Abstract
In honey bees (Apis mellifera), methyl palmitate (MP), methyl oleate (MO), methyl linoleate (ML), and methyl linolenate (MLN) are important pheromone components of the capping pheromones triggering the capping behavior of worker bees. In this study, we compared the amounts of these four pheromone components in the larvae of workers and drones, prior to be capped, in the process of being capped and had been capped. The amounts of MP, MO, and MLN peaked at the capping larval stage, and ML was highest at capped larvae in worker larvae, whereas in drone larvae, the amounts of the four pheromone components were higher overall and increased with aging. Furthermore, we proposed de novo biosynthetic pathways for MP, MO, ML, and ML, from acetyl-CoA. Besides, stable isotope tracer 13C and deuterium were used to confirm that these capping pheromone components were de novo synthesized by larvae themselves rather than from their diets.
Similar content being viewed by others
1 Introduction
Honey bees have been instrumental in revealing both the significance and the complexity of pheromonal communication systems. Pheromones help to coordinate a colony’s collective behaviour, such as foraging, defensive behaviour, and brood-rearing activity (Free and Winder 1983; Free 1987; Vallet et al. 1991; Breed et al. 2004; Hunt 2007; Stout and Goulson 2001; Slessor et al. 2005; Maisonnasse et al. 2010). The subtleties and complexities of bee pheromonal communication are perhaps best illustrated by the pheromonal communication between workers and developing larvae.
Honey bee larvae are entirely dependent on their attendant adult workers. Larvae release capping pheromone components, form their salivary glands containing ten methyl or ethyl fatty acid esters (Le Conte et al. 1989, 1990, 2006; Trouiller et al. 1991). The composition of this pheromone varies with larval age and sex (worker or drone), and workers adjust their behavioral responses to larvae accordingly (Free and Winder 1983; Le Conte 1994; Slessor et al. 2005). When larvae reach the stage of pupation, their cells are closed off by workers constructing a thin wax cap over the cell so that the larvae pupate in a clean and stable environment. Four of the components of BEP are particularly important in triggering the capping behaviour of workers. These are methyl palmitate (MP), methyl oleate (MO), methyl linoleate (ML), and methyl linolenate (MLN) (Le Conte et al. 1990). Production of these components by larvae increases quite dramatically when the larva reaches the developmental stage prior to pupation (Trouiller et al. 1991).
Currently, our knowledge of how pheromones are synthesized and released by worker and drone larvae at different developmental stages are limited. To fulfill the gaps, here we examined changes in the BEP components related to cell capping in both worker and drone larvae pre, during and post-cell capping. In parallel, we analyzed gene expression changes in larvae across the cell capping stage as a first step to the identification of biosynthetic pathways in bees for pheromone components.
2 Materials and methods
2.1 Experimental honey bee colonies
Honey bee colonies (Apis mellifera) used throughout this study were maintained at the Honeybee Research Institute, Jiangxi Agricultural University, Nanchang, China (28.46° N, 115.49° E), according to standard beekeeping techniques.
2.2 Gas chromatograph-mass spectroscopy analysis
Amounts of capping pheromone components were compared in the following sample groups: (i) non-capped larvae of 4-day-old (4th) instar of workers and drones; (ii) capping-stage worker and drone larvae sampled as wax caps were being constructed over their cells; (iii) capped worker and drone larvae sampled from cells that had been capped completely and had a thin silk cocoon under the wax cover, which were approximately 8-day-old instar for worker and 9-day-old instar for drones. There were four biological replicates of each group, with each replicate sourced from a different colony. Each sample contained groups of larvae to provide enough biological material for gas chromatograph-mass spectroscopy (GC/MS) analysis. For each sample, we collected 40 4th instar worker and drone larvae, 20 capping-stage and capped worker larvae, and 5 capping-stage and capped drone larvae from brood frames.
The whole larvae of each sample group freshly collected from their wax cells were placed in (without body washes) glass vials with 3 ml dichloromethane, respectively. Twenty microliters of methyl nonadecanoate (≥ 98%, AccuStandard, 10 μg/ml) was added to each vial as the internal standard. Afterwards, the glass vials with larvae were shaken lightly for 30 min using an oscillator (120 r/min), and then, the solution was transferred into a clean vial and concentrated under nitrogen flow to about 20 μl. A 1-μl sample was injected into the GC/MS.
A GC/MS system (7890A/5975C, Agilent Technologies Inc., Santa Clara, CA, USA) with an HP-Innowax chromatographic column (60-m, 0.25-mm, 0.25-μm film thickness, 19091N-136, Agilent Technologies) was used. Helium was used as carrier gas at a constant flow rate of 1.0 ml/min. Samples were analyzed with time split injection mode and split ratio of 2:1. The column temperature was programmed as follows: 120 °C for 1 min, then rising to 180 °C at a rate of 5 °C/min, and then rising to 230 °C at a rate of 2 °C/min, finally cooling to 250 °C at a rate of 5 °C/min and held constant for 10 min. The temperature of the sample injection port was 250 °C, and the pressure of helium was 13.5552 psi. Sample ionization was performed in electron impact ionization mode at 70 eV. The temperature of ion source, electrode stem, and transmission line were 250 °C, 150 °C, and 250 °C, respectively. Single-ion monitoring chromatograms were reconstructed at the base peak for MP, MO, ML, and MLN and the internal standard: mass-to-charge ratio (m/z) 270, 264, 294, 292, and 312.
For each compound of interest, peak areas were converted to amounts by reference to calibration curves for each compound. MP, MO, ML, and MLN (≥ 99.5%, Sigma) were used as external standards. Seven concentrations of each external standard (0.25 μg/ml, 0.5 μg/ml, 1 μg/ml, 2 μg/ml, 5 μg/ml, 10 μg/ml, and 25 μg/ml) were used to construct calibration curves. All of the correlation coefficients of the four calibration curves were > 99.96% (Figure S1), which suggested that the GC/MS system was stable enough for the later quantitative analysis of MP, MO, ML, and MLN from honey bee larvae.
2.3 RNA-Seq data analysis
Samples from the groups described above were also used for RNA-sequence analysis to analyze differences in gene expression between groups. For each sample, we collected six workers or drone larvae and created three biological replicates with each replicate sourced from a different colony. The RNA was extracted with Trizol. The samples were flash frozen in liquid nitrogen and stored at − 80 °C until use.
RNA extraction, RNA sequencing, and data analysis were performed according to the methods described in He et al. (2016). Briefly, the raw counts for each gene were normalized with Fragments Per Kilobase of transcript per Million (FPKM) to calculate the P value for significantly regulated genes with DESeq packages. The P value was further corrected for multiple comparisons with false discovery rate (FDR). Significantly differentially expressed genes (DEGs) between different samples were defined as genes with a FDR ≤ 0.05 and the absolute value of log2 ratio ≥ 1.
2.4 Identification of pheromone biosynthesis pathway
The longest transcripts of significantly differentially expressed genes were converted to protein sequences and then compared with the Kyoto Encyclopedia of Genes and Genomes (KEGG) protein database by using Basic Local Alignment Search Tool (BLAST) with E-value < 10−5 set as the cutoff criterion. To identify putatively enriched metabolic pathways in the RNA-seq data, the outcome of the KEGG mapping was analyzed with the KEGG Orthology Based Annotation System 2.0 (KOBAS 2.0). The KOBAS 2.0 (Xie et al. 2011) references the provided gene database against multiple existing databases of metabolic and signaling pathways to identify pathways that are enriched in the RNA-seq samples.
2.5 Verification of DEGs with qRT-PCR analysis
Six genes, which participate in the inferred biosynthetic pathways of MP, MO, ML, and MLN (Table SI) and a “housekeeping gene” (β-actin), were selected for qPCR analysis. For each group, 3 larvae from each of three colonies were sampled, yielding three biological replicates per group. Samples were flash frozen in liquid nitrogen and stored at − 80 °C until used. Detailed methods for qRT-PCR are described in our previous study (Qin et al. 2014). Primers for these genes were designed by Primer 5.0 (Table SI). For each gene, three technical replicates were performed for each sample. The Ct value for each sample was obtained by calculating the mean of three technical replicates. Relative expression levels between samples were calculated using the formula 2−ΔΔCt reported by Liu and Saint (2002) and then square-root-transformed to normalize the distribution of the data before ANOVA analysis.
2.6 Stable isotopic tracing
While insects are able to synthesize fatty acids de novo, they can also get them from their diets. In order to provide further evidence to demonstrate that these capping pheromone components were de novo synthesized by larvae themselves rather than from their diets, stable isotope tracer method was used.
In this experiment, the basic larval food (BLD) consists of 50% royal jelly, 6% fructose, 6% glucose, 1% yeast extract, and 37% ddH2O (Vandenberg and Shimanuki 1987). The glucose in the BLD was consist of 20% 13C-glucose (Sigma) and 80% 12C-glucose. Pheromone components were collected from the following sample groups: (i) larvae reared with BLD as control group (C); (ii) larvae reared with BLD and 13C-glucose (Sigma) as treatment group 1 (T1); (iii) larvae reared with BLD with 0.25% 2H-methanol (Sigma) as treatment group 2 (T2). Larval food of all groups was stored at − 80 °C to keep fresh and was pre-warmed at 35 °C for 15 min before usage.
One-day worker larvae were removed from the comb and transferred to 24-well cell culture plates with 200 μl of larval food per well and then were placed in the incubator at 35 °C and 90% relative humidity. Every 12 h, larvae were fed with 100-μl fresh food per well. When began to defecate, larvae were removed from the cell culture plates, dried on Kimwipe™ tissues, and transferred to the pupation plates with Kimwipe™ tissues on the bottom (Kucharski et al. 2008). Six hours later, every 10 larvae of the same group were freshly collected from the pupation plates to a glass vial with 3 ml dichloromethane, respectively. The abundances of 13C and 2H of the capping pheromone was analyzed using GC/MS system (7890A/5975C, Agilent Technologies Inc., Santa Clara, CA, USA) by Shanghai Research Institute of Chemical Industry CO., LTD. Sample treatment method and experiment condition were the same as the GC/MS analysis as described previously. Three biological replicates were made for each group. The abundances of 13C and 2H in control and treated groups were calculated by amounts of 13C- and 2H-marked capping pheromone against the total amount of related pheromone, respectively. Three capping pheromones were detected with 13C and 2H and one pheromone, ML, failed.
2.7 Statistics
StatView 5.01 (SAS Institute, Cary, NC, USA) was used to analyze all data from the GC/MS and qRT-PCR of each group, by using the ANOVA followed by a Fisher PLSD test. Differences between groups were considered to be significant at the probability level of 0.05%.
3 Results
3.1 Levels of capping pheromone components in larvae of different developmental stages
The amounts of MP, MO, ML, and MLN in samples of worker larvae are shown in Figure 1. The levels of MP, MO, and MLN were significantly higher in capping-stage worker larvae compared with the 4th instar and capped larvae. Additionally, the amount of ML was significantly higher in capped larvae than in 4th instar and capping-stage larvae.
Results for drone larvae are shown in Figure 2. In drones, the amounts of all four pheromone components increased with age from 4th instar to capped larvae and were significantly higher in capped larvae than in 4th instar and capping-stage larvae.
3.2 RNA sequence analysis in larvae of different capping stage
Summaries of RNA sequence sample quality are shown in Table SII. In the libraries, the alignment rate between the reads and reference genome was all above 90.00%. The Pearson correlation coefficient of capped WL-R2 and capped DL-R1 was too low with the other two replicate samples; thus, they were eliminated in the following analysis (Table SIII). But beyond that, the Pearson correlation coefficients among the other three biological replicates of all experimental groups were all ≥ 0.75 (Table SIII), which is a conventionally accepted threshold for valid replicates (Tarazona et al. 2011) indicating that there was acceptable sequencing quality and repeatability among the biological replicates of each group.
The number of DEGs varied dramatically among developmental stages (using a threshold of log2 ratio ≥ 1, FDR < 0.01, Table SIV). The top 5 significantly enriched KEGG pathways of DEGs were glycolysis/gluconeogenesis (ko00010); citrate cycle (TCA cycle, ko00020); pentose phosphate pathway (ko00030); valine, leucine, and isoleucine biosynthesis (ko00290); and glutathione metabolism (ko00480). These five pathways were not involved in pheromone biosynthesis pathways.
However, two KEGG pathways were identified possibly relating to the long carbon chain biosynthesis: one for fatty acid elongation (ko00062) and one for biosynthesis of unsaturated fatty acid (ko01040), which we predicted might be involved in the biosynthesis of four capping pheromone components. The fatty acid elongation pathway involves in the de novo synthesis of hexadecanoate (C16) from acetyl-coenzyme A (acetyl-CoA), and the biosynthesis of unsaturated fatty acid pathway is involved in the formation of one or more double-bond unsaturated fatty acids such as oleic acid and linoleic acid by using C16 as a precursor. This pathway has been reported to be involved in the pheromone biosynthesis of some insect species (Roelofs and Bjostad 1984; Ando et al. 1988; Tang et al. 1989).
In this study, 6 genes Kat (GB50970: 3-ketoacyl-CoA thiolase, mitochondrial), Hadha (GB45128: trifunctional enzyme subunit alpha, mitochondrial), Echs1 (GB40460: enoyl coenzyme A hydratase short chain 1, mitochondrial), Mecr (GB43706: probable trans-2-enoyl-CoA reductase, mitochondrial), Ppt2 (GB40083: lysosomal thioesterase PPT2 homolog), and Ppt1 (GB49609: palmitoyl-protein thioesterase 1) were enriched in the fatty acid elongation pathway, and Vlcecr (GB46772: very-long-chain enoyl-CoA reductase), Vlchacd2 (GB54258: very-long-chain (3R)-3-hydroxyacyl-CoA dehydratase 2), and four acyl-CoA Δ11 desaturase-like genes (Δ11-desaturases, GB48193, GB48194, GB48195, and GB51238) were enriched in the biosynthesis of unsaturated fatty acid pathway. All these 12 genes were significantly differentially expressed in at least one comparison among 4th instar and capping and capped larvae (Table SV). In particular, GO annotation showed that two Δ11-desaturase genes (GB48193 and GB48195) had Δ9-desaturase activity (Table SV). We therefore proposed these as candidate genes for the process of biosynthesis of MP, MO, ML, and MLN in honey bee larvae (Figure 3).
Proposed biosynthetic pathways involved in the synthesis of capping pheromone components and expression of related genes in worker and drone larvae of different capping stages. The grids illustrate the gene expression levels for 4th instar and capping and capped larvae of workers and drones. Greyscale density of each cell shows the absolute expression magnitude of honey bee larvae, with the FPKM values 0–10, 10–20, 20–40, 40–80, 80–160, 160–320, 320–640, 640–1280, 1280–2560, and 2560–5500 represented by gray scale levels, respectively.
We compared our four candidate Δ11-desaturase genes GB48193, GB48194, GB48195, and GB51238 with domestic silkworm (Bombyx mori) acyl-CoA delta (11) desaturase by the protein BLAST using the National Center for Biotechnology Information (NCBI) website. Protein BLAST results showed that these four genes were high homology with Bombyx mori acyl-CoA delta (11) desaturase with positives of 62%, 46%, 63%, and 67%, respectively.
3.3 Results of qRT-PCR
Figure 4 shows the results of the qRT-PCR analyses alongside the results of RNA sequence analyses for four genes from the fatty acid elongation pathway and two of the Δ11-desaturase-like genes. The results of the qRT-PCR analysis were concordant with the results of the RNA sequence. Furthermore, the general expression tendency showed that these genes had a higher relative expression level both in 4th instar and capping larvae than that in capped larvae.
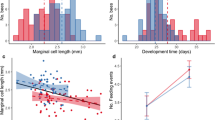
Expression levels of six genes possibly implicated in the biosynthesis of capping pheromone components in larvae of different capping stages (4th instar, capping, and capped). Estimates of relative gene expression level assessed with RNA-seq and qRT-PCR are shown for each gene. Different superscripts (a, b, and c) to bars indicate significant differences in expression between capping stages (qRT-PCR: P < 0.05, RNA-seq: FDR ≤ 0.05). Relative expression data of qRT-PCR were transformed by square root transformation and were presented here after transformation. Each bar corresponds to a single group represented as the mean ± S.E. of its biological replicates.
3.4 Isotope abundance analysis
The abundances of 13C and 2H from different capping pheromone components are shown in Table I. On account of the natural isotopic abundance, 13C and 2H were detected from control groups as well. Nevertheless, for MP, MO, and MLN (ML was not detected for its low abundance), there was at least one of the abundances of 13C or 2H in treatment groups that was significantly higher than that in control groups.
4 Discussion
MP, MO, ML, and MLN have been reported to be important signals for honey bee capping behaviour (Le Conte et al. 1990). Our data is consistent with Trouiller et al. (1992), indicating that honey bee larvae increase the amounts of capping pheromone components at a critical larval stage for wax capping. Yan et al. (2009) also reported a similar releasing trend of pheromone levels in Apis cerana worker and drone larvae. However, Trouiller et al. (1992) found capping pheromone decreased in Apis mellifera drone larvae post-capping (8–10 instars). Drone cells are normally located at the edge of natural brood frames (Winston 1991), and in this study, we sampled drones from the edges of natural frames. Trouiller et al. (1992) caged a queen onto a drone frame to lay eggs and probably collected drone samples from central areas. There are differences in temperature and temperature variability across brood frames, as well as differences in larval attendance rates by workers. Whether position on the frame influences pheromone accumulation is presently unknown, but would be worthy of further study.
Varroa mite is the worst pest of Apis mellifera, resulting in high death rate of honey bees around the world (Le Conte et al. 2010). Three BEP components have been reported to be attractive Varroa mites in honey bee colonies (MP has very strong attractive response, EP and ML have weaker response, Le Conte et al. 1989). In the present study, the amounts of MP and ML in drone larvae were dramatically higher compared with worker larvae at all stages (Figures 1 and 2). Our study supports earlier suggestions that these BEP components may be used by Varroa to differentiate and select drone brood at the capping stage from worker brood (Le Conte et al. 1989).
In many insect species, acetyl-CoA is the precursor of several divergent metabolic pathways leading to the biosynthesis of fatty acids and esters (Mcgee and Spector 1975; Cripps et al. 1986; Moshitzky et al. 2003). For example, in silkworm and some species, methyl esters (including methyl hexadecanoate) and the sex pheromone bombykol were de novo synthesized from acetyl-CoA to hexadecanoate or octadecanoate, respectively (Ando et al. 1988; Tang et al. 1989). Furthermore, similar pathways were found in the methyl esters of leptospira and soybean (Stern et al. 1969; Bachlava et al. 2009). These studies support the reliability of our results on the biosynthesis pathway of four honey bee capping pheromone components.
In many insects and animals, stearic acid is generally synthesized via two pathways (Wang et al. 2002): in mitochondria, stearic acid can be biosynthesized from palmitic acid by mainly five steps (condensation, reduction, dehydration, reduction, and release); in endoplasmic reticulum, stearic acid can be directly biosynthesized by using palmitic acid as substrate. For the unsaturated methyl esters, it is well-known that Δ9-desaturase, Δ12-desaturase, and Δ15-desaturase are the relative desaturases for the synthesis of MO, ML, and MN (Wang et al. 2002). In this study, 4 Δ11-desaturase genes were enriched in the proposed biosynthetic pathway of MO, ML, and MN in honey bees. We suspect that these four Δ11-desaturase genes may play the same function as Δ9-desaturase, Δ12-desaturase, and Δ15-desaturase in the biosynthetic process of these three unsaturated methyl esters. Firstly, the Δ11-desaturase is a biofunctional enzyme having for example Δ11 and Δ12 desaturation activities (Serra et al. 2006), playing an important role in insect pheromone biosynthesis (Lofstedt et al. 1986; Rodriguez et al. 1992; Foster 1998). Secondly, GO annotation showed that two Δ11-desaturase gene (GB48193 and GB48195) had stearoyl-CoA 9-desaturase activities (Table SV). Moreover, there are no Δ9-desaturase, Δ12-desaturase, and Δ15-desaturase genes in the GenBank database, but 12 Δ11-desaturase genes have been annotated. We mapped all 12 honey bee Δ11-desaturase genes to house cricket’s (Acheta domesticus) Δ9-desaturase which biological function has been confirmed (Riddervold et al. 2002). The protein of GB48195 had the highest homology of 66% with the Δ9-desaturase of house cricket and could not map to its Δ11-desaturase. Therefore, the gene GB48195 may be the Δ9-desaturase ortholog of honey bees. Eventually, Roelofs and Bjostad (1984) showed that the oleate, linoleate, and linolenate in the red banded leaf roller moth, cabbage looper moth, and the domestic silkworm are de novo synthesized and desaturated by a delta (11) desaturase enzyme. Consequently, it is believed that these four Δ11-desaturase genes are involved in the biosynthetic pathways of honey bee capping pheromone components. For the last step of fatty acids becoming methyl esters, Castillo et al. (2012) showed that honey bee ethyl oleate was synthesized by dehydration of oleic acid and free ethanol under regulating of genes GB11403 and GB13365. In our study, the gene GB13365 (updated BeeBase: GB43508, lipase member H-A-like) was significantly differentially expressed among three larval groups. However, whether this gene participates in regulating the biosynthetic process of honey bee methyl esters from fatty acid and free methanol still requires further investigations. While insects are able to synthesize fatty acids de novo, they can also get them from their diets. Results of stable isotope tracing of 13C and 2H provided further evidence to demonstrate that these capping pheromone components were de novo synthesized by larvae themselves rather than from their diets and made a definite connection between the gene expression seen and the biosynthesis of the methyl esters. However, it is a complex process from gene regulation to pheromone production. The regulation of gene expression can be carried out on multiple levels including gene, transcription, post-transcription, translation, and post-translation. According to our results, we proposed these 12 genes as candidate genes for the process of biosynthesis of MP, MO, ML, and MLN in honey bee larvae, but the exact regulatory mechanism still requires further investigations.
In summary, this study showed that honey bee Apis mellifera worker and drone larvae increased their capping pheromone components during the wax capping process. Drone larvae released significantly higher pheromones than worker larvae, which attracts more Varroa mites during wax capping. We also identified putative biosynthetic pathways for these pheromones and predicted that these pheromones are de novo from acetyl-CoA. Our findings contribute information on how brood pheromones are biosynthesized and released in honey bees and how these pheromones influence the host selection of honey bee pests.
References
Ando, T., Hase, T., Arima, R., Uchiyama, M. (1988). Biosynthetic pathway of bombykol, the sex pheromone of the female silkworm moth. Agric. Bioi. Chern. 52(2), 473–478.
Bachlava, E., Dewey, R.E., Burton, J.W., Cardinal, A.J. (2009). Mapping candidate genes for oleate biosynthesis and their association with unsaturated fatty acid seed content in soybean. Mol. Breeding 23(2), 337–347.
Breed, M.D., Guzman-Novoa, E., Hunt, G.J. (2004) Defensive behavior of honey bees: organization, genetics, and comparisons with other bees. Annu. Rev. Entomol. 49, 271–298.
Castillo, C., Chen, H., Graves, C., Maisonnasse, A., Le, C.Y., Plettner, E. (2012). Biosynthesis of ethyl oleate, a primer pheromone, in the honey bee (Apis mellifera L.). Insect Biochem. Molec. 42(6), 404–416.
Cripps, C., Blomquist, G.J., Renobales, M.D. (1986). De novo biosynthesis of linoleic acid in insects. BBA-Lipid Lipid Met. 876(3), 572–580.
Foster, S.P. (1998). Sex pheromone biosynthesis in the tortricid moth planotortrix excessana (walker) involves chain-shortening of palmitoleate and oleate. Arch. Insect Biochem. 37(2), 158–167.
Free, J.B. (1987) Pheromones of Social Bees. Chapman & Hall, London
Free, J.B., Winder, M.E. (1983) Brood recognition by honey bee Apis mellifera workers. Anim. Behav. 31, 539–545.
He, X.J., Zhang, X.C., Jiang, W.J., Barron, A.B., Zhang, J.H., Zeng, Z.J. (2016) Starving honey bee (Apis mellifera) larvae signal pheromonally to worker bees. Sci. Rep. 6, 22359.
Hunt, G.J. (2007) Flight and fight: a comparative view of the neurophysiology and genetics of honey bee defensive behavior. J. Insect Physiol. 53, 399–410.
Kucharski, R. Maleszka, J., Foret, S., Maleszka, R. (2008) Nutritional control of reproductive status in honey bees via DNA methylation. Science 319(5871): 1827–30.
Le Conte, Y. (1994) The Recognition of Larvae by Worker Honeybees. Naturwissenschaften. 81, 462–465.
Le Conte, Y., Arnold, G., Troiler, J., Masson, C., Chappe, B., Ourisson, G. (1989) Attraction of the parasitic mite Varroa to the drone larvae of honey bees by simple aliphatic esters. Science, 245, 638–639.
Le Conte, Y., Arnold, G., Trouiler, J., Masson, C. (1990) Identification of a brood pheromone in honeybees. Naturwissenschaften.77, 334-336.
Le Conte Y., Bécard, J.M., Costagliola G., de Vaublanc, G., El Maâtaoui, M., Crauser, D., Plettner, E., Slessor, K.M. (2006) Larva salivary glands are a source of primer and releaser pheromone in honey bee (Apis mellifera L.). Naturwissenschaften. 93, 237–241.
Le Conte, Y., Ellis, M., Ritter, W. (2010). Varroa mites and honey bee health: can Varroa, explain part of the colony losses? Apidologie, 41(3), 353–363.
Liu, W., Saint, D.A. (2002) A new quantitative method of real time reverse transcription polymerase chain reaction assay based on simulation of polymerase chain reaction kinetics. Anal. Biochem. 1, 52–59.
Lofstedt, C., Elmfors, A., Sjögren, M., Wijk, E. (1986). Confirmation of sex pheromone biosynthesis from (16-d3) palmitic acid in the turnip moth using capillary gas chromatography. Cell Mol. Life Sci. 42(9), 1059–1061.
Maisonnasse, A., Lenior, J.C., Beslay, D., Crauser, D., Le Conte, Y. (2010) E-β-ocimene, a volatile brood pheromone involved in social regulation in the honey bee colony (Apis mellifera). PLoS One. 5, 1–7.
Mcgee, R., Spector, A. A. (1975). Fatty acid biosynthesis in erlich cells. the mechanism of short term control by exogenous free fatty acids. J. Biol. Chem. 250(14), 5419.
Moshitzky, P., Miloslavski, I., Aizenshtat, Z., Applebaum, S. W. (2003). Methyl palmitate: a novel product of the medfly (ceratitis capitata) corpus allatum. Insect Biochem. Mol. Biol. 33(12), 1299–306.
Qin, Q.H., Han, X., Liu, H., Zhang, S.W., Zeng, Z.J. (2014) Expression levels of glutamate and serotonin receptor genes in the brain of different behavioural phenotypes of worker honeybee (Apis mellifera).Türk. Entomol. Derg. 38 (4), 431–441.
Riddervold, M. H., Tittiger, C., Blomquist, G. J., Borgeson, C. E. (2002). Biochemical and molecular characterizaton of house cricket (acheta domesticus, orthoptera: gryllidae) Δ9 desaturase. Insect Biochem. Mol. Biol. 32(12), 1731–1740.
Rodriguez, F., Hallahan, D. L., Pickett, J. A., Camps, F. (1992). Characterization of the Δ11-palmitoyl-coa-desaturase from spodoptera littoralis (lepidoptera: noctuidae). Insect Biochem. Mol. Biol. 22(2), 143–148.
Roelofs, W., Bjostad, L. (1984). Biosynthesis of lepidopteran pheromones. Bioorg. Chem. 12(4), 279–298.
Serra, M., Piña, B., Bujons, J., Camps, F., Fabriàs, G. (2006). Biosynthesis of 10, 12-dienoic fatty acids by a bifunctional Δ11desaturase in Spodoptera littoralis. Insect Biochem. Mol. Biol. 36(8), 634–641.
Slessor, K.N., Winston, M.L., Le Conte, Y. (2005) Pheromone communication in the honey bee (Apis mellifera L.). J. Chem. Ecol. 31, 2731–2745.
Stern, N., Shenberg, E., Tietz, A. (1969). Studies on the metabolism of fatty acids in leptospira: the biosynthesis of Δ9-and Δ11 -monounsaturated acids. European J. Biochem. 8(1), 101–108.
Stout, J.C., Goulson, D. (2001) The use of conspecific and interspecific scent marks by foraging bumble-bees and honeybees. Anim. Behav. 62, 183–189.
Tang, J.D., Charlton, R.E., Jurenka, R.A., Wolf, W.A., Phelan, P.L., Sreng, L., Roelofs, W.L. (1989). Regulation of pheromone biosynthesis by a brain hormone in two moth species. Proc. Natl. Acad. Sci. USA. 86(6), 1806–10.
Tarazona, S., García-Alcalde, F., Dopazo, J., Ferrer, A., Conesa, A. (2011). Differential expression in RNA-seq: a matter of depth. Genome Res. 21, 2213–2223.
Trouiller, J., Arnold, G., Le Conte, Y., Masson, C., Chappe, B. (1991) Temporal pheromonal and kairomonal secretion in the brood of honeybees. Naturwissenschaften. 78(8), 368–370.
Trouiller, J., Arnold, G., Chappe, B., Le Conte, Y., Masson, C. (1992). Semiochemical basis of infestation of honey bee brood by Varroa jacobsoni. J Chem. Ecol. 18(11), 2041–53.
Vallet, A., Cassier, P., Lensky, Y. (1991) Ontogeny of the fine structure of the mandibular glands of the honeybee (Apis mellifera L.) workers and the pheromonal activity of 2-heptanone. J. Insect Physiol. 37, 789–804.
Vandenberg, J.D., Shimanuki, H. (1987) Technique for rearing worker honeybees in the laboratory. J. Apicult. Res. 26(2), 90–97.
Wang, J. Y., Zhu, S. G., Xu, C. F. Biological chemistry (Third edition). China higher education press, Beijing, 2002.
Winston, M. (1991) The Biology of the Honey Bee. Harvard University Press, Cambridge, MA, USA.
Xie, C., Mao, X., Huang, J., Ding, Y., Wu, J.M., Dong, S., Kong, L., Gao, G., Li, C.Y., Wei, L.P. (2011). KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic. Acids. Res. 39, W316-W322.
Yan, W.Y., Le Conte, Y., Beslay, D., Zeng, Z.J. (2009). Identification of brood pheromone in Chinese honeybee [Apis cerana cerana (Hymenoptera: apidae)]. Scientia Agricultura Sinica 42(6), 2250–2254. (In Chinese).
Acknowledgments
We thank Prof. Junwu Ma for help on biosynthetic pathway analysis and Dr. Qiang Huang for revising the manuscript.
Contributions
QHQ, XJH, and WJJ conducted all experiments; ZJJ designed the experiments; QHQ, XJH, and ABB wrote the paper; and LG participated in experiments and data analysis.
Funding
This work was supported by the National Natural Science Foundation of China (31572469, 31872432) and the Earmarked Fund for China Agriculture Research System (CARS-44-KXJ15).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Klaus Hartfelder
Les phéromones d’operculation et les voies de biosynthèse suspectées chez les larves d'ouvrières et de mâles d’ Apis mellifera
Abeille domestique / larve / operculation / phéromones / voies de synthèse
Die Verdeckelungspheromone und deren vermuteten Biosynthesewege bei Arbeiterinnen- und Drohnenlarvern der Honigbiene Apis mellifera
Honigbiene / Larven / Verdeckelung / Pheromone / Synthesewege
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
All sequencing data has been submitted to GenBank under accession numbers SRR4051855, SRR4051820, bioproject PRJNA339509
Electronic supplementary material
Figure S1.
The correlation coefficients of the four standard curves. (PNG 25 kb)
Table SI.
Primers for qRT-PCR. (XLS 29 kb)
Table SII.
Statistics of RNA sequencing in worker and drone larvae. (DOCX 19 kb)
Table SIII.
Pearson correlation coefficient among three biological replicates of each larval group. (XLSX 15 kb)
Table SIV.
The number of significantly differentially expressed genes in larvae of different capping stage. (DOCX 17 kb)
Table SV.
Expression of genes relating to the biosynthesis of capping pheromone components in larvae of different capping stages. (XLSX 26 kb)
Rights and permissions
About this article
Cite this article
Qin, QH., He, XJ., Barron, A.B. et al. The capping pheromones and putative biosynthetic pathways in worker and drone larvae of honey bees Apis mellifera. Apidologie 50, 793–803 (2019). https://doi.org/10.1007/s13592-019-00686-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-019-00686-9