Abstract
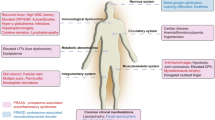
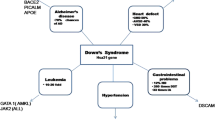
Shwachman-Diamond syndrome (SDS) is a rare inherited disease mainly caused by mutations in the Shwachman-Bodian-Diamond Syndrome (SBDS) gene. However, it has recently been reported that other genes, including DnaJ heat shock protein family (Hsp40) member C21 (DNAJC21), elongation factor-like 1 (EFL1) and signal recognition particle 54 (SRP54) are also associated with an SDS-like phenotype. Interestingly, SBDS, DNAJC21, EFL1 and SRP54 are involved in ribosome biogenesis: SBDS, through direct interaction with EFL1, promotes the release of the eukaryotic initiation factor 6 (eIF6) during ribosome maturation, DNAJC21 stabilizes the 80S ribosome, and SRP54 facilitates protein trafficking. These findings strengthen the postulate that SDS is a ribosomopathy. SDS is a multiple-organ disease mainly characterized by bone marrow failure, bone malformations, pancreatic insufficiency and cognitive disorders. Almost 15–20% of patients with SDS present myelodysplastic syndrome with a high risk of acute myeloid leukemia (AML) transformation. Unfortunately, besides bone marrow transplantation, no gene-based therapy for SDS has yet been developed. This review aims to recapitulate the recent findings on the molecular mechanisms of SDS underlying bone marrow failure, hematopoiesis and AML development and to draw a realistic picture of current perspectives.


Similar content being viewed by others
References
Boocock GR, Morrison JA, Popovic M, Richards N, Ellis L, Durie PR, et al. Mutations in SBDS are associated with Shwachman-Diamond syndrome. Nat Genet. 2003;33(1):97–101.
Cipolli M. Shwachman-Diamond syndrome: clinical phenotypes. Pancreatology. 2001;1(5):543–8.
Ginzberg H, Shin J, Ellis L, Morrison J, Ip W, Dror Y, et al. Shwachman syndrome: phenotypic manifestations of sibling sets and isolated cases in a large patient cohort are similar. J Pediatr. 1999;135(1):81–8.
Dror Y, Durie P, Ginzberg H, Herman R, Banerjee A, Champagne M, et al. Clonal evolution in marrows of patients with Shwachman-Diamond syndrome: a prospective 5-year follow-up study. Exp Hematol. 2002;30(7):659–69.
Maserati E, Minelli A, Pressato B, Valli R, Crescenzi B, Stefanelli M, et al. Shwachman syndrome as mutator phenotype responsible for myeloid dysplasia/neoplasia through karyotype instability and chromosomes 7 and 20 anomalies. Genes Chromosomes Cancer. 2006;45(4):375–82.
Mäkitie O, Ellis L, Durie PR, Morrison JA, Sochett EB, Rommens JM, et al. Skeletal phenotype in patients with Shwachman-Diamond syndrome and mutations in SBDS. Clin Genet. 2004;65(2):101–12.
Toiviainen-Salo S, Mäyränpää MK, Durie PR, Richards N, Grynpas M, Ellis L, et al. Shwachman-Diamond syndrome is associated with low-turnover osteoporosis. Bone. 2007;41(6):965–72.
Kerr EN, Ellis L, Dupuis A, Rommens JM, Durie PR. The behavioral phenotype of school-age children with shwachman diamond syndrome indicates neurocognitive dysfunction with loss of Shwachman-Bodian-Diamond syndrome gene function. J Pediatr. 2010;156(3):433–8.
Perobelli S, Nicolis E, Assael BM, Cipolli M. Further characterization of Shwachman-Diamond syndrome: psychological functioning and quality of life in adult and young patients. Am J Med Genet A. 2012;158A(3):567–73.
Perobelli S, Alessandrini F, Zoccatelli G, Nicolis E, Beltramello A, Assael BM, et al. Diffuse alterations in grey and white matter associated with cognitive impairment in Shwachman-Diamond syndrome: evidence from a multimodal approach. Neuroimage Clin. 2015;7:721–31.
Bezzerri V, Bardelli D, Morini J, Vella A, Cesaro S, Sorio C, et al. Ataluren-driven restoration of shwachman-bodian-diamond syndrome protein function in shwachman-diamond syndrome bone marrow cells. Am J Hematol. 2018;93:527–36.
Zhang S, Shi M, Hui CC, Rommens JM. Loss of the mouse ortholog of the shwachman-diamond syndrome gene (Sbds) results in early embryonic lethality. Mol Cell Biol. 2006;26(17):6656–63.
Kuijpers TW, Alders M, Tool AT, Mellink C, Roos D, Hennekam RC. Hematologic abnormalities in Shwachman Diamond syndrome: lack of genotype-phenotype relationship. Blood. 2005;106(1):356–61.
Burwick N, Shimamura A, Liu JM. Non-Diamond Blackfan anemia disorders of ribosome function: Shwachman Diamond syndrome and 5q-syndrome. Semin Hematol. 2011;48(2):136–43.
Finch AJ, Hilcenko C, Basse N, Drynan LF, Goyenechea B, Menne TF, et al. Uncoupling of GTP hydrolysis from eIF6 release on the ribosome causes Shwachman-Diamond syndrome. Genes Dev. 2011;25(9):917–29.
Weis F, Giudice E, Churcher M, Jin L, Hilcenko C, Wong CC, et al. Mechanism of eIF6 release from the nascent 60S ribosomal subunit. Nat Struct Mol Biol. 2015;22(11):914–9.
Austin KM, Gupta ML, Coats SA, Tulpule A, Mostoslavsky G, Balazs AB, et al. Mitotic spindle destabilization and genomic instability in Shwachman-Diamond syndrome. J Clin Invest. 2008;118(4):1511–8.
Orelio C, Verkuijlen P, Geissler J, van den Berg TK, Kuijpers TW. SBDS expression and localization at the mitotic spindle in human myeloid progenitors. PLoS One. 2009;4(9):e7084.
Dror Y, Freedman MH. Shwachman-Diamond syndrome marrow cells show abnormally increased apoptosis mediated through the Fas pathway. Blood. 2001;97(10):3011–6.
Watanabe K, Ambekar C, Wang H, Ciccolini A, Schimmer AD, Dror Y. SBDS-deficiency results in specific hypersensitivity to Fas stimulation and accumulation of Fas at the plasma membrane. Apoptosis. 2009;14(1):77–89.
Morini J, Babini G, Mariotti L, Baiocco G, Nacci L, Maccario C, et al. Radiosensitivity in lymphoblastoid cell lines derived from Shwachman-Diamond syndrome patients. Radiat Prot Dosimetry. 2015;166(1–4):95–100.
Tourlakis ME, Zhang S, Ball HL, Gandhi R, Liu H, Zhong J, et al. In vivo senescence in the sbds-deficient murine pancreas: cell-type specific consequences of translation insufficiency. PLoS Genet. 2015;11(6):e1005288.
Schaballie H, Renard M, Vermylen C, Scheers I, Revencu N, Regal L, et al. Misdiagnosis as asphyxiating thoracic dystrophy and CMV-associated haemophagocytic lymphohistiocytosis in Shwachman-Diamond syndrome. Eur J Pediatr. 2013;172(5):613–22.
Keogh SJ, McKee S, Smithson SF, Grier D, Steward CG. Shwachman-Diamond syndrome: a complex case demonstrating the potential for misdiagnosis as asphyxiating thoracic dystrophy (Jeune syndrome). BMC Pediatr. 2012;12:48.
Qiu XB, Shao YM, Miao S, Wang L. The diversity of the DnaJ/Hsp40 family, the crucial partners for Hsp70 chaperones. Cell Mol Life Sci. 2006;63(22):2560–70.
Young JC, Agashe VR, Siegers K, Hartl FU. Pathways of chaperone-mediated protein folding in the cytosol. Nat Rev Mol Cell Biol. 2004;5(10):781–91.
Hennessy F, Nicoll WS, Zimmermann R, Cheetham ME, Blatch GL. Not all J domains are created equal: implications for the specificity of Hsp40-Hsp70 interactions. Protein Sci. 2005;14(7):1697–709.
Lo KY, Li Z, Bussiere C, Bresson S, Marcotte EM, Johnson AW. Defining the pathway of cytoplasmic maturation of the 60S ribosomal subunit. Mol Cell. 2010;39(2):196–208.
Tummala H, Walne AJ, Williams M, Bockett N, Collopy L, Cardoso S, et al. DNAJC21 mutations link a cancer-prone bone marrow failure syndrome to corruption in 60s ribosome subunit maturation. Am J Hum Genet. 2016;99(1):115–24.
Dhanraj S, Matveev A, Li H, Lauhasurayotin S, Jardine L, Cada M, et al. Biallelic mutations in DNAJC21 cause Shwachman-Diamond syndrome. Blood. 2017;129:1557–62.
D’Amours G, Lopes F, Gauthier J, Saillour V, Nassif C, Wynn R, et al. Refining the phenotype associated with biallelic DNAJC21 mutations. Clin Genet. 2018;94(2):252–8.
Morini J, Nacci L, Babini G, Cesaro S, Valli R, Ottolenghi A, et al. Whole exome sequencing discloses heterozygous variants in the DNAJC21 and EFL1 genes but not in SRP54 in 6 out of 16 patients with Shwachman-Diamond Syndrome carrying biallelic SBDS mutations. Br J Haematol. 2018. https://doi.org/10.1111/bjh.15594.
Stepensky P, Chacón-Flores M, Kim KH, Abuzaitoun O, Bautista-Santos A, Simanovsky N, et al. Mutations in EFL1, an SBDS partner, are associated with infantile pancytopenia, exocrine pancreatic insufficiency and skeletal anomalies in aShwachman-Diamond like syndrome. J Med Genet. 2017;54(8):558–66.
Tan QK, Cope H, Spillmann RC, Stong N, Jiang YH, McDonald MT, et al. Further evidence for the involvement of EFL1 in a Shwachman-Diamond-like syndrome and expansion of the phenotypic features. Cold Spring Harb Mol Case Stud. 2018. https://doi.org/10.1101/mcs.a003046.
Carapito R, Konantz M, Paillard C, Miao Z, Pichot A, Leduc MS, et al. Mutations in signal recognition particle SRP54 cause syndromic neutropenia with Shwachman-Diamond-like features. J Clin Invest. 2017;127(11):4090–103.
Keenan RJ, Freymann DM, Stroud RM, Walter P. The signal recognition particle. Annu Rev Biochem. 2001;70:755–75.
Khincha PP, Savage SA. Genomic characterization of the inherited bone marrow failure syndromes. Semin Hematol. 2013;50(4):333–47.
Dror Y, Freedman MH. Shwachman-Diamond syndrome: an inherited preleukemic bone marrow failure disorder with aberrant hematopoietic progenitors and faulty marrow microenvironment. Blood. 1999;94(9):3048–54.
Mercuri A, Cannata E, Perbellini O, Cugno C, Balter R, Zaccaron A, et al. Immunophenotypic analysis of hematopoiesis in patients suffering from Shwachman-Bodian-Diamond Syndrome. Eur J Haematol. 2015;95(4):308–15.
Blair A, Sutherland HJ. Primitive acute myeloid leukemia cells with long-term proliferative ability in vitro and in vivo lack surface expression of c-kit (CD117). Exp Hematol. 2000;28(6):660–71.
Heinrich MC, Dooley DC, Keeble WW. Transforming growth factor beta 1 inhibits expression of the gene products for steel factor and its receptor (c-kit). Blood. 1995;85(7):1769–80.
Pierelli L, Scambia G, Bonanno G, Rutella S, Puggioni P, Battaglia A, et al. CD34 +/CD105 + cells are enriched in primitive circulating progenitors residing in the G0 phase of the cell cycle and contain all bone marrow and cord blood CD34 +/CD38low/- precursors. Br J Haematol. 2000;108(3):610–20.
Hudson E, Aldor T. Pancreatic insufficiency and neutropenia with associated immunoglobulin deficit. Arch Intern Med. 1970;125(2):314–6.
Aggett PJ, Harries JT, Harvey BA, Soothill JF. An inherited defect of neutrophil mobility in Shwachman syndrome. J Pediatr. 1979;94(3):391–4.
Sacchi F, Maggiore G, Marseglia G, Marconi M, Nespoli L, Siccardi AG. Association of neutrophil and complement defects in two twins with Shwachman syndrome. Helv Paediatr Acta. 1982;37(2):177–81.
Repo H, Savilahti E, Leirisalo-Repo M. Aberrant phagocyte function in Shwachman syndrome. Clin Exp Immunol. 1987;69(1):204–12.
Dror Y, Ginzberg H, Dalal I, Cherepanov V, Downey G, Durie P, et al. Immune function in patients with Shwachman-Diamond syndrome. Br J Haematol. 2001;114(3):712–7.
Bannon SA, DiNardo CD. Hereditary predispositions to myelodysplastic syndrome. Int J Mol Sci. 2016. https://doi.org/10.3390/ijms17060838.
Swerdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R, et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood. 2016 05;127(20):2375–90.
Myers KC, Davies SM, Shimamura A. Clinical and molecular pathophysiology of Shwachman-Diamond syndrome: an update. Hematol Oncol Clin North Am. 2013;27(1):117–28, ix.
Valli R, Pressato B, Marletta C, Mare L, Montalbano G, Curto FL, et al. Different loss of material in recurrent chromosome 20 interstitial deletions in Shwachman-Diamond syndrome and in myeloid neoplasms. Mol Cytogenet. 2013;6(1):56.
Liu JM. A clinical algorithm predicts hematological complications in Shwachman-Diamond syndrome? Expert Rev Hematol. 2012;5(4):373–5.
Pressato B, Valli R, Marletta C, Mare L, Montalbano G, Lo Curto F, et al. Deletion of chromosome 20 in bone marrow of patients with Shwachman-Diamond syndrome, loss of the EIF6 gene and benign prognosis. Br J Haematol. 2012;157(4):503–5.
Bezzerri V, Vella A, Calcaterra E, Finotti A, Gasparello J, Gambari R, et al. New insights into the Shwachman-Diamond Syndrome-related haematological disorder: hyper-activation of mTOR and STAT3 in leukocytes. Sci Rep. 2016;6:33165.
Ravera S, Dufour C, Cesaro S, Bottega R, Faleschini M, Cuccarolo P, et al. Evaluation of energy metabolism and calcium homeostasis in cells affected by Shwachman-Diamond syndrome. Sci Rep. 2016;6:25441.
Chapuis N, Tamburini J, Green AS, Willems L, Bardet V, Park S, et al. Perspectives on inhibiting mTOR as a future treatment strategy for hematological malignancies. Leukemia. 2010;24(10):1686–99.
Ma L, Teruya-Feldstein J, Behrendt N, Chen Z, Noda T, Hino O, et al. Genetic analysis of Pten and Tsc2 functional interactions in the mouse reveals asymmetrical haploinsufficiency in tumor suppression. Genes Dev. 2005;19(15):1779–86.
Hoshii T, Matsuda S, Hirao A. Pleiotropic roles of mTOR complexes in haemato-lymphopoiesis and leukemogenesis. J Biochem. 2014;156(2):73–83.
Kusaba H, Ghosh P, Derin R, Buchholz M, Sasaki C, Madara K, et al. Interleukin-12-induced interferon-gamma production by human peripheral blood T cells is regulated by mammalian target of rapamycin (mTOR). J Biol Chem. 2005;280(2):1037–43.
Mitchell TJ, John S. Signal transducer and activator of transcription (STAT) signalling and T-cell lymphomas. Immunology. 2005;114(3):301–12.
Coffer PJ, Koenderman L, de Groot RP. The role of STATs in myeloid differentiation and leukemia. Oncogene. 2000;19(21):2511–22.
Calò V, Migliavacca M, Bazan V, Macaluso M, Buscemi M, Gebbia N, et al. STAT proteins: from normal control of cellular events to tumorigenesis. J Cell Physiol. 2003;197(2):157–68.
Yu H, Liu X, Huang J, Zhang Y, Hu R, Pu J. Comparison of read-through effects of aminoglycosides and PTC124 on rescuing nonsense mutations of HERG gene associated with long QT syndrome. Int J Mol Med. 2014;33(3):729–35.
O’Shea JJ, Holland SM, Staudt LM. JAKs and STATs in immunity, immunodeficiency, and cancer. N Engl J Med. 2013;368(2):161–70.
Tourlakis ME, Zhong J, Gandhi R, Zhang S, Chen L, Durie PR, et al. Deficiency of Sbds in the mouse pancreas leads to features of Shwachman-Diamond syndrome, with loss of zymogen granules. Gastroenterology. 2012;143(2):481–92.
Zambetti NA, Bindels EM, Van Strien PM, Valkhof MG, Adisty MN, Hoogenboezem RM, et al. Deficiency of the ribosome biogenesis gene Sbds in hematopoietic stem and progenitor cells causes neutropenia in mice by attenuating lineage progression in myelocytes. Haematologica. 2015;100(10):1285–93.
Venkatasubramani N, Mayer AN. A zebrafish model for the Shwachman-Diamond syndrome (SDS). Pediatr Res. 2008;63(4):348–52.
Provost E, Wehner KA, Zhong X, Ashar F, Nguyen E, Green R, et al. Ribosomal biogenesis genes play an essential and p53-independent role in zebrafish pancreas development. Development. 2012;139(17):3232–41.
Danilova N, Sakamoto KM, Lin S. Ribosomal protein S19 deficiency in zebrafish leads to developmental abnormalities and defective erythropoiesis through activation of p53 protein family. Blood. 2008;112(13):5228–37.
McGowan KA, Li JZ, Park CY, Beaudry V, Tabor HK, Sabnis AJ, et al. Ribosomal mutations cause p53-mediated dark skin and pleiotropic effects. Nat Genet. 2008;40(8):963–70.
Uechi T, Nakajima Y, Chakraborty A, Torihara H, Higa S, Kenmochi N. Deficiency of ribosomal protein S19 during early embryogenesis leads to reduction of erythrocytes in a zebrafish model of Diamond-Blackfan anemia. Hum Mol Genet. 2008;17(20):3204–11.
Dror Y. P53 protein overexpression in Shwachman-Diamond syndrome. Arch Pathol Lab Med. 2002;126(10):1157–8 (author reply 8).
Lindsley RC, Saber W, Mar BG, Redd R, Wang T, Haagenson MD, et al. Prognostic mutations in myelodysplastic syndrome after stem-cell transplantation. N Engl J Med. 2017 02;376(6):536–47.
Dror Y, Donadieu J, Koglmeier J, Dodge J, Toiviainen-Salo S, Makitie O, et al. Draft consensus guidelines for diagnosis and treatment of Shwachman-Diamond syndrome. Ann N Y Acad Sci. 2011;1242:40–55.
Gabrilove JL, Jakubowski A, Fain K, Grous J, Scher H, Sternberg C, et al. Phase I study of granulocyte colony-stimulating factor in patients with transitional cell carcinoma of the urothelium. J Clin Invest. 1988;82(4):1454–61.
Negrin RS, Haeuber DH, Nagler A, Kobayashi Y, Sklar J, Donlon T, et al. Maintenance treatment of patients with myelodysplastic syndromes using recombinant human granulocyte colony-stimulating factor. Blood. 1990;76(1):36–43.
Rosenberg PS, Alter BP, Bolyard AA, Bonilla MA, Boxer LA, Cham B, et al. The incidence of leukemia and mortality from sepsis in patients with severe congenital neutropenia receiving long-term G-CSF therapy. Blood. 2006;107(12):4628–35.
Ryan NJ, Lo JH. Delamanid: first global approval. Drugs. 2014;74(9):1041–5.
Zainal Abidin N, Haq IJ, Gardner AI, Brodlie M. Ataluren in cystic fibrosis: development, clinical studies and where are we now? Expert Opin Pharmacother. 2017;18(13):1363–71.
Kerem E, Konstan MW, De Boeck K, Accurso FJ, Sermet-Gaudelus I, Wilschanski M, et al. Ataluren for the treatment of nonsense-mutation cystic fibrosis: a randomised, double-blind, placebo-controlled phase 3 trial. Lancet Respir Med. 2014;2(7):539–47.
Finkel RS, Flanigan KM, Wong B, Bönnemann C, Sampson J, Sweeney HL, et al. Phase 2a study of ataluren-mediated dystrophin production in patients with nonsense mutation Duchenne muscular dystrophy. PLoS One. 2013;8(12):e81302.
Pranke I, Bidou L, Martin N, Blanchet S, Hatton A, Karri S, et al. Factors influencing readthrough therapy for frequent cystic fibrosis premature termination codons. ERJ Open Res. 2018. https://doi.org/10.1183/23120541.00080-2017.
Floquet C, Hatin I, Rousset JP, Bidou L. Statistical analysis of readthrough levels for nonsense mutations in mammalian cells reveals a major determinant of response to gentamicin. PLoS Genet. 2012;8(3):e1002608.
Linde L, Boelz S, Nissim-Rafinia M, Oren YS, Wilschanski M, Yaacov Y, et al. Nonsense-mediated mRNA decay affects nonsense transcript levels and governs response of cystic fibrosis patients to gentamicin. J Clin Invest. 2007;117(3):683–92.
Pibiri I, Lentini L, Tutone M, Melfi R, Pace A, Di Leonardo A. Exploring the readthrough of nonsense mutations by non-acidic Ataluren analogues selected by ligand-based virtual screening. Eur J Med Chem. 2016;122:429–35.
Pibiri I, Lentini L, Melfi R, Gallucci G, Pace A, Spinello A, et al. Enhancement of premature stop codon readthrough in the CFTR gene by Ataluren (PTC124) derivatives. Eur J Med Chem. 2015;101:236–44.
Sürün D, von Melchner H, Schnütgen F. CRISPR/Cas9 genome engineering in hematopoietic cells. Drug Discov Today Technol. 2018;28:33–9.
Shammas C, Menne TF, Hilcenko C, Michell SR, Goyenechea B, Boocock GR, et al. Structural and mutational analysis of the SBDS protein family. Insight into the leukemia-associated Shwachman-Diamond Syndrome. J Biol Chem. 2005;280(19):19221–9.
Delaporta P, Sofocleous C, Economou M, Makis A, Kostaridou S, Kattamis A. The Greek registry of Shwachman Diamond-Syndrome: Molecular and clinical data. Pediatr Blood Cancer. 2017;64:e26630.
Acknowledgements
The authors are grateful to Enrico Valletta (unit of Pediatrics, Azienda USL della Romagna, Italy) for helpful discussions, to Emily Pintani (Italian Shwachman-Diamond Syndrome Registry) for the excellent data management and to the Italian SDS Registry for providing us with some of the patient genetic data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
VB and MC are the inventors of the specific use of ataluren in SDS (patent provisional number 62/393,747 USA).
Funding
This review was supported by the “Associazione Italiana Sindrome di Shwachman” (Grant #AISS-2017 to MC and VB) and the Italian Ministry of Health (Grant GR-2016-02363570 to VB).
Rights and permissions
About this article
Cite this article
Bezzerri, V., Cipolli, M. Shwachman-Diamond Syndrome: Molecular Mechanisms and Current Perspectives. Mol Diagn Ther 23, 281–290 (2019). https://doi.org/10.1007/s40291-018-0368-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-018-0368-2