Abstract
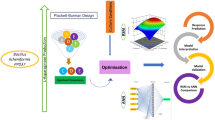
The utilization of bioluminescent bacteria has served as a powerful bioassay in environmental monitoring as a result of simplicity of use, quick response and high sensitivity on the basis of the luminescence being produced. In the present study, the optimization process for the luminescence production of a newly isolated Photobacterium sp. NAA-MIE using response surface methodology (RSM) and artificial neural network (ANN) is investigated. RSM and ANN are the most favored techniques and efficient methods for the optimization of medium components, especially for nonlinear systems. This analysis presents the comparative evaluation between RSM and ANN aimed at their particular empirical modeling and predictive ability. RSM predicted the optimized condition with four major variables at glycerol concentration of 0.16%, NaCl of 2.49%, pH at 7.4 and tryptone of 2.97% with 260,748.94 RLU of the maximum luminescence production. The accuracy of the RSM model equations was measured with R2 of 0.9001 and adjusted R2 of 0.8001. ANN predicted optimum condition at a glycerol concentration of 0.10%, NaCl of 2.46%, pH at 7.5 and tryptone of 1.97% with 237,481.32 RLU. The study demonstrated that ANN accomplishes more than RSM with a higher R2 value of 0.968 and low mean percentage error and root mean square error, which were 0.003 and 3.66, respectively. Both models are presented as suitable predictive models on the basis of mathematical modeling to optimize and improve biological systems for future upscaling processes with ANN as more predictive and having more fitting potentiality compared to RSM.








Similar content being viewed by others
References
Hastings JW (1995) Bioluminescence. In: Sperelakis N (ed) Cell physiology source book. Academic Press, San Diego, pp 665–681
Meighen EA (1991) Molecular biology of bacterial bioluminescence. Microbiol Rev 55:123–142
Hastings JW (2004) Bioluminescence. The website for the molecular imaging program at Stanford. https://mips.stanford.edu/public/abstracts/hastings/pdf. Accessed 22 Feb 2005
Vetrova E, Esimbekova E, Remmel N, Kotova S, Beloskov N, Kratasyuk V, Gitelson I (2007) A bioluminescent signal system: detection of chemical toxicants in water. Luminescence 22(3):206–214
Johnson BT, Blaise C, Férard F (2005) Microtox acute toxicity test. In: Small-scale freshwater toxicity investigations. Springer, Dordrecht, pp 69–105
Girotti S, Bolelli L, Roda A, Gentilomi G, Musiani M (2002) Improved detection of toxic chemicals using bioluminescent bacteria. Anal Chim Acta 471:113–120
Lee BS, Lee JG, Shin DH, Kim EK (2001) Statistical optimization of bioluminescence of Photobacterium phosphoreum KCTC2852. J Biosci Bioeng 92:72–76
Waters P, Lloyd D (1985) Salt, pH, and temperature dependencies of growth and bioluminescence of three species of luminous bacteria analysed on gradient plates. J Gen Microbiol 131:2865–2869
Nunes-Halldorson VS, Duran NL (2003) Bioluminescent bacteria: lux genes as environmental biosensors. Braz J Microbiol 34:91–96
Hassan SHA, Oh SE (2010) Improved detection of toxic chemicals by Photobacterium phosphoreum using modified Boss medium. J Photochem Photobiol B Biol 101(1):16–21
Witek-Krowiak A, Chojnacka K, Podstawczyk D, Dawiec A, Pokomeda K (2014) Application of response surface methodology and artificial neural network methods in modelling and optimization of biosorption process. Bioresour Technol 160:150–160
Muhamad MH, Abdullah SRS, Mohamad AB, Rahman RA, Kadhum AAH (2013) Application of response surface methodology (RSM) for optimisation of COD, NH 3-N and 2, 4-DCP removal from recycled paper wastewater in apilot-scale granular activated carbon sequencing batch biofilm reactor (GAC-SBBR). J Environ Manag 121:179–190
Seraman S, Rajendran A, Thangavelu V (2010) Statistical optimization of anticholesterolemic drug lovastatin production by the red mold Monascus purpureus. Food Bioprod Process 88:266–276
Ebrahimpour A, Rahman RN, Ch’ng DHE, Basri M, Salleh AB (2008) A modeling study by response surface methodology and artificial neural network on culture parameters optimization for thermostable lipase production from a newly isolated thermophilic Geobacillus sp. Strain ARM. BMC Biotechnol 8(1):96
Gaur R, Gupta A, Khare SK (2008) Lipase from solvent tolerant Pseudomonas aeruginosa strain: production optimization by response surface methodology and application. Bioresour Technol 99:4796–4802
Bas D, Boyacı İH (2007) Modeling and optimization II: comparison of estimation capabilities of response surface methodology with artificial neural networks in a biochemical reaction. J Food Eng 78:846–854
Beg QK, Saxena RK, Gupta R (2002) Kinetic constants determination for an alkaline protease from Bacillus mojavensis using response surface methodology. Biotechnol Bioeng 78:289–295
Senanayake SPJN, Shahidi F (2002) Lipase-catalyzed incorporation of docosahexaenoic acid (DHA) into borage oil: optimization using response surface methodology. Food Chem 7:115–123
Dutta JR, Dutta PK, Banerjee R (2004) Optimization of culture parameters for extracellular protease production from a newly isolated Pseudomonas sp. Using response surface and artificial neural network models. Process Biochem 39(12):2193–2198
Basri M, Rahman RN, Ebrahimpour A, Salleh AB, Gunawan ER, Rahman MB (2007) Comparison of estimation capabilities of response surface methodology (RSM) with artificial neural network (ANN) in lipase-catalyzed synthesis of palm-based wax ester. BMC Biotechnol 7(1):53
Bingol D, Hercan M, Elevli S, Kılıç E (2012) Comparison of the results of response surface methodology and artificial neural network for the biosorption of lead using black cumin. Bioresour Technol 112:111–115
Sanusi SNA, Halmi MIE, Abdullah SRS, Hassan HA, Hamzah FM, Idris M (2016) Comparative process optimization of pilot-scale total petroleum hydrocarbon (TPH) degradation by Paspalum scrobiculatum L. Hack using response surface methodology (RSM) and artificial neural networks (ANNs). Ecol Eng 97:524–534
Halmi MIE, Jirangon H, Johari WLW, Abdul Rachman AR, Shukor MY, Syed MA (2014) Comparison of Microtox and Xenoassay light as a near real time river monitoring assay for heavy metals. Sci World J. https://doi.org/10.1155/2014/834202
Thakre NA, Shanwar AS (2015) Promising biological indicator of heavy metal pollution: bioluminescent bacterial strains isolated and characterized from marine niches of Goa India. Indian J Microbiol 55(3):327–332
Miller RV, Day MJ (2004) Microbial evolution: gene establishment, survival, and exchange. ASM Press, Washington
Hasan HA, Abdullah SRS, Kamarudin SK, Kofli NT (2011) Response surface methodology for optimization of simultaneous COD, NH4C–N and Mn2C removal from drinking water by biological aerated filter. Desalination 275(1):50–61
Bezerra MA, Santelli RE, Oliveira EP, Villar LS, Escaleira LA (2008) Response surface methodology (RSM) as a tool for optimization in analytical chemistry. Talanta 76:965–977
Kannan S, Slochanal SMR, Subburaj P, Padhy NP (2004) Application of particle swarm optimization technique and its variants to generation expansion planning problem. Electr Power Syst Res 70:203–210
Zheng Y, Wang A (2010) Removal of heavy metals using polyvinyl alcohol semi-IPN poly (acrylic acid)/tourmaline composite optimized with response surface methodology. Chem Eng J 162(1):186–193
Wu Y, Zhou S, Qin F, Ye X, Zheng K (2010) Modeling physical and oxidative removal properties of Fenton process for treatment of landfill leachate using response surface methodology (RSM). J Hazard Mater 180:456–465
Lee W, Yusof S, Hamid NSA, Baharin BS (2006) Optimizing conditions for enzymatic clarification of banana juice using response surface methodology (RSM). J Food Eng 73:55–63
Maran JP, Manikandan S, Thirugnanasambandham K, Nivetha CV, Dinesh R (2013) Box-Behnken design based statistical modeling for ultrasound-assisted extraction of corn silk polysaccharide. Carbohydr Polym 92:604–611
Karthikeyan R, Balasubramanian V (2010) Predictions of the optimized friction stir spot welding process parameters for joining AA2024 aluminum alloy using RSM. Int J Adv Manuf Technol 51:173–183
Jadhav S, Surwase S, Phugare S, Jadhav J (2013) Response surface methodology mediated optimization of Remazol orange decolorization in plain distilled water by Pseudomonas aeruginosa BCH. Int J Environ Sci Technol 10:181–190
Ferreira S, Duarte AP, Ribeiro MH, Queiroz JA, Domingues FC (2009) Response surface optimization of enzymatic hydrolysis of Cistus ladanifer and Cytisus striatus for bioethanol production. Biochem Eng J 45:192–200
Hu Z-H, Yu H-Q, Zheng J-C (2006) Application of response surface methodology for optimization of acidogenesis of cattail by rumen cultures. Bioresour Technol 97:2103–2109
Kaur B, Kumar B, Garg N, Kaur N (2015) Statistical optimization of conditions for decolorization of synthetic dyes by cordyceps militaris MTCC 3936 using RSM. BioMed Res Int. https://doi.org/10.1155/2015/536745
Bashir MJ, Aziz HA, Yusoff MS, Adlan MN (2010) Application of response surface methodology (RSM) for optimization of ammoniacal nitrogen removal from semi-aerobic landfill leachate using ion exchange resin. Desalination 254:154–161
Linko S, Zhu YH, Linko P (1999) Applying neural networks as software sensors for enzyme engineering. Trends Biotechnol 17:155–162
Kalali A, Ebadi T, Rabbani A, Moghaddam SS (2011) Response surface methodology approach to the optimization of oil hydrocarbon polluted soil remediation using enhanced soil washing. Int J Environ Sci Technol 8:389–400
Gomez F, Sartaj M (2014) Optimization of field scale biopiles for bioremediation of petroleum hydrocarbon contaminated soil at low temperature conditions by response surface methodology (RSM). Int Biodeterior Biodegrad 89:103–109
Farmer J, Hickman-Brenner F (1992) The genera Vibrio and Photobacterium. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer K-H (eds) The prokaryotes, 2nd edn, vol III. Springer-Verlag, New York, pp 2952–3011
Soto W, Gutierrez J, Remmenga MD, Nishiguchi MK (2009) Salinity and temperature effects on physiological responses of Vibrio fischeri from diverse ecological niches. Microb Ecol 57:140–150. https://doi.org/10.1007/s00248-008-9412-9
Kumar A (2010) Isolation of bioluminescent bacteria from Bay of Bengal and their molecular characterization. PhD thesis. Department of Biotech, AMET University, Chennai, India
Onorati F, Mecozzi M (2004) Effects of two diluents in the Microtox toxicity bioassay with marine sediments. Chemosphere 54(5):679–687
Hong Y, Chen Z, Zhang B, Zhai Q (2010) Isolation of Photobacterium sp. LuB-1 and its application in rapid assays for chemical toxicants in water. Lett Appl Microbiol 51(3):308–312
Prosser CL (1973) Comparative physiology, 3rd edn. Saunders College, Philadelphia
Acknowledgements
This study was assisted by the research grant from Putra Grant (GP-IPM/2017/9532800) and Yayasan Pak Rashid Grant UPM (6300893-10201).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Significance Statement
The comparative medium optimization of luminescent production by Photobacterium sp. NAA-MIE was carried out using RSM and ANN. Both models are presented as suitable predictive models ANN is more precise and reproducible compared to RSM.
Rights and permissions
About this article
Cite this article
Adnan, N.A., Halmi, M.I.E., Gani, S.S.A. et al. Statistical Modeling for the Optimization of Bioluminescence Production by Newly Isolated Photobacterium sp. NAA-MIE. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 90, 797–810 (2020). https://doi.org/10.1007/s40011-019-01154-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-019-01154-1