Abstract
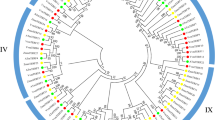
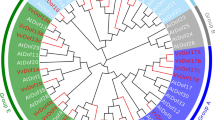
Plant mitochondrial transcription termination factor (mTERF) is a large and important family with valuable roles in organizing of organelle gene expression under the various stresses. In this study, genome wide analysis of mTERF regulatory genes was achieved and twenty-five potential mTERF genes response water deficit in grape (Vitis vinifera L.) were determined. Most of them were targeted to organelles genome especially mitochondria. It was found that Grape mTERFs were clustered into six main groups based on phylogenetic analysis. As a result of comprehensive expression analysis of these genes, using RNA-seq data in this study revealed that these genes have various expression profiles. With this study, various important roles of mTERF genes of grape under the water deficit stress were reported at first in grape. All results were found will be useful for elucidating the roles of mTERF genes in the growth, development and stress response of grape and fundamental for functional genomic studies.







Similar content being viewed by others
Abbreviations
- BP:
-
Biological processes
- CC:
-
Cellular component
- MF:
-
Molecular functions
- m-TERF:
-
Mitochondrial transcription termination factor
- NJ:
-
Neighbor-joining
References
Babiychuk E, Vandepoele K, Wissing J, Garcia-Diaz M, De Rycke R, Akbari H, Joubès J, Beeckman T, Jänsch L, Frentzen M (2011) Plastid gene expression and plant development require a plastidic protein of the mitochondrial transcription termination factor family. Proc Natl Acad Sci 108(16):6674–6679
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34(Web Server issue):W369–W373. https://doi.org/10.1093/nar/gkl198
Baloglu MC, Eldem V, Hajyzadeh M, Unver T (2014) Genome-wide analysis of the bZIP transcription factors in cucumber. PLoS ONE 9(4):e96014
Caraux G, Pinloche S (2005) PermutMatrix: a graphical environment to arrange gene expression profiles in optimal linear order. Bioinformatics 21(7):1280–1281. https://doi.org/10.1093/bioinformatics/bti141
Chaves M, Zarrouk O, Francisco R, Costa J, Santos T, Regalado A, Rodrigues M, Lopes C (2010) Grapevine under deficit irrigation: hints from physiological and molecular data. Ann Bot 105(5):661–676
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2(4):953–971
FAO (2014) FAOSTAT. Food and Agriculture Organization of the United Nations, Rome, Italy
Fernandez-Silva P, Martinez-Azorin F, Micol V, Attardi G (1997) The human mitochondrial transcription termination factor (mTERF) is a multizipper protein but binds to DNA as a monomer, with evidence pointing to intramolecular leucine zipper interactions. EMBO J 16(5):1066–1079
Ghan R, Van Sluyter SC, Hochberg U, Degu A, Hopper DW, Tillet RL, Schlauch KA, Haynes PA, Fait A, Cramer GR (2015) Five omic technologies are concordant in differentiating the biochemical characteristics of the berries of five grapevine (Vitis vinifera L.) cultivars. BMC Genom 16(1):946
Goodstein DM, Shu SQ, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi chuan = Hereditas/Zhongguo yi chuan xue hui bian ji 29(8):1023–1026
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35(Web Server issue):W585–W587. https://doi.org/10.1093/nar/gkm259
Jefferys BR, Kelley LA, Sternberg MJ (2010) Protein folding requires crowd control in a simulated cell. J Mol Biol 397(5):1329–1338
Kim M, Lee U, Small I, des Francs-Small CC, Vierling E (2012) Mutations in an Arabidopsis mitochondrial transcription termination factor-related protein enhance thermotolerance in the absence of the major molecular chaperone HSP101. Plant Cell 24(8):3349–3365
Kleine T (2012) Arabidopsis thaliana mTERF proteins: evolution and functional classification. Front Plant Sci 3:233
Kushwaha H, Gupta S, Singh VK, Rastogi S, Yadav D (2011) Genome wide identification of Dof transcription factor gene family in sorghum and its comparative phylogenetic analysis with rice and Arabidopsis. Mol Biol Rep 38(8):5037–5053. https://doi.org/10.1007/s11033-010-0650-9
Letunic I, Bork P (2011) Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478
Lijavetzky D, Carbonero P, Vicente-Carbajosa J (2003) Genome-wide comparative phylogenetic analysis of the rice and Arabidopsis Dof gene families. BMC Evol Biol 3:17. https://doi.org/10.1186/1471-2148-3-17
Luo M, Gao Z, Li H, Li Q, Zhang C, Xu W, Song S, Ma C, Wang S (2018) Selection of reference genes for miRNA qRT-PCR under abiotic stress in grapevine. Sci Rep 8(1):4444
Lynch M, Conery JS (2003) The evolutionary demography of duplicate genes. J Struct Funct Genom 3(1–4):35–44
Meskauskiene R, Würsch M, Laloi C, Vidi PA, Coll NS, Kessler F, Baruah A, Kim C, Apel K (2009) A mutation in the Arabidopsis mTERF-related plastid protein SOLDAT10 activates retrograde signaling and suppresses 1O2-induced cell death. Plant J 60(3):399–410
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628. https://doi.org/10.1038/nmeth.1226
Pagliarani C, Vitali M, Ferrero M, Vitulo N, Incarbone M, Lovisolo C, Valle G, Schubert A (2017) Accumulation of microRNAs differentially modulated by drought is affected by grafting in grapevine. Plant Physiol 173:2180–2195
Quesada V, Sarmiento-Mañús R, González-Bayón R, Hricová A, Pérez-Marcos R, Graciá-Martínez E, Medina-Ruiz L, Leyva-Díaz E, Ponce MR, Micol JL (2011) Arabidopsis RUGOSA2 encodes an mTERF family member required for mitochondrion, chloroplast and leaf development. Plant J 68(4):738–753
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33(Web Server issue):W116–W120. https://doi.org/10.1093/nar/gki442
Roberti M, Bruni F, Polosa PL, Manzari C, Gadaleta MN, Cantatore P (2006) MTERF3, the most conserved member of the mTERF-family, is a modular factor involved in mitochondrial protein synthesis. Biochim Biophys Acta (BBA) Bioenerg 1757(9):1199–1206
Roberti M, Polosa PL, Bruni F, Manzari C, Deceglie S, Gadaleta MN, Cantatore P (2009) The MTERF family proteins: mitochondrial transcription regulators and beyond. Biochim Biophys Acta (BBA) Bioenerg 1787(5):303–311
Robles P, Micol JL, Quesada V (2012) Unveiling plant mTERF functions. Mol Plant 5(2):294–296
Sazegari S, Niazi A, Ahmadi FS (2015) A study on the regulatory network with promoter analysis for Arabidopsis DREB-genes. Bioinformation 11(2):101
Schönfeld C, Wobbe L, Borgstädt R, Kienast A, Nixon PJ, Kruse O (2004) The nucleus-encoded protein MOC1 is essential for mitochondrial light acclimation in Chlamydomonas reinhardtii. J Biol Chem 279(48):50366–50374
Snyman MC, Solofoharivelo M-C, Souza-Richards R, Stephan D, Murray S, Burger JT (2017) The use of high-throughput small RNA sequencing reveals differentially expressed microRNAs in response to aster yellows phytoplasma-infection in Vitis vinifera cv.‘Chardonnay’. PloS ONE 12(8):e0182629
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78
Yang Z, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17(1):32–43
Yang Z, Gu S, Wang X, Li W, Tang Z, Xu C (2008) Molecular evolution of the CPP-like gene family in plants: insights from comparative genomics of Arabidopsis and rice. J Mol Evol 67(3):266–277. https://doi.org/10.1007/s00239-008-9143-z
Zhang Y (2005) miRU: an automated plant miRNA target prediction server. Nucleic Acids Res 33(Web Server issue):W701–W704. https://doi.org/10.1093/nar/gki383
Zhao Y, Cai M, Zhang X, Li Y, Zhang J, Zhao H, Kong F, Zheng Y, Qiu F (2014) Genome-wide identification, evolution and expression analysis of mTERF gene family in maize. PLoS ONE 9(4):e94126
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no potential conflicts of interest
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
İnal, B., İlhan, E., Büyük, İ. et al. Transcriptome wide characterization of water deficit responsive grape mTERF transcription. J. Plant Biochem. Biotechnol. 29, 102–113 (2020). https://doi.org/10.1007/s13562-019-00520-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-019-00520-4