for benefit from taxanes in the NCIC-CTG MA21 clinical trial. Oncotarget. 2016;7:49099–106.
Zhang W, Mao J-H, Zhu W, Jain AK, Liu K, Brown JB, Karpen GH. Centromere and kinetochore gene misexpression predicts cancer patient survival and response to radiotherapy and chemotherapy. Nat Commun. 2016;7:12619.
Bolhaqueiro ACF, Ponsioen B, Bakker B, Klaasen SJ, Kucukkose E, van Jaarsveld RH, et al. Ongoing chromosomal instability and karyotype evolution in human colorectal cancer organoids. Nat Genet. 2019;51:824–34.
Johnson SD, McClelland SE. Watching cancer cells evolve. Nature. 2019;570:166–7.
Smid M, Hoes M, Sieuwerts AM, Sleijfer S, Zhang Y, Wang Y, et al. Patterns and incidence of chromosomal instability and their prognostic relevance in breast cancer subtypes. Breast Cancer Res Treat. 2011;128:23–30.
A’Hern RP, Jamal-Hanjani M, Szász AM, Johnston SRD, Reis-Filho JS, Roylance R, Swanton C. Taxanes benefit in breast cancer- a role for grade and chromosome stability. Nat Rev Clin Oncol. 2013;10:357–64.
Martinez-Arribas F, Nuñez-Villar MJ, Lucas AR, Sanchez J, Tejerina A, Schneider J. The S-phase fraction of the aneuploidy cell subpopulation is the biologically relevant one in aneuploidy breast cancers. Breast Cancer Res Treat. 2005;92:77–80.
Persons DL, Robinson RA, Hsu PH, Seelig SA, Borell TJ, Hartmann LC, Jenkins RB. Chromosome-specific aneusomy in carcinoma of the breast. Clin Cancer Res. 1996;2:883–8.
Thiru P, Kern DM, McKinley KL, Monda JK, Rago F, Su K-C, et al. Kinetochore genes are coordinately up-regulated in human tumors as part of a FoxM1-related cell division program. Mol Biol Cell. 2014;25:1983–94.
Pfister K, Pipka JL, Chiang C, Liu Y, Clark RA, et al. Identification of drivers of aneuploidy in breast tumors. Cell Rep. 2018;23:2758–69.
Nik-Zainal S, Morganella S. Mutational signatures in breast: the problem at the DNA level. Clin Cancer Res. 2017;23:2617–29.
Roylance R, Endesfelder D, Gorman P, Burrell RA, Sander J, Tomlinson I, et al. Relationship of extreme chromosomal instability with long-term survival in a retrospective analysis of primary breast cancer. Cancer Epidemiol Biomark Prev. 2011;20:2183–94.
Jamal-Hanjani M, A’Hern R, Birkbak NJ, et al. Extreme chromosomal instability forecasts improved outcome in ER-negative breast cancer: a prospective validation cohort study from the TACT trial. Ann Oncol. 2015;26:1340–6.
Nath S, Chowdhury A, Dey S, Roychoudhury A, Ganguly A, Bhattacharyya D, Roychoudhury S. Deregulation of Rb-E2F1 axis causes chromosomal instability by engaging the transactivation function of Cdc20-anaphase-promoting complex/cyclosome. Mol Cell Biol. 2015;35:356–69.
Amato A, Schillaci T, Lentini L, Di Leonardo A. CENPA overexpression promotes genome instability in pRb-depleted human cells. Mol Cancer. 2009;8:119.
Fridlyand J, Snijders AM, Ylstra B, Li H, Olshen A, Segraves R, et al. Breast tumor copy number aberration phenotypes and genomic instability. BMC Cancer. 2006;6:96.
Schvartzman J-M, Duijf PHG, Sotillo R, Coker C, Benezra R. Mad2 is a critical mediator of the chromosome instability observed upon Rb and p53 pathway inhibition. Cancer Cell. 2011;19:701–14.
Berenzeno IM, Piñeiro R, Castillo SD, Pearce W, McGranahan N, Dewhurst SM, et al. Oncogenic PI3CA induces centrosome amplification and tolerance to genome doubling. Nat Commun. 2017;8:1773.
Crockford A, Zalmas LP, Grönroos E, et al. Cyclin D mediates tolerance of genome-doubling in cancers with functional p53. Ann Oncol. 2017;28:149–56.
Aylon Y, Oren M. p53: guardian of ploidy. Mol Oncol. 2011;5:315–23.
Gronroos E, López-García C. Tolerance of chromosomal instability in cancer: mechanisms and therapeutic opportunities. Cancer Res. 2018;78:6529–35.
Bertheau P, Lehmann-Che J, Varna M, et al. p53 in breast cancer subtypes and new insights into response to chemotherapy. Breast. 2013;22:S27–9.
Thompson SL, Compton DA. Proliferation of aneuploid human cells is limited by a p53-dependent mechanism. J Cell Biol. 2010;188:369–81.
Mikule K, Delaval B, Kaldis P, Jurcyzk A, Hergert P, Doxsey S. Loss of centrosome integrity induces p38-p53-p211-dependent G1-S arrest. Nat Cell Biol. 2007;9:160–70.
Pati D, Haddad BR, Haegele A, Thompson H, Kittrell FS, Shepard A, et al. Hormone-induced chromosomal instability in p53-null mammary epithelium. Cancer Res. 2004;64:5608–16.
Burds AA, Schulze Lutum A, Sorger PK. Generating chromosome instability through the simultaneous deletion of Mad2 and p53. Proc Natl Acad Sci USA. 2005;102:11296–301.
Rausch T, Jones DTW, Zapatka M, Stütz AM, Zichner T, Weischenfeldt J, et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell. 2012;148:59–71.
Engeland K. Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ. 2018;25:114–32.
Voutsadakis IA. Chromosome 17 centromere amplification and chromosomal instability (CIN) in breast cancer: Pathogenic and therapeutic implications. Neoplasma. 2019 (in press).
Sadasivam S, DeCaprio JA. The DREAM complex: master coordinator of cell cycle-dependent gene expression. Nat Rev Cancer. 2013;13:585–95.
Weiss MB, Vitolo MI, Mohseni M, Rosen DM, Denmeade SR, Park BH, et al. Deletion of p53 in human mammary epithelial cells causes chromosomal instability and altered therapeutic response. Oncogene. 2010;29:4715–24.
Sage J, Straight AF. RB’s original CIN? Genes Dev. 2010;24:1329–33.
Oser MG, Fonseca R, Chakraborty AA, Brough R, Spektor A, Jennings RB, et al. Cells lacking the RB1 tumor suppressor gene are hyperdependent on Aurora B kinase for survival. Cancer Discov. 2019;9:230–47.
Gong X, Du J, Parsons SH, Merzug FF, Webster Y, Iversen PW, et al. Aurora A kinase inhibition is synthetic lethal with loss of the RB1 tumor suppressor gene. Cancer Discov. 2019;9:248–63.
Fukagawa T, Earnshaw WC. The centromere: chromatin foundation for the kinetochore machinery. Dev Cell. 2014;30:496–508.
Cheeseman IM. The kinetochore. Cold Spring Harb Perspect Biol. 2014;6:a015826.
Sacristan C, Kops GJPL. Joined at the hip: kinetochores, microtubules, and spindle assembly checkpoint signaling. Trends Cell Biol. 2015;25:21–8.
Schuyler SC, Wu Y-F, Kuan VJ-W. The Mad1-Mad2 balancing act—a damaged spindle checkpoint in chromosome instability and cancer. J Cell Sci. 2012;125:4197–4206.
Wang y, Jin F, Higgins R, McNight K. The current view for the silencing of the spindle assembly checkpoint. Cell Cycle. 2014;13:1694–1701.
Foley EA, Kapoor TM. Microtubule attachment and spindle assembly checkpoint signalling at the kinetochore. Nat Rev Mol Cell Biol. 2013;14:25–37.
Liu D, Vader G, Vromans MJ, Lampson MA, Lens SM. Sensing chromosome bi-orientation by spatial separation of Aurora B kinase from kinetochore substrates. Science. 2009;323:1350–3.
Zhao G, Cheng Y, Gui P, Cui M, Liu W, Wang W, et al. Dynamic acetylation of the kinetochore-associated protein HEC1 ensures accurate microtubule-kinetochore attachment. J Biol Chem. 2019;294:576–92.
Shepperd LA, Meadows JC, Sochaj AM, Lancaster TC, Zou J, Buttrick GJ, et al. Phosphodependent recruitment of Bub1 and Bub3 to Spc7/KNL1 by Mph1 kinase maintains the spindle checkpoint. Curr Biol. 2012;22:891–9.
Yamagishi Y, Yang CH, Tanno Y, Watanabe Y. MPS1/Mph1 phosphorylates the kinetochore protein KNL1/Spc7 to recruit SAC components. Nat Cell Biol. 2012;14:746–52.
Zhao Y, Chen R-H. Mps1 phosphorylation by MAP kinase is required for kinetochore localization of spindle-checkpoint proteins. Curr Biol. 2006;16:1764–9.
Yeh C-H, Yu Z-C, Chen P-H, Cheng Y-C, Shieh S-Y. Phosphorylation at threonine 288 by cell cycle checkpoint kinase 2 (CHK2) controls human monopolar spindle 1 (Mps1) kinetochore localization. J Biol Chem. 2014;289:15319–27.
Sullivan LL, Chew K, Sullivan BA. Α satellite DNA variation and function of the human centromere. Nucleus. 2017;8:331–9.
Dambacher S, Deng W, Hahn M, Sadic D, Fröhlich JJ, Nuber A, et al. CENP-C facilitates the recruitment of M18BP1 to centromeric chromatin. Nucleus. 2012;3:101–10.
Hara M, Fukagawa T. Kinetochore assembly and disassembly during mitotic entry and exit. Curr Opin Cell Biol. 2018;52:73–81.
Sharma AB, Dimitrov S, Hamiche A, Van Dyck E. Centromeric and ectopic assembly of CENP-A chromatin in health and cancer: old marks and new tracks. Nucleic Acids Res. 2019;47:1051–69.
Giunta S, Funabiki H. Integrity of the human centromere DNA repeats is protected by CENP-A, CENP-C, and CENP-T. Proc Natl Acad Sci USA. 2017;114:1928–33.
Hoffman S, Fachinetti D. A time out for CENP-A. Mol Cell Oncol. 2017;4:e1293596.
Hoffman S, Dumont M, Barra V, Ly P, Nechemia-Arbely Y, McMahon MA, et al. CENP-A is dispensable for mitotic centromere function after initial centromere/kinetochore assembly. Cell Rep. 2016;17:2394–404.
Mellone BG, Zhang W, Karpen GH. Frodos found: behold the CENP-A “ring” bearers. Cell. 2009;137:409–12.
Montes de Oca R, Gurard-Levin ZA, Berger F, Rehman H, Martel E, Corpet A, et al. The histone chaperone HJURP is a new independent prognostic marker for luminal A breast carcinoma. Mol Oncol. 2015;9:657–674.
Shrestha RL, Ahn GS, Staples MI, Sathyan KM, Karpova TS, Foltz DR, Basrai MA. Mislocalization of centromeric histone H3 variant CENP-A contributes to chromosomal instability (CIN) in human cells. Oncotarget. 2017;8:46781–800.
Curtis C, Shah SP, Chin S-F, Turashvili G, Rueda OM, Dunning MJ, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486:346–52.
Network Cancer Genome Atlas. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:269.
Howman EV, Fowler KJ, Newson AJ, Redward S, MacDonald AC, Kalitsis P, Choo KH. Early disruption of centromeric chromatin organization in centromere protein A (Cenpa) null mice. Proc Natl Acad Sci USA. 2000;97:1148–53.
Klare K, Weir JR, Basilico F, Zimniak T, Massimiliano L, Ludwigs N, et al. CENP-C is a blueprint for constitutive centromere-associated network assembly within human kinetochores. J Cell Biol. 2015;210:11–22.
Liao W-T, Feng Y, Li M-L, Liu G-L, Li M-Z, Zeng M-S, Song L-B. Overexpression of centromere protein H is significantly associated with breast cancer progression and overall patient survival. Chin J Cancer. 2011;30:627–37.
Tomonaya T, Matsushita K, Yamaguchi S, Oohashi T, Shimada H, Ochiai T, et al. Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res. 2003;63:3511–6.
Thangavelu PU, Lin C-Y, Vaidyanathan S, Nguyen THM, Dray E, Duijf PHG. Overexpression of the E2F target gene CENPI promotes chromosome instability and predicts poor prognosis in estrogen receptor-positive breast cancer. Oncotarget. 2017;8:62167–82.
Oliveras-Ferraros C, Vazquez-Martin A, Cuyàs E, Corominas-Faja B, Rodriguez-Gallego E, Fernández-Arroyo S, et al. Acquired resistance to metformin in breast cancer cells triggers transcriptome reprogramming toward a degradome-related metastatic stem-like profile. Cell Cycle. 2014;13:1132–44.
Ding M, Jiang J, Yang F, Zheng F, Fang J, Wang Q, et al. Holliday junction recognition protein interacts with and specifies the centromeric assembly of CENP-T. J Biol Chem. 2019;294:968–80.
Nishino T, Rago F, Hori T, Tomii K, Cheeseman IM, Fukagawa T. CENP-T provides a structural platform for outer kinetochore assembly. EMBO J. 2013;32:424–36.
Lee S, Gang J, Jeon SB, Choo SH, Lee B, Kim YG, et al. Molecular cloning and functional analysis of a novel oncogene, cancer-upregulated gene 2 (CUG2). Biochem Biophys Res Commun. 2007;360:633–9.
Nishino T, Takeuchi K, Gascoigne KE, Suzuki A, Hori T, Oyama T, et al. CENP-T-W-S-X forms a unique centromeric chromatin structure with a histone-like fold. Cell. 2012;148:487–501.
Lin S-Y, Lv Y-B, Mao G-X, Chen X-J, Peng F. The effect of centromere protein U silencing by lentiviral mediated RNA interference on the proliferation and apoptosis of breast cancer. Oncol Lett. 2018;16:6721–8.
Zhu D, Xu S, Deyanat-Yazdi G, Peng SX, Barnes LA, Narla RK, et al. Synthetic lethal strategy identifies a potent and selective TTK and CLK1/2 inhibitor for treatment of triple-negative breast cancer with a compromised G1-S checkpoint. Mol Cancer Ther. 2018;17:1727–38.
Endo H, Ikeda K, Urano T, Horie-Inoue K, I noue S. Terf/TRIM17 stimulates degradation of kinetochore protein ZWINT and regulates cell proliferation. J Biochem. 2012;151:139–144.
Yuan W, Xie S, Wang M, Pan S, Huang X, Xiong M, et al. Bioinformatic analysis of prognostic value of ZW10 interacting protein in lung cancer. Onco Targ Ther. 2018;11:1683–95.
Ying H, Xu Z, Chen M, Zhou S, Liang X, Cai X. Overexpression of Zwint predicts poor prognosis and promotes the proliferation of hepatocellular carcinoma by regulating cell-cycle-related proteins. Onco Targ Ther. 2018;11:689–702.
Ng CKY, Martelotto LG, Gauthier A, Wen H-C, Piscuoglio S, Lim RS, et al. Intra-tumor genetic heterogeity and alternative driver genetic alterations in breast cancers with heterogeneous HER2 gene amplification. Genome Biol. 2015;16:107.
Bièche I, Vacher S, Lallemand F, Tozlu-Kara S, Bennani H, Beuzelin M, et al. Expression analysis of mitotic checkpoint genes in breast carcinoma: role of NDC80/HEC1 in early breast tumorigenicity, and a two-gene signature for aneuploidy. Mol Cancer. 2011;10:23.
Yang K, Gao J, Luo M. Identification of key pathways and hub genes in basal-like breast cancer using bioinformatics analysis. Onco Targ Ther. 2019;12:1319–31.
Ur-Rehman S, Gao Q, Mitsopoulos C, Zvelebil M. ROCK: a resource for integrative breast cancer data analysis. Breast Cancer Res Treat. 2013;139:907–21.
Tang H, Xiao G, Behrens C, Schiller J, Allen J, Chow C-W, et al. A 12-gene set predicts survival benefits from adjuvant chemotherapy in non-small cell lung cancer patients. Clin Cancer Res. 2013;19:1577–86.
van Deursen JM. Rb loss causes cancer by driving mitosis Mad. Cancer Cell. 2007;11:1–3.
Yuan B, Xu Y, Woo J-H, Wang Y, Bae YK, Yoon D-S, et al. Increased expression of mitotic checkpoint genes in breast cancer cells with chromosomal instability. Clin Cancer Res. 2006;12:405–10.
Scintu M, Vitale R, Prencipe M, Gallo AP, Bonghi L, Valori VM, et al. Genomic instability and increased expression of BUB1B and MAD2L1 genes in ductal breast carcinoma. Cancer Lett. 2007;254:298–307.
Sun Q, Zhang X, Liu T, Liu X, Geng J, He X, et al. Increased expression of mitotic arrest deficient-like 1 (MAD1L1) is associated with poor prognosis and insensitive to Taxol treatment in breast cancer. Breast Cancer Res Treat. 2013;140:323–30.
Ryan SD, Britigan EMC, Zasadil LM, Witte K, Audhya A, Roopra A, Weaver BA. Up-regulation of the mitotic checkpoint component Mad1 causes chromosomal instability and resistance to microtubule poisons. Proc Natl Acad Sci USA. 2012;109:E2205–14.
Sotillo R, Hernando E, Díaz-Rodríguez E, Teruya-Feldstein J, Cordón-Cardo C, Lowe SW, Benezra R. Mad2 overexpression promotes aneuploidy and tumorigenesis in mice. Cancer Cell. 2007;11:9–23.
Rowald K, Mantovan M, Passos J, Buccitelli C, Mardin BR, Korbel JO, et al. Negative selection and chromosome instability induced by Mad2 overexpression delay breast cancer but facilitate oncogene-independent outgrowth. Cell Rep. 2016;15:2679–91.
Takagi K, Miki Y, Shibahara Y, Nakamura Y, Ebata A, Watanabe M, et al. BUB1 immunolocalization in breast carcinoma: its nuclear localization as a potent prognostic factor of the patients. Horm Canc. 2013;4:92–102.
Maciejczyk A, Szelachowska J, Czapiga B, Matkowski R, Haloń A, Györffy B, Surowiak P. Elevated BUBR1 expression is associated with poor survival in early breast cancer patients: 15-year follow-up analysis. J Histochem Cytochem. 2013;61:330–9.
Baker DJ, Dawlaty MM, Wijshake T, Jeganathan KB, Malureanu L, van Ree JH, et al. Increased expression of BubR1 protects against aneuploidy and cancer and extends healthy lifespan. Nat Cell Biol. 2013;15:96–102.
Ocaña A, Pérez-Peña J, Díez-González L, Sánchez-Corrales V, Templeton A, Seruga B, et al. Transcriptomic analyses identify association between mitotic kinases, PDZ-binding kinase and BUB1, and clinical outcome in breast cancer. Breast Cancer Res Treat. 2016;156:1–8.
Turner N, Lambros MB, Horlings HM, Pearson A, Sharpe R, Natrajan r, et al. Integrative molecular profiling of triple negative breast cancers identifies amplicon drivers and potential therapeutic targets. Oncogene. 2010;29:2013–2023.
Yan H, Zhu S, Song C, Liu N, Kang J. Bone morphogenic protein (BMP) signaling regulates mitotic checkpoint protein levels in human breast cancer cells. Cell Signal. 2012;24:961–8.
Karra H, Repo H, Ahonen I, Löyttyniemi E, Pitkänen R, Lintunen M, et al. Cdc20 and securing overexpression predict short-term breast cancer survival. Br J Cancer. 2014;110:2905–13.
Daniel J, Coulter J, Woo J-H, Wilsbach K, Gabrielson E. High levels of the Mps1 checkpoint protein are protective of aneuploidy in breast cancer cells. Proc Natl Acad Sci USA. 2011;108:5384–9.
Xu Q, Xu Y, Pan B, Wu L, Ren X, Zhou Y, et al. TTK is a favorable prognostic biomarker for triple-negative breast cancer survival. Oncotarget. 2016;7:81815–29.
King JL, Zhang B, Li Y, Li KP, Ni JJ, Saavedra HI, Dong J-T. TTK promotes mesenchymal signaling via multiple mechanisms in triple negative breast cancer. Oncogenesis. 2018;7:69.
Voutsadakis IA. The network of pluripotency, epithelial mesenchymal transition and prognosis of breast cancer. Breast Cancer Targ Ther. 2015;7:303–19.
Berto A, Yu J, Morchoisne-Bolhy S, Bertipaglia C, Vallee R, Dumont J, et al. Disentangling the molecular determinants for cenp-F localization to nuclear pores and kinetochores. EMBO Rep. 2018;19:e44742.
O’Brien SL, Fagan A, Fox EJP, Millikan RC, Culhane AC, Brennan DJ, et al. CENP-F expression is associated with poor prognosis and chromosomal instability in patients with primary breast cancer. Int J Cancer. 2007;120:1434–43.
van der Waal MS, Hengeveld RCC, van der Horst A, Lens SMA. Cell division control by the chromosomal passenger complex. Exp Cell Res. 2012;318:1407–20.
Yang C, Tang X, Guo X, Niikura Y, Kitagawa K, Cui K, et al. Aurora B mediated ATM serine 1403 phosphorylation is required for mitotic ATM activation and the spindle checkpoint. Mol Cell. 2011;44:597–608.
Wheatley SP, Altieri DC. Survivin at a glance. J Cell Sci. 2019;132:jcs223826.
Wiedemuth R, Klink B, Töpfer K, Schröck E, Schackert G, Tatsuka M, Temme A. Survivin safeguards chromosome numbers and protects from aneuploidy independently from p53. Mol Cancer. 2014;13:107.
Tsukahara T, Tanno Y, Watanabe Y. Phosphorylation of the CPC by Cdk1 promotes chromosome bi-orientation. Nature. 2010;467:719–23.
Gohard FH, St-Cyr DJ, Tyers M, Earnshaw WC. Targeting the INCENP IN-box-Aurora B interaction to inhibit CPC activity in vivo. Open Biol. 2014;4:140163.
Huang H, Lampson M, Efimov A, Yen TJ. Chromosome instability in tumor cells due to defects in Aurora B mediated error correction at kinetochores. Cell Cycle. 2018;17:2622–36.
Zhang Y, Jiang C, Li H, Lv F, Li X, Qian X, et al. Elevated Aurora B expression contributes to chemoresistance and poor prognosis in breast cancer. Int J Clin Exp Pathol. 2015;8:751–7.
Larsen SL, Yde CW, Laenkholm A-V, Rasmussen BB, Duun-Henriksen AK, Bak M, et al. Aurora kinase B is important for antiestrogen resistant cell growth and a potential biomarker for tamoxifen resistant breast cancer. BMC Cancer. 2015;15:239.
Gully CP, Zhang F, Chen J, Yeung JA, Velazquez-Torres G, Wang E, et al. Antineoplastic effects of an Aurora B kinase inhibitor in breast cancer. Mol Cancer. 2010;9:42.
Yamanaka Y, Heike T, Kumada T, Shibata M, Takaoka Y, Kitano A, et al. Loss of Borealin/DasraB leads to defective cell proliferation, p53 accumulation and early embryonic lethality. Mech Dev. 2008;125:441–50.
Ma CX, Gao F, Luo J, Northfelt DW, Goetz M, Forero A, et al. NeoPalAna: neoadjuvant palbociclib, a cyclin-dependent kinase 4/6 inhibitor, and anastrozole for clinical stage 2 or 3 estrogen receptor-positive breast cancer. Clin Cancer Res. 2017;23:4055–65.
Dominguez-Brauer C, Thu KL, Mason JM, Blaser H, Bray MR, Mak TW. Targeting mitosis in cancer: emerging strategies. Mol Cell. 2015;60:524–36.
Thu KL, Silvester J, Elliott MJ, Ba-alawi W, Duncan MH, Elia AC, et al. Disruption of the anaphage-promoting complex confers resistance to TTK inhibitors in triple-negative breast cancer. Proc Natl Acad Sci USA. 2018;115:E1570–7.
Zhang C-Z, Spektor A, Cornils H, Francis JM, Jackson EK, Liu S, et al. Chromothripsis from DNA damage in micronuclei. Nature. 2015;522:179–84.
Knouse KA, Amon A. The micronucleus gets its big break. Nature. 2015;522:162–3.
Mackenzie KJ, Carroll P, Martin C-A, Murina O, Fluteau A, Simpson DJ, et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature. 2017;548:461–5.
Paul A, Edwards J, Pepper C, Mackay S. Inhibitory-Κb kinase (IKK) α and nuclear factor-κB (NFκB)-inducing kinase (NIK) as anti-cancer drug targets. Cells. 2018;7:176.
Voutsadakis IA. Immune blockade inhibitors and the radiation abscopal effect in gastrointestinal cancers. World J Gastrointest Oncol. 2018;10:221–7.
Narang P, Chen M, Sharma AA, Anderson KS, Wilson MA. The neoepitope landscape of breast cancer: implications for immunotherapy. BMC Cancer. 2019;19:200.
Taylor AM, Shih J, Ha G, Gao GF, Zhang X, Berger AC, et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell. 2018;33:676–89.
Bobustuc GC, Kassam AB, Rovin RA, Jeudy S, Smith JS, Isley B, et al. MGMT inhibition in ER positive breast cancer leads to CDC2, TOP2A, AURKB, CDC20, KIF20A, Cyclin A2, Cyclin B2, Cyclin D1, ERα and survivin inhibition and enhances response to temozolamide. Oncotarget. 2018;9:29727–42.
Swanton C, Nicke B, Schuett M, Eklund AC, Ng C, Li Q, et al. Chromosomal instability determines taxane response. Proc Natl Acad Sci USA. 2009;106:8671–6.
Janssen A, Kops GJPL, Medema RH. Elevating the frequency of chromosome mis-segregation as a strategy to kill tumor cells. Proc Natl Acad Sci USA. 2009;106:19108–13.
Schmidt M, Budirahardja Y, Klompmaker R, Medema RH. Ablation of the spindle assembly checkpoint by a compound targeting Mps1. EMBO Rep. 2005;6:866–72.
Zasadil LM, Andersen KA, Yeum D, Rocque GB, Wilke LG, Tevaarwerk AJ, et al. Cytotoxicity of paclitaxel in breast cancer is due to chromosome missegration on multipolar spindles. Sci Transl Med 2014;6:229ra43.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
IAV declares no conflicts of interest related to this article.
Funding
No funding was received from any source to perform this work.
Rights and permissions
About this article
Cite this article
Voutsadakis, I.A. Clinical Implications of Chromosomal Instability (CIN) and Kinetochore Abnormalities in Breast Cancers. Mol Diagn Ther 23, 707–721 (2019). https://doi.org/10.1007/s40291-019-00420-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-019-00420-2