Abstract
Since the discovery and classification of non-coding RNAs, their roles have gained great attention. In this respect, microRNAs and long non-coding RNAs have been firmly demonstrated to be linked to regulation of gene expression and onset of human diseases, including rare genetic diseases; therefore they are suitable targets for therapeutic intervention. This issue, in the context of rare genetic diseases, is being considered by an increasing number of research groups and is of key interest to the health community. In the case of rare genetic diseases, the possibility of developing personalized therapy in precision medicine has attracted the attention of researchers and clinicians involved in developing “orphan medicinal products” and proposing these to the European Medicines Agency (EMA) and to the Food and Drug Administration (FDA) Office of Orphan Products Development (OOPD) in the United States. The major focuses of these activities are the evaluation and development of products (drugs, biologics, devices, or medical foods) considered to be promising for diagnosis and/or treatment of rare diseases or conditions, including rare genetic diseases. In an increasing number of rare genetic diseases, analysis of microRNAs and long non-coding RNAs has been proven a promising strategy. These diseases include, but are not limited to, Duchenne muscular dystrophy, cystic fibrosis, Rett syndrome, and β-thalassemia. In conclusion, a large number of approaches based on targeting microRNAs and long non-coding RNAs are expected in the field of molecular diagnosis and therapy, with a facilitated technological transfer in the case of rare genetic diseases, in virtue of the existing regulation concerning these diseases.
Similar content being viewed by others
Since non-coding RNAs’ discovery and classification, their roles in genetic diseases have gained great attention. |
Both microRNAs (miRNAs) and long non-coding RNAs (lncRNAs) can be considered excellent markers for advanced diagnostic protocols. |
Both miRNAs and lncRNAs are suitable targets for therapeutic intervention. |
Diagnostics based on miRNA and lncRNA profiles might be a key factor for the development of personalized therapy in precision medicine of genetic diseases. |
1 Key Roles of Non-coding RNAs in Gene Regulation
The expression of eukaryotic genes is a multistep and complex process, finely regulated by many different factors and levels of control [1], including (but not limited to) transcription factors (TFs), promoter sequences and enhancers [2,3,4], post-transcriptional regulation, and control of translation [5]. In this context, non-coding RNAs (ncRNAs) play a very important role [6,7,8]; they are composed of several classes, among which microRNAs (miRNAs) [9,10,11] and long non-coding RNAs (lncRNAs) [12,13,14,15,16] are of key interest.
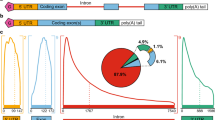
MicroRNAs (19–25 nucleotides in length) are a family of small ncRNAs with the important function of inhibiting or reducing protein synthesis after formation of complexes with target messenger RNAs (mRNAs) through base pairing [9,10,11, 17, 18]. Target mRNAs can hybridize to complementary miRNAs at the 3′-untranslated region (3′-UTR), coding sequences (CDS) and 5′-untranslated region (5′-UTR) [17, 18]. When this biological function is considered in its deep complexity, it can be concluded that more than 60% of human mRNAs are regulated by miRNAs [17, 18]. The miRNA/mRNA interaction causes translational repression or mRNA degradation, depending on the degree of complementarity at the level of RNA-induced silencing complex (RISC) [19, 20]. At present, the number of human miRNAs identified and deposited in the miRBase databases (miRBase v.22, http://www.mirbase.org) is more than 2500 [21,22,23]. Several reports conclusively support the concept that miRNAs are involved in the onset and progression of rare genetic diseases [24,25,26,27,28,29,30,31,32].
As already mentioned, lncRNAs have been also deeply implicated in the control of gene expression [12,13,14,15,16]. lncRNAs are characterized by a length of more than 200 nucleotides, and their effects on gene expression in the development of diseases are based on a variety of mechanisms of action, including recruitment of chromatin modifiers, controlled recruitment of TFs (both activators and repressors of gene transcription), regulation of RNA splicing, regulation of chromosome looping, control of mRNA translation and decay, and miRNA sponging [17,18,19,20,21]. Figure 1 summarizes the interplay between miRNAs, TFs, and lncRNAs.
2 Non-coding RNAs as Possible Targets in Theranostics of Genetic Diseases
While miRNAs are firmly established as being involved in rare genetic diseases [24,25,26,27,28,29,30,31,32], examples of lncRNAs affecting gene expression in these diseases are fewer, but this research field is rapidly growing. Two major issues are related to the finding that ncRNAs are possible markers of rare genetic disease: (1) molecular diagnosis based on ncRNAs might open new avenues in the fields of diagnosis, prognosis, patient stratification, and response to therapy; (2) identification of molecular targets will help in developing novel therapeutic approaches, possibly leading to novel strategies in precision medicine.
In the field of molecular diagnosis, in addition to well-established technologies, ncRNAs are very important markers for non-invasive diagnostic approaches, having implications in diagnosis and/or prognosis. Among these approaches, we would like to mention molecular analyses, which are an alternative to standard biopsies, known as “non-invasive liquid biopsies”, and aim to quantify and determine ncRNA patterns in body fluids (Fig. 2). For instance, the characterization of circulating cell-free miRNAs in serum helps in concluding diagnostic and prognostic assessments, based on the fact that disease-specific miRNAs can be identified in extracellular body fluids (such as serum, plasma, and saliva), supporting their potential as biomarkers [24,25,26,27, 30, 33,34,35]. The presence of miRNAs in body fluids is based on the fact that extracellular miRNAs are protected by exosome-like structures and small intraluminal vesicles released by a large variety of cells [36, 37].
Liquid biopsy versus standard biopsy in human pathologies. Analysis of non-coding circulating RNAs is helpful for patient stratification and to verify disease staging and response to treatment. The liquid biopsy approach has been applied to some rare genetic diseases, such as DMD. DMD Duchenne muscular dystrophy
Also lncRNAs were reported to be of great interest for liquid biopsy, especially in the field of diagnostics of cancer [38,39,40]. While circulating lncRNAs have been proposed as useful markers in Crohn’s disease [41], gynecological diseases [42], coronary artery disease [43], Sjögren’s syndrome [44], and osteoporosis [45], their application to stratification of patients affected by rare genetic diseases is under investigation and limited to a few examples [46,47,48,49].
With respect to translational medicine, miRNAs are suitable targets for therapeutic intervention, as summarized in Fig. 3 [50,51,52,53,54,55,56,57]. A forced decrease of miRNA activity can be obtained using miRNA-inhibitor oligomers (such as in miRNA antisense therapy based on RNA, DNA, locked nucleic acid [LNA] and other DNA analogs) [58,59,60,61,62,63], miRNA sponges [64,65,66] and mowers [67], or through miRNA masking that interferes with miRNA function by masking the miRNAs binding site of target mRNAs through hybridization with complementary molecules [68, 69]. In contrast, the use of modified miRNA mimetics, either synthetic or produced by plasmid or lentiviral vectors, might lead to potentiation of miRNA function (miRNA replacement therapy) [70,71,72,73].
Targeting lncRNAs is a more complicated process and largely depends on the mechanism of action of the studied lncRNA. Inhibition of the recruitment of chromatin modifiers and/or TFs can be achieved with molecules altering the scaffold of lncRNAs; antisense molecules can be employed for alteration of the lncRNA-mediated regulation of RNA splicing, regulation of chromosome looping, and control of mRNA translation and decay; inhibition of miRNA sponging can be obtained either by knocking down lncRNAs with the use of antisense molecules activating RNAse H or by competing with the lncRNA/miRNA interaction.
3 Examples of Non-coding RNAs in Rare Genetic Diseases
In this section, we will consider selected examples of miRNAs and lncRNAs that could be considered biomarkers of rare genetic diseases. The involvement of ncRNAs has been proposed in studies based on a variety of biological samples, including tissue samples, primary cells (including stem cells) and body-derived fluids (including serum, plasma, and saliva). When available, examples of targeting of ncRNA for therapeutic protocols development will be considered. Table 1 [74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116] and Table 2 [46,47,48, 117,118,119,120,121,122,123,124,125,126,127,128,129,130] report a more extensive analysis of the available examples of miRNAs (Table 1) and lncRNAs (Table 2) in rare genetic diseases.
3.1 Duchenne Muscular Dystrophy
Duchenne muscular dystrophy (DMD) is a lethal X-linked neuromuscular disease caused by mutations in the dystrophin gene [131]. DMD is the most common muscular dystrophy affecting children and, characterized by a rapid progression of muscle degeneration, causing, in the absence of medical intervention, loss of ambulation and death within the second decade of life. Li et al. applied the “liquid biopsy strategy” to DMD and demonstrated that circulating miRNAs are biomarkers useful for the assessment of the clinical status of DMD patients [132]. This possibility is relevant for the development of non-invasive biomarkers in diagnostic/prognostic protocols and very useful for the replacement of invasive muscle biopsy in DMD patients. Serum levels of six muscle-specific miRNAs (miR-1, miR-206, miR-133, miR-499, miR-208a, and miR-208b, also known as myomiRs) were analyzed by Li et al. [132] and found to be all elevated in DMD patients. More recently, digital reverse transcription polymerase chain reaction (RT-PCR) was employed for quantification of miR-30c and miR-181a and validated for detection of these DMD serum biomarkers [34]. According to this and similar studies, a limited and validated list of “dystromirs” (miR-1, miR-133a, miR-133b, miR-206, and miR-31) have been proposed as serum biomarkers not only for monitoring the disease severity in DMD [27], but also to monitor progression of the disease and response to therapies [26]. Moreover, the study of the miRNA pattern allowed identification of strong differences between Becker’s muscular dystrophy and DMD patients [33]. In line with these findings, three serum biomarkers, miR-1, miR-133a, and miR-206, were characterized for limb-girdle muscular dystrophy, Becker’s muscular dystrophy and facioscapulohumeral muscular dystrophy [33].
With respect to possible applications in therapy, in vitro approaches have demonstrated that miRNA therapy can be successfully combined with other strategies. For instance Hildyard and Wells reported that miR-31 targeting might be of great interest [133], considering that translational suppression of dystrophin mRNA is caused by miR-31, which is elevated in the muscle of DMD patients. Therefore, they verified whether oligonucleotide-mediated exon skipping (leading to increase of dystrophin mRNA) could be conducted together with inhibition of miR-31 activity (leading to stabilization of the increased dystrophin mRNA levels), obtaining, at least in cell culture models, increased dystrophin production. Another very interesting approach involving miRNA targeting in DMD was recently reported by Mishra et al., and was based on the observation that the overexpression of utrophin, the autosomal homolog of dystrophin, can to some extent compensate for the dystrophin absence [134]. Since several miRNAs (including let-7c) are able to inhibit utrophin mRNA, the hypothesis by Mishra et al. was that a forced inhibition of utrophin/let-7c interaction was potentially able to “repress the repression,” therefore increasing utrophin expression [134]. Following this hypothesis, an oligonucleotide was designed and validated for efficient annealing with the 3′-UTR of the utrophin mRNA. This was associated with inhibition of let-7c miRNA binding, upregulation of utrophin expression, and improvement of the dystrophic phenotype in vivo. A final remark is related to the opposing roles of different miRNAs on the progression of fibrosis in DMD. Zanotti et al. found that miR-21 and miR-29 were differentially regulated in muscle biopsies from DMD patients [135]. In Duchenne muscle biopsies, miR-21 expression was significantly increased, and correlated directly with transcript levels of the COL1A1 and COL6A1 genes. On the other hand, miR-29a and miR-29c were significantly reduced in Duchenne muscle and myoblasts, and miR-29 target transcripts, COL3A1, FBN1 and YY1, significantly increased. Therefore miR-21 silencing on one hand and miR-29 mimicking on the other were able to reduce fibrosis in mdx mice, suggesting that pharmacological modulation of their expression has therapeutic potential for reducing fibrosis in this condition [135].
The involvement of lncRNAs in DMD was the subject of a limited number of studies, based on novel technologies such as transcriptome sequencing (RNA-seq), which highlight the possible importance of these novel RNAs [46]. For example, Ballarino et al. showed that the human lncRNA linc-MD1 is expressed during early stages of normal myoblast differentiation as well as in human primary myoblasts from DMD patients [117]. Twayana et al. clarified that its mechanism of action was based on “sponging” miR-133 and miR-135 [118]. Interestingly, linc-MD1 is also the host transcript of miR-133b, and their biogenesis is mutually exclusive. Furthermore, Bovolenta et al. identified a variety of novel lncRNAs (both sense and antisense), whose expression mirrored that of the DMD gene [136]. These transcripts are intronic in origin, specifically localized within the nucleus, and are transcribed contextually with dystrophin isoforms or primed by MyoD-induced myogenic differentiation.
3.2 Amyotrophic Lateral Sclerosis
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease characterized by loss of motor neurons, denervation of target muscles, muscle atrophy, and paralysis [137]. Potential therapeutic targets for ALS were reported by Di Pietro et al., who found that changes in the expression of miR-206, miR-208b and miR-499 occur in the skeletal muscle of patients during progression of the ALS disease [29], and by Waller et al., who reported that serum miRNAs miR-206, 143-3p and 374b-5p are promising ALS biomarkers [30]. For the therapeutic point of view, it was reported that deficiency of miR-206 in ALS mouse models accelerates disease progression [86], supporting the concept that efficient regeneration of neuromuscular synapses after acute nerve injury requires miR-206. Therefore, miR-206 mimicking could be beneficial in ALS therapy.
A few reports are available on lncRNAs in ALS [47, 120, 138]. In this respect, Gagliardi et al. performed a whole transcriptome RNA-seq analysis to investigate the regulation of non-coding and coding RNAs in sporadic ALS (SALS) patients, mutated ALS patients and peripheral blood mononuclear cells (PBMC) isolated from matched controls [47]. A total of 293 differentially expressed lncRNAs were found in SALS patients, whereas a limited number of lncRNAs were deregulated in mutated patients [47]. These data highlighted the importance of extending the knowledge on the significance of regulatory lncRNA classes in the understanding of ALS disease. This study offers a starting point for new investigations into the pathogenic mechanisms involved in ALS disease.
3.3 Cystic Fibrosis
Cystic fibrosis (CF) is an autosomal recessive genetic disorder affecting in addition to the lungs, other organs, such as the pancreas, liver, kidneys, and intestine [139]. As a result of frequent and heavy lung infection, CF patients exhibit important difficulty in breathing and frequent coughs with extensive mucus [140]. CF is caused by several classes of mutations of the gene for the CF transmembrane conductance regulator (CFTR) protein. With respect to the CF pathology, miRNA-based epigenetic regulation of expression of the CFTR gene and CFTR regulators has been recently proposed by different groups after extensive studies performed using CF primary bronchial epithelial cells in vitro or from bronchial brushings ex vivo [88,89,90,91,92,93,94,95,96]. The general conclusion is that miRNAs are involved in CF. For instance, the expression of miR-145 and miR-494 was found anti-regulated with that of CFTR gene [88]. The interplay between miRNAs and TFs regulating the CFTR gene is supported by a study focusing on genome-wide expression of miRNAs in primary non-CF bronchial epithelial cells and demonstrating reduced expression of miR-138, a downregulator of SIN3A (a transcriptional repressor of CFTR) [90]. A direct effect of miRNAs on CFTR expression was on the other hand supported by the finding of a high expression in CF cells of miR-494 and miR-509-3p (able to interact with the 3′-UTR sequence of the CFTR mRNA) [91]. These in vitro findings were confirmed in ex vivo analyses demonstrating an increased expression of miR-494, miR-223 and miR-145 in CF brushings of airway cells [92]. Taken together, different miRNAs whose expression was found changed in CF primary bronchial epithelial cells can reduce CFTR expression, either by direct (miR-145, miR-223, miR-494, and miR-509-3p) or indirect (miR-138) effects. In this respect, it should be underlined that in addition to direct interaction with CFTR mRNA, miRNAs might regulate CFTR through binding to 3′-UTR sequences of mRNAs coding CFTR regulators, such as a large series of proteins known to interact with the CFTR protein in the quality-control mechanisms to regulate its folding and misfolding [141]. Among the so-termed “CFTR interactome”, some proteins have been shown to interact with CFTR to stabilize its expression on the apical membrane of the epithelial cells [142, 143]. CFTR regulators include, but are not limited to, NHERF1, NHERF2, ezrin, PDZK1, PDZK2, and Shank2. The data presented by Fabbri et al. [93] showed that a peptide nucleic acid (PNA) directed against miR-145-5p causes miR-145-5p inhibition and increases the miR-145-5p regulated CFTR, after quantification of CFTR expression by quantitative reverse transcription PCR (RT-qPCR) and Western blotting. Therefore, miRNA targeting in CF might be considered as a useful strategy to increase CFTR content, supporting the possible application in the personalized therapy of cystic fibrosis. With respect to miR-145-5p involvement in CFTR expression, we have to underline that the promoter of this miRNA is under strong negative control, one of the involved TFs being Tcf4, as schematically represented in Fig. 4 [93, 144]. Interestingly, Tcf4 binds to an intronic enhancer present in the intron I of the CFTR gene [145]. Therefore, low expression of Tcf4 (as it occurs in CF) [146,147,148] might lead to low transcription of the CFTR gene, increased miR-145-5p transcription, and high miR-145-5p—dependent down modulation of CFTR mRNA. From the theoretical point of view, miR-145 might cause an increase of CFTR in consideration of the fact that miR-145-5p downregulates Sp1 [149], a TF modulating CFTR gene transcription (see Fig. 4). Therefore, PNAs targeting miR-145-5p might interfere with the direct binding of miR-145-5p to the 3′-UTR CFTR mRNA sequences; in addition, the same PNA might indirectly potentiate CFTR by interfering with the miR-145-5p/Sp1 mRNA interactions (Fig. 4). Furthermore, given the central role of Tcf4 in enhancing CFTR gene transcription and inhibiting miR-145-5p expression, a possible therapeutic strategy might consider enhancing Tcf4 [144, 145]. In this respect, considering that miR-155 is among the possible Tcf4 targeting miRNAs [150, 151], the development of anti-miR-155 PNAs might be of interest in the future [151].
The proposed interplay between transcription factors, CFTR regulation, miR-145-5p transcription and miR-145-5p target molecules [93]. This might have important implications for drug design, suggesting development of approaches able to perform multi-targeting of microRNAs involved in complex activities. CFTR cystic fibrosis transmembrane conductance regulator
In addition to miR-145-5p, several other miRNAs have been proposed to downregulate CFTR expression, such as miR-494, miR-509-3p, miR-101, and miR-433 [88,89,90,91,92,93,94,95,96, 152, 153]. Therefore, screening of PNAs targeting these miRNAs, identification of the most active, and combined treatments using the more efficient should be considered to reach a CFTR increase compatible with physiological effects.
Also lncRNAs are implicated in CF. In one study, McKiernan et al. demonstrated that lncRNAs are aberrantly expressed in vivo in the CF bronchial epithelium [121]. In a second study, the possible role of lncRNAs in CF was analyzed on CF cells infected by Pseudomonas aeruginosa [122], the leading cause of chronic lung infection in CF patients, associated with an exaggerated inflammatory response. Since lncRNAs are deeply involved in regulating innate immune response, Balloy et al. hypothesized that CF cells exposed to P. aeruginosa could express a specific lncRNA signature. This study identified temporal expression signatures of 26 lncRNA transcripts differentially expressed after 6 h of P. aeruginosa infection, which differentiate CF from non-CF cells [122]. The differential expression of two lncRNAs (BLACAT1 and MEG9) was independently validated using real-time PCR.
3.4 Rett Syndrome
Rett syndrome (RTT) is a complex X-linked neurological disorder associated with mutations in the gene coding for the methyl CpG binding protein 2 (Mecp2 gene) [154], causing a severe phenotype. The high frequency (one out of every 10,000–15,000 live births) makes the RTT disorder the second most frequent cause of mental retardation in females. The miRNA miR-199a was found by Tsujimura et al. to link MeCP2 with mammalian target of rapamycin (mTOR) signaling, and its dysregulation leads to RTT phenotypes. In RTT, miRNAs dysregulation caused by Mecp2 absence has been reported. In particular, it was demonstrated that the most commonly disrupted miRNAs were miR-146a, miR-146b, miR-130, miR-122a, miR-342, and miR-409 (downregulated) and miR-29b, miR-329, miR-199b, miR-382, miR-296, miR-221 and miR-92 (upregulated) [155]. As far as lncRNAs are concerned, the study by Petazzi et al. focused on the impact of Mecp2 impairment on lncRNAs pattern using an RTT mouse model [124]. Using a microarray platform, an aberrant lncRNA transcriptome was identified in the brain of RTT mice. In particular, the transcripts of the AK081227 and AK087060 genes were found to be upregulated in Mecp2-null mice brains. Furthermore, this study highlighted that the overexpression of AK081227 mediated by the Mecp2 loss was associated with the downregulation of its host coding protein gene, the gamma-aminobutyric acid receptor subunit Rho 2 (Gabrr2) [124]. These observations support the concept that dysregulation of lncRNAs transcription upon Mecp2 loss might contribute to the neurological RTT phenotype.
3.5 Sézary Syndrome
This is a rare and aggressive leukemic variant of cutaneous T-cell lymphoma (CTCL). This disease is characterized by the presence of neoplastic lymphocytes named Sézary cells in the skin, lymph nodes, and peripheral blood [156, 157]. Studies based on deep-sequencing demonstrated that the miR-199a2/214 cluster represents the vast majority of aberrantly expressed miRNAs in Sézary syndrome (SS) [31]. On the other hand, Ballabio and colleagues reported that a high proportion of SS-associated miRNAs were downregulated, consistent with previously reported abnormalities of the genomic copy number [100]. Altered miR-223 levels were found to discriminate SS samples from healthy controls; inhibition of apoptosis was found to be associated in SS samples with downregulation of miR-342 and was hypothesized to be involved in the pathogenesis of this disease [100]. The involvement of miRNAs in SS was also strongly supported by another study in which a miRNA signature was identified that was potentially able to discriminate patients with unfavorable and favorable outcomes [101]. Among the miRNAs identified, miR-21 appears to be important. In line with this hypothesis, van der Fits and colleagues demonstrated that miR-21 expression is increased in SS cells when compared with cells from healthy subjects. Functional studies focusing on miR-21 in SS were able to demonstrate that silencing of miR-21 in SS cells is associated with increased apoptosis, suggesting a role for miR-21 in the leukomogenic process. These studies propose miR-21 as a possible molecular target in protocols finalized for the therapeutic treatment of SS [102].
As far as lncRNAs are concerned, among the few studies available, one reported by Lee et al. was based on bioinformatics, which allowed the identification of SS-associated lncRNAs [158]. To our knowledge, this was, among the studies on SS, the only one that, starting from the characterization of the SS transcriptome, supports an implication of lncRNA dysregulation in SS.
3.6 Multiple Osteochondromas
Multiple Osteochondromas (MOs) are characterized by a high number of mutations affecting the EXT1/EXT2 genes [104]. At the same time, a significant population of MO patients does not exhibit alterations in the EXT genes. Intra- and interfamilial variability of the MO phenotype is present, for which the molecular basis is largely unknown. For this reason, appropriate diagnostic/prognostic markers and personalized clinical management for both benign and malignant lesions are not available so far [104]. In this respect, a miRNA signature could help in patient stratification. Zuntini et al. reported the miRNA pattern of MO, differentiating disease-specific from normal cartilage miRNA signatures [104]. In particular, a signature of eight miRNAs (miR-21, miR-140, miR-145, miR-214, miR-195, miR-199a, miR-451, and miR-483) was associated with MO and was useful to distinguish MO patients from unaffected subjects. The obtained data indicate that differentially expressed miRNAs may contribute to the molecular signaling responsible for normal proliferation and differentiation of chondrocytes, playing a role in both pathogenesis and clinical outcome [104]. To the best of our knowledge, no examples of lncRNAs associated with MOs are so far available.
3.7 β-Thalassemia
β-Thalassemia is a recessive genetic disease caused by more than 300 mutations affecting the β-globin gene [159]. This leads to absence or lower production of the adult β-globin and, consequently, hemoglobin. The management of β-thalassemia patients is mostly based on blood transfusion, chelation therapy and, alternatively, on bone marrow transplantation [160]. Recently, novel therapeutic options are explored, such as gene therapy and fetal hemoglobin (HbF) induction [161]. Despite the fact that these approaches are promising, these therapeutic strategies are at present still under deep experimental development, involving a limited number of clinical trials. Furthermore, it should be considered that new HbF inducers and/or new approaches for reactivation of γ-globin genes are required, since important limitations are exhibited by hydroxyurea (the most used HbF inducer in β-thalassemia), including (1) toxicity; (2) lack of response by about 50% of β-thalassemia patients; and (3) development of drug resistance with long-term treatments [161].
In this respect, of great interest is the demonstration of the fact that the transcriptional regulation of the human γ-globin genes is under the control of several transcriptional repressors, including MYB, BCL11A, KLF-1, KLF-2, Sp1, and LYAR [162,163,164]. This has greatly facilitated the identification of specific targets for the development of possible pre-clinical approaches to reactivate γ-globin gene expression (and consequently HbF production) by targeting these TFs. Among possible additional strategies for HbF potentiation is the use of miRNA-based approaches. In fact, miRNAs involved in the control of γ-globin gene repressors have been identified and validated. Examples are miR-486-3p (targeting BCL11A mRNA), miR-23a (targeting KLF-2) miR-15a and miR-16-1 (targeting MYB mRNA), and miR-27a (targeting Sp1) [110] (Fig. 5).
While with respect to other HbF-associated miRNAs, information is still lacking, miR-210 has been suggested by different groups to be somehow involved in HbF production. It has been reported that miR-210 is involved in erythroid differentiation and, possibly, in γ-globin gene upregulation [165,166,167,168,169]. For instance, a recent paper by Bavelloni et al. [168] confirmed the data published by Bianchi et al. [166], demonstrating an increase in miR-210 in K562 cells treated with mithramycin, one of the most powerful HbF inducers in erythroid cells. Sarakul et al. demonstrated upregulation of miR-210 under hypoxic conditions in K562 cells and in β-thalassemia/HbE (hemoglobin E) progenitor cell cultures [167]. Inhibition of miR-210 expression leads to reduction in γ-globin gene expression, indicating that miR-210 contributes to hypoxia-induced erythroid differentiation. The possible role of miR-210 in HbF production was also proposed in the communication by Sawant et al. that suggested that HbF induction by hydroxycarbamide works through miR-210 in sickle cell anemia patients [169].
In a recent study, we analyzed a coding sequence of BCL11A mRNA as a possible target of miR-210 [110]. This miR-210 site of BCL11A mRNA is conserved through molecular evolution, possibly indicating that this sequence plays a key biological function. Interactions between miR-210 and the miR-210 BCL11A mRNA site were demonstrated by surface plasmon resonance (SPR)-based biomolecular interaction analysis (BIA), and forced expression of miR-210 leads to a decrease in BCL11A mRNA and an increase in γ-globin mRNA content in erythroid cells, including erythroid precursors isolated from β-thalassemia patients. While most validated miRNA/mRNA interactions involve the 3′-UTR of the target mRNAs [170,171,172,173,174,175,176,177,178], functional interactions between miRNAs and coding sequences of target mRNAs have been reported in several studies [179,180,181]. Our study suggests that, in addition to the already reported binding of miR-486-3p to the 3′-UTR sequence of the BCL11A mRNA, also the coding mRNA sequence of BCL11A can be targeted by miRNAs (such as miR-210). Therefore, upregulation of γ-globin gene expression can be achieved by different miRNA-based approaches, including the use of pre-miRNA targeting the 3′-UTR region of the BCL11A mRNA, or the use of pre-miR-210, targeting the coding region of the mRNA. Both approaches (that at least in theory can be combined) might lead to BCL11A downregulation and induction of γ-globin gene expression (Fig. 6), in consideration of the fact that BCL11A is a transcriptional repressor of γ-globin genes. This is of interest, since inhibition of BCL11A is a recognized strategy for HbF induction for treatment of β-thalassemia, as suggested by several papers and patent applications [110, 173].
Scheme outlining the possible miRNA mimicking approach for targeting BCL11A mRNA. Downregulation of BCL11A gene expression will lead to upregulation of γ-globin mRNA [110]. mRNA messenger RNA, miRNA microRNA
One of very few examples of validated lncRNAs in β-thalassemia [48, 129, 130] was described by Morrison et al., who characterized a novel 1283 bp transcript (HMI-LNCRNA) that was transcribed from the enhancer region of the MYB gene coding for one of the major transcriptional repressor of β-globin genes [129]. Within erythroid cells, HMI-LNCRNA was almost exclusively present in the nucleus, and was significantly higher in erythroblasts derived from cultured adult peripheral blood CD34+ cells, which expressed adult hemoglobin (HbA), compared to erythroblasts from cultured cord blood CD34+ cells, which expressed much more HbF. Downregulation of HMI-LNCRNA in HUDEP-2 cells significantly upregulated HbF expression both at the mRNA (200-fold) and protein levels. Therefore, HMI-LNCRNA might be a potential therapeutic target for HbF induction treatment in sickle cell disease and β-thalassemia.
4 Concluding Remarks
Since ncRNAs’ discovery and classification, their roles have gained great attention for several reasons, including their possible employment as novel disease markers as well as novel therapeutic targets. In this respect, miRNA and lncRNAs are key players (see Fig. 1), as they have been firmly linked to regulation of gene expression and to the onset of human diseases. While the majority of examples available in the literature are focused on cancer, examples of the involvement of ncRNAs in genetic diseases (including rare genetic diseases) are growing in number. Most of the examples available are based on the analysis of miRNAs, and only a limited (but growing) number have been reported for lncRNAs.
As far as applications in diagnostic and therapeutic protocols are concerned, both miRNAs and lncRNAs can be considered excellent markers for advanced diagnostic protocols, including liquid biopsy (see Fig. 2). In addition, progress has been made with respect to genetic mutations of miRNAs as well as of miRNA-binding sites affecting the 3′-UTR sequences of miRNA-regulated genes. Since these genetic alterations can be detected in rare cellular subpopulations, taking advantage of the knowledge driven by liquid biopsy approaches, we would expect future applications also in non-invasive prenatal diagnosis, which is particularly relevant in the case of genetic diseases and is based on the detection of genetic mutations in fetal cells circulating within the mother’s peripheral blood.
With respect to translational medicine, miRNAs and lncRNAs are suitable targets for therapeutic intervention [51,52,53,54,55,56,57]. Inhibition of miRNA activity has been achieved by different approaches [58,59,60,61,62,63,64,65,66,67,68,69]. Increase of miRNA function (miRNA replacement therapy) was also considered [70,71,72,73]. Also in the case of therapeutic interventions (as it occurs for diagnostic protocols), the examples of targeting lncRNAs for the development of therapeutic approaches are fewer compared with miRNAs. This is probably related to the fact that targeting in this case largely depends on the mechanism of action of the studied lncRNA. For instance, lncRNAs could act by sponging miRNAs, recruitment of chromatin modifiers, controlled recruitment of TFs (both activators and repressors of gene transcription), regulation of RNA splicing, regulation of chromosome looping, and control of mRNA translation and decay [12,13,14,15,16]. Very different approaches are therefore necessary depending on the lncRNA mechanism of action in the target cells.
Considering the key role of miRNAs and lncRNAs also in rare genetic diseases, a final comment should be focused on the issue of development of protocols based on these molecules. This issue is considered of key interest by an increasing number of research groups and patient associations. In the case of rare genetic diseases (unlike drugs and protocols developed for cancer, infectious diseases, and cardiovascular pathologies), it should be considered that even if excellent and innovative therapeutic strategies are available exhibiting high activity levels and promising specificity in pre-clinical tests, they are not expected to attract the interest of large pharmaceutical companies in consideration of the limited number of patients to be treated [182, 183]. The importance of taking this issue into consideration is also strongly related to the possibility of developing personalized therapy in precision medicine of rare genetic diseases [184]. In this context, the possibility to develop and propose “orphan medicinal products” [185] might be a key action, considering the deep advantages associated with these designations [186,187,188]. In Europe, the European Medicines Agency (EMA) continuously evaluates the applications for Orphan Medicinal Product designation on the basis of reports obtained from the Committee of the Orphan Medicinal Products (COMP). A similar activity is coordinated in United States by the Food and Drug Administration (FDA) Office of Orphan Products Development (OOPD) [204–206]. The major focuses of these activities are the evaluation and development of products (drugs, biologics, devices, or medical foods) considered to be promising for diagnosis and/or treatment of rare diseases or conditions, including rare genetic diseases. In conclusion, a large number of approaches are expected in the field of molecular diagnosis and therapy based on targeting miRNAs and lncRNAs, with a facilitated technological transfer in the case of rare genetic diseases.
In our opinion, the analysis of miRNAs and lncRNAs will be a key diagnostic and prognostic tool in the near future, helping clinicians to choose the therapeutic strategy to be activated in patients stratified with respect to the production of these ncRNA species. The protocols that will be developed are expected to significantly impact in the field of personalized therapy and precision medicine. In this respect, optimization of delivery systems, combined treatment targeting multiple miRNAs, and/or the development of protocols involving miRNA replacement (affecting pathology-linked downregulated miRNAs) associated with antagomiRNA therapy (affecting pathology-linked upregulated miRNAs) are expected to significantly contribute to the set-up of advanced therapeutic protocols in the future.
References
Amaral PP, Dinger ME, Mercer TR, Mattick JS. The eukaryotic genome as an RNA machine. Science. 2008;319:1787–9.
Ganapathi M, Srivastava P, Das Sutar SK, Kumar K, Dasgupta D, Pal Singh G, et al. Comparative analysis of chromatin landscape in regulatory regions of human housekeeping and tissue specific genes. BMC Bioinform. 2005;6:126.
Maston GA, Evans SK, Green MR. Transcriptional regulatory elements in the human genome. Annu Rev Genom Hum Genet. 2006;7:29–59.
Ong CT, Corces VG. Enhancer function: new insights into the regulation of tissue-specific gene expression. Nat Rev Genet. 2011;12:283–93.
Jackson RJ, Hellen CU, Pestova TV. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat Rev Mol Cell Biol. 2010;11:113–27.
Barrett LW, Fletcher S, Wiltons SD. Regulation of eukaryotic gene expression by the untranslated gene regions and other non-coding elements. Cell Mol Life Sci. 2012;69:3613–34.
Taft RJ, Pang KC, Mercer TR, Dinger M, Mattick JS. Non-coding RNAs: regulators of disease. J Pathol. 2009;220:126–39.
Morris KV. RNA-directed transcriptional gene silencing and activation in human cells. Oligonucleotides. 2009;19:299–306.
Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11:597–610.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351–79.
Clark MB, Mattick JS. Long noncoding RNAs in cell biology. Semin Cell Dev Biol. 2011;22:366–76.
Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 2009;23:1494–504.
Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152:1298–307.
Kung JT, Colognori D, Lee JT. Long noncoding RNAs: past, present, and future. Genetics. 2013;193:651–69.
Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014;5:e944014.
Maziere P, Enright AJ. Prediction of microRNA targets. Drug Discov Today. 2007;12:452–8.
Witkos TM, Koscianska E, Krzyzosiak WJ. Practical aspects of microRNA target prediction. Curr Mol Med. 2011;11:93–109.
Sun K, Lai EC. Adult-specific functions of animal microRNAs. Nat Rev Genet. 2013;14:535–48.
Chekulaeva M, Filipowicz W. Mechanisms of miRNA-mediated post-transcriptional regulation in animal cells. Curr Opin Cell Biol. 2009;21:452–60.
Friedländer MR, Lizano E, Houben AJS, Bezdan D, Báñez-Coronel M, Kudla G. Evidence for the biogenesis of more than 1000 novel human microRNAs. Genome Biol. 2014;15:R57.
Londin E, Loher P, Telonis AG, Quann K, Clark P, Jing Y. Analysis of 13 cell types reveals evidence for the expression of numerous novel primate-and tissue-specific microRNAs. Proc Natl Acad Sci USA. 2015;112:E1106–15.
Kozomara A, Griffiths-Jones S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014;42:D68–73.
Salvatore M, Magrelli A, Taruscio D. The role of microRNAs in the biology of rare diseases. Int J Mol Sci. 2011;12:6733–42.
Hu J, Kong M, Ye Y, Hong S, Cheng L, Jiang L. Serum miR-206 and other muscle-specific microRNAs as non-invasive biomarkers for Duchenne muscular dystrophy. J Neurochem. 2014;129:877–83.
Giordani L, Sandoná M, Rotini A, Puri PL, Consalvi S, Saccone V. Muscle-specific microRNAs as biomarkers of Duchenne muscular dystrophy progression and response to therapies. Rare Dis. 2014;2:e974969.
Zaharieva IT, Calissano M, Scoto M, Preston M, Cirak S, Feng L, et al. Dystromirs as serum biomarkers for monitoring the disease severity in Duchenne muscular dystrophy. PLoS One. 2013;8:e80263.
Recabarren-Leiva D, Alarcón M. New insights into the gene expression associated to amyotrophic lateral sclerosis. Life Sci. 2018;193:110–23.
Di Pietro L, Baranzini M, Berardinelli MG, Lattanzi W, Monforte M, Tasca G, et al. Potential therapeutic targets for ALS: MIR206, MIR208b and MIR499 are modulated during disease progression in the skeletal muscle of patients. Sci Rep. 2017;7:9538.
Waller R, Goodall EF, Milo M, Cooper-Knock J, Da Costa M, Hobson E, et al. Serum miRNAs miR-206, 143-3p and 374b-5p as potential biomarkers for amyotrophic lateral sclerosis (ALS). Neurobiol Aging. 2017;55:123–31.
Qin Y, Buermans HP, van Kester MS, van der Fits L, Out-Luiting JJ, Osanto S, et al. Deep-sequencing analysis reveals that the miR-199a2/214 cluster within DNM3os represents the vast majority of aberrantly expressed microRNAs in Sézary syndrome. J Investig Dermatol. 2012;132:1520–2.
Finotti A, Gambari R. Recent trends for novel options in experimental biological therapy of β-thalassemia. Expert Opin Biol Ther. 2014;14:1443–54.
Matsuzaka Y, Kishi S, Aoki Y, Komaki H, Oya Y, Takeda S, et al. Three novel serum biomarkers, miR-1, miR-133a, and miR-206 for limb-girdle muscular dystrophy, facioscapulohumeral muscular dystrophy, and Becker muscular dystrophy. Environ Health Prev Med. 2014;19:452–8.
Llano-Diez M, Ortez CI, Gay JA, Álvarez-Cabado L, Jou C, Medina J, et al. Digital PCR quantification of miR-30c and miR-181a as serum biomarkers for Duchenne muscular dystrophy. Neuromuscul Disord. 2017;27:15–23.
Anaya-Segura MA, Rangel-Villalobos H, Martínez-Cortés G, Gómez-Díaz B, Coral-Vázquez RM, Zamora-González EO, et al. Serum levels of microRNA-206 and novel mini-STR assays for carrier detection in Duchenne muscular dystrophy. Int J Mol Sci. 2016;17:E1334.
Grasedieck S, Sorrentino A, Langer C, Buske C, Döhner H, Mertens D, et al. Circulating microRNAs in hematological diseases: principles, challenges, and perspectives. Blood. 2013;121:4977–84.
Weiland M, Gao XH, Zhou L, Mi QS. Small RNAs have a large impact: circulating microRNAs as biomarkers for human diseases. RNA Biol. 2012;9:850–9.
Zhou X, Yin C, Dang Y, Ye F, Zhang G. Identification of the long non-coding RNA H19 in plasma as a novel biomarker for diagnosis of gastric cancer. Sci Rep. 2015;5:11516.
Isin M, Ozgur E, Cetin G, Erten N, Aktan M, Gezer U, et al. Investigation of circulating lncRNAs in B-cell neoplasms. Clin Chim Acta. 2014;431:255–9.
Guo F, Yu F, Wang J, Li Y, Li Y, Li Z, et al. Expression of MALAT1 in the peripheral whole blood of patients with lung cancer. Biomed Rep. 2015;3:309–12.
Chen D, Liu J, Zhao HY, Chen YP, Xiang Z, Jin X. Plasma long noncoding RNA expression profile identified by microarray in patients with Crohn’s disease. World J Gastroenterol. 2016;22:4716–31.
Wang WT, Sun YM, Huang W, He B, Zhao YN, Chen YQ. Genome-wide long non-coding RNA analysis identified circulating lncRNAs as novel non-invasive diagnostic biomarkers for gynecological disease. Sci Rep. 2016;6:23343.
Cai Y, Yang Y, Chen X, Wu G, Zhang X, Liu Y, et al. Circulating ‘lncRNA OTTHUMT00000387022’ from monocytes as a novel biomarker for coronary artery disease. Cardiovasc Res. 2016;112:714–24.
Sandhya P, Joshi K, Scaria V. Long noncoding RNAs could be potential key players in the pathophysiology of Sjögren’s syndrome. Int J Rheum Dis. 2015;18:898–905.
Tong X, Gu PC, Xu SZ, Lin XJ. Long non-coding RNA-DANCR in human circulating monocytes: a potential biomarker associated with postmenopausal osteoporosis. Biosci Biotechnol Biochem. 2015;79:732–7.
Perry MM, Muntoni F. Noncoding RNAs and Duchenne muscular dystrophy. Epigenomics. 2016;8:1527.
Gagliardi S, Zucca S, Pandini C, Diamanti L, Bordoni M, Sproviero D, et al. Long non-coding and coding RNAs characterization in peripheral blood mononuclear cells and spinal cord from amyotrophic lateral sclerosis patients. Sci Rep. 2018;8:2378.
Ma J, Liu F, Du X, Ma D, Xiong L. Changes in lncRNAs and related genes in β-thalassemia minor and β-thalassemia major. Front Med. 2017;11:74–86.
Bayoumi AS, Aonuma T, Teoh JP, Tang YL, Kim IM. Circular noncoding RNAs as potential therapies and circulating biomarkers for cardiovascular diseases. Acta Pharmacol Sin. 2018;39(7):1100–9.
Ghelani HS, Rachchh MA, Gokani RH. MicroRNAs as newer therapeutic targets: a big hope from a tiny player. J Pharmacol Pharmacother. 2012;3:217–27.
Brown BD, Naldini L. Exploiting and antagonizing microRNA regulation for therapeutics and experimental applications. Nat Rev Genet. 2009;10:578–85.
Fabbri E, Brognara E, Borgatti M, Lampronti I, Finotti A, Bianchi N, et al. miRNA therapeutics: delivery and biological activity of peptide nucleic acids targeting miRNAs. Epigenomics. 2011;3:733–45.
Seto AG. The road toward microRNA therapeutics. Int J Biochem Cell Biol. 2010;42:1298–305.
Bader AG, Lammers P. The therapeutic potential of microRNAs. Innov Pharm Technol. 2011;52–5.
Van Rooij E, Kauppinen S. Development of microRNA therapeutics is coming of age. EMBO Mol Med. 2014;6:851–64.
Berindan-Neagoe I, Monroig Pdel C, Pasculli B, Calin GA. MicroRNAome genome: a treasure for cancer diagnosis and therapy. CA Cancer J Clin. 2014;64:311–36.
Czech MP. MicroRNAs as therapeutic targets. N Engl J Med. 2006;354:1194–5.
Weiler J, Hunziker J, Hall J. Anti-miRNA oligonucleotides (AMOs): ammunition to target miRNAs implicated in human disease? Gene Ther. 2006;13:496–502.
Lu Y, Xiao J, Lin H, Bai Y, Luo X, Wang Z, et al. A single antimicroRNA antisense oligodeoxyribonucleotide (AMO) targeting multiple microRNAs offers an improved approach for microRNA interference. Nucleic Acids Res. 2009;37:e24.
Lennox KA, Behlke MA. Chemical modification and design of antimiRNA oligonucleotides. Gene Ther. 2011;18:1111–20.
Obad S, dos Santos CO, Petri A, Heidenblad M, Broom O, Ruse C, et al. Silencing of microRNA families by seed-targeting tiny LNAs. Nat Genet. 2011;43:371–8.
Fabbri E, Borgatti M, Montagner G, Bianchi N, Finotti A, Lampronti I, et al. Modulation of the biological activity of microRNA-210 with peptide nucleic acids (PNAs). ChemMedChem. 2011;6:2192–202.
Brognara E, Fabbri E, Bazzoli E, Montagner G, Ghimenton C, Eccher A, et al. Uptake by human glioma cell lines and biological effects of a peptide-nucleic acids targeting miR-221. J Neurooncol. 2014;118:19–28.
Ebert MS, Neilson JR, Sharp PA. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods. 2007;4:721–6.
Ebert MS, Sharp PA. MicroRNA sponges: progress and possibilities. RNA. 2010;16:2043–50.
Kluiver J, Gibcus JH, Hettinga C, Adema A, Richter MK, Halsema N, et al. Rapid generation of microRNA sponges for microRNA inhibition. PLoS One. 2012;7:e29275.
Liu Y, Han Y, Zhang H, Nie L, Jiang Z, Fa P, et al. Synthetic miRNA-mowers targeting miR-183-96-182 cluster or miR-210 inhibit growth and migration and induce apoptosis in bladder cancer cells. PLoS One. 2012;7:e52280.
Wang Z. The principles of MiRNA-masking antisense oligonucleotides technology. Methods Mol Biol. 2011;676:43–9.
Murakami K, Miyagishi M. Tiny masking locked nucleic acids effectively bind to mRNA and inhibit binding of microRNAs in relation to thermodynamic stability. Biomed Rep. 2014;2:509–12.
Askou AL, Aagaard L, Kostic C, Arsenijevic Y, Hollensen AK, Bek T, et al. Multigenic lentiviral vectors for combined and tissue-specific expression of miRNA- and protein-based antiangiogenic factors. Mol Ther Methods Clin Dev. 2015;2:14064.
Montgomery RL, Yu G, Latimer PA, Stack C, Robinson K, Dalby CM, et al. MicroRNA mimicry blocks pulmonary fibrosis. EMBO Mol Med. 2014;6:1347–56.
Bader AG. miR-34-a microRNA replacement therapy is headed to the clinic. Front Genet. 2012;3:120.
Kwekkeboom RF, Lei Z, Doevendans PA, Musters RJ, Sluijter JP. Targeted delivery of miRNA therapeutics for cardiovascular diseases: opportunities and challenges. Clin Sci (Lond). 2014;127:351–65.
Cacchiarelli D, Legnini I, Martone J, Cazzella V, D’Amico A, Bertini E, et al. miRNAs as serum biomarkers for Duchenne muscular dystrophy. EMBO Mol Med. 2011;3:258–65.
Mizuno H, Nakamura A, Aoki Y, Ito N, Kishi S, Yamamoto K, et al. Identification of muscle-specific microRNAs in serum of muscular dystrophy animal models: promising novel blood-based markers for muscular dystrophy. PLoS One. 2011;6:e18388.
Naguibneva I, Ameyar-Zazoua M, Polesskaya A, Ait-Si-Ali S, Groisman R, Souidi M, et al. The microRNA miR-181 targets the homeobox protein Hox-A11 during mammalian myoblast differentiation. Nat Cell Biol. 2006;8:278–84.
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM, et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet. 2006;38:228–33.
van Rooij E, Sutherland LB, Thatcher JE, DiMaio JM, Naseem RH, Marshall WS, et al. Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc Natl Acad Sci USA. 2007;105:13027–32.
Eisenberg I, Eran A, Nishino I, Moggio M, Lamperti C, Amato AA, et al. Distinctive patterns of microRNA expression in primary muscular disorders. Proc Natl Acad Sci USA. 2007;104:17016–21.
Greco S, De Simone M, Colussi C, Zaccagnini G, Fasanaro P, Pescatori M, et al. Common micro-RNA signature in skeletal muscle damage and regeneration induced by Duchenne muscular dystrophy and acute ischemia. FASEB J. 2009;23:3335–46.
Yuasa K, Hagiwara Y, Ando M, Nakamura A, Takeda S, Hijikata T. MicroRNA-206 is highly expressed in newly formed muscle fibers: implications regarding potential for muscle regeneration and maturation in muscular dystrophy. Cell Struct Funct. 2008;33:163–9.
Wang L, Zhou L, Jiang P, Lu L, Chen X, Lan H, et al. Loss of miR-29 in myoblasts contributes to dystrophic muscle pathogenesis. Mol Ther. 2012;20:1222–33.
Gambardella S, Rinaldi F, Lepore SM, Viola A, Loro E, Angelini C, et al. Overexpression of microRNA-206 in the skeletal muscle from myotonic dystrophy type 1 patients. J Transl Med. 2010;8:48.
Fritegotto C, Ferrati C, Pegoraro V, Angelini C. Micro-RNA expression in muscle and fiber morphometry in myotonic dystrophy type 1. Neurol Sci. 2017;38:619–25.
Hervé M, Ibrahim EC. MicroRNA screening identifies a link between NOVA1 expression and a low level of IKAP in familial dysautonomia. Dis Model Mech. 2016;9:899–909.
Williams AH, Valdez G, Moresi V, Qi X, McAnally J, Elliott JL, et al. MicroRNA-206 delays ALS progression and promotes regeneration of neuromuscular synapses in mice. Science. 2009;326:1549–54.
Paco S, Casserras T, Rodríguez MA, Jou C, Puigdelloses M, Ortez CI, et al. Transcriptome analysis of Ullrich congenital muscular dystrophy fibroblasts reveals a disease extracellular matrix signature and key molecular regulators. PLoS One. 2015;10:e0145107.
Gillen AE, Gosalia N, Leir SH, Harris A. MicroRNA regulation of expression of the cystic fibrosis transmembrane conductance regulator gene. Biochem J. 2011;438:25–32.
Hassan F, Nuovo GJ, Crawford M, Boyaka PN, Kirkby S, Nana-Sinkam SP, et al. MiR-101 and miR-144 regulate the expression of the CFTR chloride channel in the lung. PLoS One. 2012;7:e50837.
Ramachandran S, Karp PH, Jiang P, Ostedgaard LS, Walz AE, Fisher JT, et al. A microRNA network regulates expression and biosynthesis of wild-type and DeltaF508 mutant cystic fibrosis transmembrane conductance regulator. Pro Natl Acad Sci USA. 2012;109:13362–7.
Ramachandran S, Karp PH, Osterhaus SR, Jiang P, Wohlford-Lenane C, Lennox KA, et al. Post-transcriptional regulation of cystic fibrosis transmembrane conductance regulator expression and function by microRNAs. Am J Respir Cell Mol Biol. 2013;49:544–51.
Oglesby IK, Chotirmall SH, McElvaney NG, Greene CM. Regulation of cystic fibrosis transmembrane conductance regulator by microRNA-145, -223, and -494 is altered in ΔF508 cystic fibrosis airway epithelium. J Immunol. 2013;190:3354–62.
Fabbri E, Tamanini A, Jakova T, Gasparello J, Manicardi A, Corradini R, et al. A peptide nucleic acid against microRNA miR-145-5p enhances the expression of the cystic fibrosis transmembrane conductance regulator (CFTR) in Calu-3 cells. Molecules. 2017;23:E71.
Pierdomenico AM, Patruno S, Codagnone M, Simiele F, Mari VC, Plebani R, et al. microRNA-181b is increased in cystic fibrosis cells and impairs lipoxin A4 receptor-dependent mechanisms of inflammation resolution and antimicrobial defense. Sci Rep. 2017;7:13519.
Sonneville F, Ruffin M, Coraux C, Rousselet N, Le Rouzic P, Blouquit-Laye S, et al. MicroRNA-9 downregulates the ANO1 chloride channel and contributes to cystic fibrosis lung pathology. Nat Commun. 2017;8:710.
Fabbri E, Borgatti M, Montagner G, Bianchi N, Finotti A, Lampronti I, et al. Expression of microRNA-93 and Interleukin-8 during Pseudomonas aeruginosa-mediated induction of proinflammatory responses. Am J Respir Cell Mol Biol. 2014;50:1144–55.
Urdinguio RG, Fernandez AF, Lopez-Nieva P, Rossi S, Huertas D, Kulis M, et al. Disrupted microRNA expression caused by Mecp2 loss in a mouse model of Rett syndrome. Epigenetics. 2010;5:656–63.
Kim JD, Lee A, Choi J, Park Y, Kang H, Chang W, et al. Epigenetic modulation as a therapeutic approach for pulmonary arterial hypertension. Exp Mol Med. 2015;47:e175.
Harafuji N, Schneiderat P, Walter MC, Chen YW. miR-411 is up-regulated in FSHD myoblasts and suppresses myogenic factors. Orphanet J Rare Dis. 2013;8:55.
Ballabio E, Mitchell T, van Kester MS, Taylor S, Dunlop HM, Chi J, et al. MicroRNA expression in Sezary syndrome: identification, function, and diagnostic potential. Blood. 2010;116:1105–13.
Narducci MG, Arcelli D, Picchio MC, Lazzeri C, Pagani E, Sampogna F, et al. MicroRNA profiling reveals that miR-21, miR486 and miR-214 are upregulated and involved in cell survival in Sézary syndrome. Cell Death Dis. 2011;2:e151.
van der Fits L, van Kester MS, Qin Y, Out-Luiting JJ, Smit F, Zoutman WH, et al. MicroRNA-21 expression in CD4 + T cells is regulated by STAT3 and is pathologically involved in Sézary syndrome. J Investig Dermatol. 2011;131:762–8.
Guibinga GH. MicroRNAs: tools of mechanistic insights and biological therapeutics discovery for the rare neurogenetic syndrome Lesch–Nyhan disease (LND). Adv Genet. 2015;90:103–31.
Zuntini M, Salvatore M, Pedrini E, Parra A, Sgariglia F, Magrelli A, et al. MicroRNA profiling of multiple osteochondromas: identification of disease-specific and normal cartilage signatures. Clin Genet. 2010;78:507–16.
Manca S, Magrelli A, Cialfi S, Lefort K, Ambra R, Alimandi M, et al. Oxidative stress activation of miR-125b is part of the molecular switch for Hailey–Hailey disease manifestation. Exp Dermatol. 2011;20:932–7.
Id Said B, Malkin D. A functional variant in miR-605 modifies the age of onset in Li-Fraumeni syndrome. Cancer Genet. 2015;208:47–51.
Magrelli A, Azzalin G, Salvatore M, Viganotti M, Tosto F, Colombo T, et al. Altered microRNA expression patterns in hepatoblastoma patients. Transl Oncol. 2009;2:157–63.
Meseguer S, Martínez-Zamora A, García-Arumí E, Andreu AL, Armengod ME. The ROS-sensitive microRNA-9/9* controls the expression of mitochondrial tRNA-modifying enzymes and is involved in the molecular mechanism of MELAS syndrome. Hum Mol Genet. 2015;24:167–84.
Feng J, Sun G, Yan J, Noltner K, Li W, Buzin CH, et al. Evidence for X-chromosomal schizophrenia associated with microRNA alterations. PLoS One. 2009;4:e6121.
Gasparello J, Fabbri E, Bianchi N, Breveglieri G, Zuccato C, Borgatti M, et al. BCL11A mRNA targeting by miR-210: a possible network regulating γ-globin gene expression. Int J Mol Sci. 2017;18:E2530.
Siwaponanan P, Fucharoen S, Sirankapracha P, Winichagoon P, Umemura T, Svasti S. Elevated levels of miR-210 correlate with anemia in β-thalassemia/HbE patients. Int J Hematol. 2016;104:338–43.
Srinoun K, Nopparatana C, Wongchanchailert M, Fucharoen S. MiR-155 enhances phagocytic activity of β-thalassemia/HbE monocytes via targeting of BACH1. Int J Hematol. 2017;106:638–47.
Leecharoenkiat K, Tanaka Y, Harada Y, Chaichompoo P, Sarakul O, Abe Y, et al. Plasma microRNA-451 as a novel hemolytic marker for β0-thalassemia/HbE disease. Mol Med Rep. 2017;15:2495–502.
Lulli V, Romania P, Morsilli O, Cianciulli P, Gabbianelli M, Testa U, et al. MicroRNA-486-3p regulates γ-globin expression in human erythroid cells by directly modulating BCL11A. PLoS One. 2013;8:e60436.
Saki N, Abroun S, Soleimani M, Kavianpour M, Shahjahani M, Mohammadi-Asl J, et al. MicroRNA expression in β-thalassemia and sickle cell disease: a role in the induction of fetal hemoglobin. Cell J. 2016;17:583–92.
Roy P, Bhattacharya G, Lahiri A, Dasgupta UB, Banerjee D, Chandra S, et al. hsa-miR-503 is downregulated in β thalassemia major. Acta Haematol. 2012;128:187–9.
Ballarino M, Cazzella V, D’Andrea D, Grassi L, Bisceglie L, Cipriano A, et al. Novel long noncoding RNAs (lncRNAs) in myogenesis: a miR-31 overlapping lncRNA transcript controls myoblast differentiation. Mol Cell Biol. 2015;35:728–36.
Twayana S, Legnini I, Cesana M, Cacchiarelli D, Morlando M, Bozzoni I. Biogenesis and function of non-coding RNAs in muscle differentiation and in Duchenne muscular dystrophy. Biochem Soc Trans. 2013;41:844–9.
Wheeler TM, Leger AJ, Pandey SK, MacLeod AR, Nakamori M, Cheng SH, et al. Targeting nuclear RNA for in vivo correction of myotonic dystrophy. Nature. 2012;488(7409):111–5.
Nishimoto Y, Nakagawa S, Hirose T, Okano HJ, Takao M, Shibata S, et al. The long non-coding RNA nuclear-enriched abundant transcript 1_2 induces paraspeckle formation in the motor neuron during the early phase of amyotrophic lateral sclerosis. Mol Brain. 2013;6:31.
McKiernan PJ, Molloy K, Cryan SA, McElvaney NG, Greene CM. Long noncoding RNA are aberrantly expressed in vivo in the cystic fibrosis bronchial epithelium. Int J Biochem Cell Biol. 2014;52:184–91.
Balloy V, Koshy R, Perra L, Corvol H, Chignard M, Guillot L, et al. Bronchial epithelial cells from cystic fibrosis patients express a specific long non-coding RNA signature upon Pseudomonas aeruginosa infection. Front Cell Infect Microbiol. 2017;7:218.
Saayman SM, Ackley A, Burdach J, Clemson M, Gruenert DC, Tachikawa K, et al. Long non-coding RNA BGas regulates the cystic fibrosis transmembrane conductance regulator. Mol Ther. 2016;24(8):1351–7.
Petazzi P, Sandoval J, Szczesna K, Jorge OC, Roa L, Sayols S, et al. G Dysregulation of the long non-coding RNA transcriptome in a Rett syndrome mouse model. RNA Biol. 2013;10:1197–203.
Sun Z, Nie X, Sun S, Dong S, Yuan C, Li Y, Xiao B, et al. Long non-coding RNA MEG3 downregulation triggers human pulmonary artery smooth muscle cell proliferation and migration via the p53 signaling pathway. Cell Physiol Biochem. 2017;42(6):2569–81.
Chen J, Guo J, Cui X, Dai Y, Tang Z, Qu J, Raj JU, et al. The long noncoding RNA LnRPT is regulated by PDGF-BB and modulates the proliferation of pulmonary artery smooth muscle cells. Am J Respir Cell Mol Biol. 2018;58(2):181–93.
Vizoso M, Esteller M. The activatory long non-coding RNA DBE-T reveals the epigenetic etiology of facioscapulohumeral muscular dystrophy. Cell Res. 2012;22(10):1413–5.
Wang W, Zhuang Q, Ji K, Wen B, Lin P, Zhao Y, et al. Identification of miRNA, lncRNA and mRNA-associated ceRNA networks and potential biomarker for MELAS with mitochondrial DNA A3243G mutation. Sci Rep. 2017;7:41639.
Morrison TA, Wilcox I, Luo HY, Farrell JJ, Kurita R, Nakamura Y, et al. A long noncoding RNA from the HBS1L-MYB intergenic region on chr6q23 regulates human fetal hemoglobin expression. Blood Cells Mol Dis. 2018;69:1–9.
Lai K, Jia S, Yu S, Luo J, He Y. Genome-wide analysis of aberrantly expressed lncRNAs and miRNAs with associated co-expression and ceRNA networks in β-thalassemia and hereditary persistence of fetal hemoglobin. Oncotarget. 2017;8:49931–43.
Falzarano MS, Scotton C, Passarelli C, Ferlini A. Duchenne muscular dystrophy: from diagnosis to therapy. Molecules. 2015;20:18168–84.
Li X, Li Y, Zhao L, Zhang D, Yao X, Zhang H, et al. Circulating muscle-specific miRNAs in Duchenne muscular dystrophy patients. Mol Ther Nucleic Acids. 2014;3:e177.
Hildyard JC, Wells DJ. Investigating synthetic oligonucleotide targeting of Mir31 in Duchenne muscular dystrophy. PLoS Curr 2016;8.
Mishra MK, Loro E, Sengupta K, Wilton SD, Khurana TS. Functional improvement of dystrophic muscle by repression of utrophin: let-7c interaction. PLoS One. 2017;12:e0182676.
Zanotti S, Gibertini S, Curcio M, Savadori P, Pasanisi B, Morandi L, et al. Opposing roles of miR-21 and miR-29 in the progression of fibrosis in Duchenne muscular dystrophy. Biochim Biophys Acta. 2015;1852:1451–64.
Bovolenta M, Erriquez D, Valli E, Brioschi S, Scotton C, Neri M, et al. The DMD locus harbours multiple long non-coding RNAs which orchestrate and control transcription of muscle dystrophin mRNA isoforms. PLoS One. 2012;7:e45328.
Kiernan MC, Vucic S, Cheah BC, Turner MR, Eisen A, Hardiman O, et al. Amyotrophic lateral sclerosis. Lancet. 2011;377:942–55.
Gagliardi S, Milani P, Sardone V, Pansarasa O, Cereda C. From transcriptome to noncoding RNAs: implications in ALS mechanism. Neurol Res Int. 2012;2012:278725.
Elborn JS. Cystic fibrosis. Lancet. 2016;388:2519–31.
Randell SH, Boucher RC, University of North Carolina Virtual Lung Group. Effective mucus clearance is essential for respiratory health. Am J Respir Cell Mol Biol. 2006;35:20–8.
Wang S, Yue H, Derin RB, Guggino WB, Li M. Accessory protein facilitated CFTR-CFTR interaction, a molecular mechanism to potentiate the chloride channel activity. Cell. 2000;103:169–79.
Peters KW, Okiyoneda T, Balch WE, Braakman I, Brodsky JL, Guggino WB, et al. CFTR Folding Consortium: methods available for studies of CFTR folding and correction. Methods Mol Biol. 2011;742:335–53.
Moyer BD, Denton J, Karlson KH, Reynolds D, Wang S, Mickle JE, et al. A PDZ-interacting domain in CFTR is an apical membrane polarization signal. J Clin Investig. 1999;104:1353–61.
Wang W, Xiao X, Chen X, Huo Y, Xi WJ, Lin ZF, et al. Tumor-suppressive miR-145 co-repressed by TCF4-β-catenin and PRC2 complexes forms double-negative regulation loops with its negative regulators in colorectal cancer. Int J Cancer. 2017;142:308–21.
Paul T, Li S, Khurana S, Leleiko NS, Walsh MJ. The epigenetic signature of CFTR expression is co-ordinated via chromatin acetylation through a complex intronic element. Biochem J. 2007;408:317–426.
Viart V, Bergougnoux A, Bonini J, Varilh J, Chiron R, Tabary O, et al. Transcription factors and miRNAs that regulate fetal to adult CFTR expression change are new targets for cystic fibrosis. Eur Respir J. 2015;45:116–28.
Vuillaumier S, Dixmeras I, Messai H, Lapoumeroulie C, Lallemand D, Gekas J, et al. Cross-species characterization of the promoter region of the cystic fibrosis transmembrane conductance regulator gene reveals multiple levels of regulation. Biochem J. 1997;327:651–62.
Pittman N, Shue G, LeLeiko NS, Walsh MJ. Transcription of cystic fibrosis transmembrane conductance regulator requires a CCAAT-like element for both basal and cAMP-mediated regulation. J Biol Chem. 1995;270:28848–57.
Zhu X, Li Y, Xie C, Yin X, Liu Y, Cao Y, et al. miR-145 sensitizes ovarian cancer cells to paclitaxel by targeting Sp1 and Cdk6. Int J Cancer. 2014;135:1286–96.
Xiang X, Zhuang X, Ju S, Zhang S, Jiang H, Mu J, et al. miR-155 promotes macroscopic tumor formation yet inhibits tumor dissemination from mammary fat pads to the lung by preventing EMT. Oncogene. 2011;30:3440–53.
Montanini L, Smerieri A, Gullì M, Cirillo F, Pisi G, Sartori C, et al. miR-146a, miR-155, miR-370, and miR-708 are CFTR-dependent, predicted FOXO1 regulators and change at onset of CFRDs. J Clin Endocrinol Metab. 2016;101:4955–63.
Amato F, Seia M, Giordano S, Elce A, Zarrilli F, Castaldo G, et al. Gene mutation in microRNA target sites of CFTR gene: a novel pathogenetic mechanism in cystic fibrosis? PLoS One. 2013;8:e60448.
Megiorni F, Cialfi S, Dominici C, Quattrucci S, Pizzuti A. Synergistic post-transcriptional regulation of the cystic fibrosis transmembrane conductance regulator (CFTR) by miR-101 and miR-494 specific binding. PLoS One. 2011;6:e26601.
Lyst MJ, Bird A. Rett syndrome: a complex disorder with simple roots. Nat Rev Genet. 2015;16:261–75.
Tsujimura K, Irie K, Nakashima H, Egashira Y, Fukao Y, Fujiwara M, et al. miR-199a links MeCP2 with mTOR signaling and its dysregulation leads to Rett syndrome phenotypes. Cell Rep. 2015;12:1887–901.
Wilcox RA. Cutaneous T-cell lymphoma: 2016 update on diagnosis, risk-stratification, and management. Am J Hematol. 2016;91:151–65.
Benoit BM, Jariwala N, O’Connor G, Oetjen LK, Whelan TM, Werth A, et al. CD164 identifies CD4 + T cells highly expressing genes associated with malignancy in Sézary syndrome: the Sézary signature genes, FCRL3, Tox, and miR-214. Arch Dermatol Res. 2017;309:11–9.
Lee CS, Ungewickell A, Bhaduri A, Qu K, Webster DE, Armstrong R, et al. Transcriptome sequencing in Sezary syndrome identifies Sezary cell and mycosis fungoides-associated lncRNAs and novel transcripts. Blood. 2012;120:3288–97.
Thein SL. Molecular basis of β thalassemia and potential therapeutic targets. Blood Cells Mol Dis. 2017;S1079–9796:30210–3.
de Dreuzy E, Bhukhai K, Leboulch P, Payen E. Current and future alternative therapies for beta-thalassemia major. Biomed J. 2016;39:24–38.
Gambari R. Foetal haemoglobin inducers and thalassaemia: novel achievements. Blood Transfus. 2010;8:5–7.
Finotti A, Breda L, Lederer CW, Bianchi N, Zuccato C, Kleanthous M, Rivella S, Gambari R. Recent trends in the gene therapy of β-thalassemia. J Blood Med. 2015;6:69–85.
Breveglieri G, Bianchi N, Cosenza LC, Gamberini MR, Chiavilli F, Zuccato C, et al. An Aγ-globin G- > A gene polymorphism associated with β039 thalassemia globin gene and high fetal hemoglobin production. BMC Med Genet. 2017;18:93.
Bianchi N, Cosenza LC, Lampronti I, Finotti A, Breveglieri G, Zuccato C, et al. Structural and functional insights on an uncharacterized Aγ-globin-gene polymorphism present in four β0-thalassemia families with high fetal hemoglobin levels. Mol Diagn Ther. 2016;20:161–73.
Bianchi N, Zuccato C, Finotti A, Lampronti I, Borgatti M, Gambari R. Involvement of miRNA in erythroid differentiation. Epigenomics. 2012;4:51–65.
Bianchi N, Finotti A, Ferracin M, Lampronti I, Zuccato C, Breveglieri G, et al. Increase of microRNA-210, decrease of raptor gene expression and alteration of mammalian target of rapamycin regulated proteins following mithramycin treatment of human erythroid cells. PLoS One. 2015;10:e0121567.
Sarakul O, Vattanaviboon P, Tanaka Y, Fucharoen S, Abe Y, Svasti S, et al. Enhanced erythroid cell differentiation in hypoxic condition is in part contributed by miR-210. Blood Cells Mol Dis. 2013;51:98–103.
Bavelloni A, Poli A, Fiume R, Blalock W, Matteucci A, Ramazzotti G, et al. PLC-beta 1 regulates the expression of miR-210 during mithramycin-mediated erythroid differentiation in K562 cells. Oncotarget. 2014;5:4222–31.
Sawant M, Colah R, Ghosh K, Nadkarni A. Does HbF induction by hydroxycarbamide work through MIR210 in sickle cell anaemia patients? Br J Haematol. 2016;173:801–3.
Li Y, Liu D, Zhang X, Li Z, Ye Y, Liu Q, et al. miR-326 regulates HbF synthesis by targeting EKLF in human erythroid cells. Exp Hematol. 2018;63:33–40.
De Antonellis P, Carotenuto M, Vandenbussche J, De Vita G, Ferrucci V, Medaglia C, et al. Early targets of miR-34a in neuroblastoma. Mol Cell Proteom. 2014;13:2114–31.
Sankaran VG, Menne TF, Scepanovic D, Vergilio JA, Ji P, Kim J, et al. MicroRNA-15a and -16-1 act via MYB to elevate fetal hemoglobin expression in human trisomy 13. Proc Natl Acad Sci USA. 2011;108:1519–24.
Pule GD, Mowla S, Novitzky N, Wonkam A. Hydroxyurea down-regulates BCL11A, KLF-1 and MYB through miRNA-mediated actions to induce γ-globin expression: implications for new therapeutic approaches of sickle cell disease. Clin Transl Med. 2016;5:15.
Ma Y, Wang B, Jiang F, Wang D, Liu H, Yan Y, Dong H, Wang F, Gong B, Zhu Y, Dong L, Yin H, Zhang Z, Zhao H, Wu Z, Zhang J, Zhou J, Yu J. A feedback loop consisting of microRNA 23a/27a and the β-like globin suppressors KLF3 and SP1 regulates globin gene expression. Mol Cell Biol. 2013;33:3994–4007.
Azzouzi I, Moest H, Winkler J, Fauchère JC, Gerber AP, Wollscheid B, Stoffel M, Schmugge M, Speer O. MicroRNA-96 directly inhibits γ-globin expression in human erythropoiesis. PLoS One. 2011;6(7):e22838.
Gu S, Jin L, Zhang F, Sarnow P, Kay MA. Biological basis for restriction of microRNA targets to the 3′ untranslated region in mammalian mRNAs. Nat Struct Mol Biol. 2009;16:144–50.
Stark A, Brennecke J, Bushati N, Russell RB, Cohen SM. Animal MicroRNAs confer robustness to gene expression and have a significant impact on 3′UTR evolution. Cell. 2005;123:1133–46.
Zhang R, Su B. Small but influential: the role of microRNAs on gene regulatory network and 3′ UTR evolution. J Genet Genom. 2009;36:1–6.
Duursma AM, Kedde M, Schrier M, le Sage C, Agami R. miR-148 targets human DNMT3b protein coding region. RNA. 2008;14:872–7.
Brümmer A, Hausser J. MicroRNA binding sites in the coding region of mRNAs: extending the repertoire of post-transcriptional gene regulation. Bioessays. 2014;36:617–26.
Hausser J, Syed AP, Bilen B, Zavolan M. Analysis of CDS-located miRNA target sites suggests that they can effectively inhibit translation. Genome Res. 2013;23:604–15.
Meekings KN, Williams CS, Arrowsmith JE. Orphan drug development: an economically viable strategy for biopharma R&D. Drug Discov Today. 2012;17:660–4.
Halffner ME, Whitley J, Moses M. Two decades of orphan product development. Nat Rev Drug Discov. 2002;1:821–5.
Fagnan DE, Gromatzky AA, Stein RM, Fernandez JM, Lo AW. Financing drug discovery for orphan diseases. Drug Discov Today. 2014;19:533–8.
Uguen D, Lönngren T, Le Cam Y, Garner S, Voisin E, Incerti C, et al. Accelerating development, registration and access to medicines for rare diseases in the European Union through adaptive approaches: features and perspectives. Orphanet J Rare Dis. 2014;9:20.
Michel M, Toumi M. Access to orphan drugs in Europe: current and future issues. Expert Rev Pharmacoecon Outcomes Res. 2012;12:23–9.
Committee for Orphan Medicinal Products and the European Medicines, Westermark K, Holm BB, Söderholm M, Llinares-Garcia J, Rivière F, et al. European regulation on orphan medicinal products: 10 years of experience and future perspectives. Nat Rev Drug Discov. 2011;10:341–9.
Hall AK, Carlson MR. The current status of orphan drug development in Europe and the US. Intractable Rare Dis Res. 2014;3:1–7.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors (AF, EF, IL, JG, MB and RG) declare they have no conflicts of interest that are directly relevant to the content of this review.
Funding
This study was partially supported by the following funds to RG: EU FP7 THALAMOSS Project (THALAssaemia MOdular Stratification System for personalized therapy of β-thalassemia; Grant number [306201]-FP7-Health-2012-INNOVATION-1), Fondazione CARIPARO and Associazione Veneta per la Lotta alla Talassemia (AVLT) Rovigo. In addition, this study was supported by Fondazione Fibrosi Cistica (FFC), Project “Revealing the microRNAs-Transcription factors network in cystic fibrosis: from microRNA therapeutics to precision medicine” (CF-miRNA-THER, FFC#7/2018). This study is also partially supported by Wellcome Trust and by Agenzia Italiana del Farmaco (AIFA). All funding bodies had no role in the design of the study, in the collection, analysis, and interpretation of data, or in writing the manuscript.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution-NonCommercial 4.0 International License (http://creativecommons.org/licenses/by-nc/4.0/), which permits any noncommercial use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Finotti, A., Fabbri, E., Lampronti, I. et al. MicroRNAs and Long Non-coding RNAs in Genetic Diseases. Mol Diagn Ther 23, 155–171 (2019). https://doi.org/10.1007/s40291-018-0380-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-018-0380-6