Abstract
Cystic fibrosis (CF) is an autosomal recessive disorder caused by mutations in the gene that encodes the cystic fibrosis transmembrane conductance regulator (CFTR) protein. The vast majority of the mortality is due to progressive lung disease. Targeted and untargeted CF breath metabolomics investigations via exhaled breath condensate (EBC) analyses have the potential to expose metabolic alterations associated with CF pathology and aid in assessing the effectiveness of CF therapies. Here, transmission-mode direct analysis in real time traveling wave ion mobility spectrometry time-of-flight mass spectrometry (TM-DART-TWIMS-TOF MS) was tested as a high-throughput alternative to conventional direct infusion (DI) electrospray ionization (ESI) and atmospheric pressure chemical ionization (APCI) methods, and a critical comparison of the three ionization methods was conducted. EBC was chosen as the noninvasive surrogate for airway sampling over expectorated sputum as EBC can be collected in all CF subjects regardless of age and lung disease severity. When using pooled EBC collected from a healthy control, ESI detected the most metabolites, APCI a log order less, and TM-DART the least. TM-DART-TWIMS-TOF MS was used to profile metabolites in EBC samples from five healthy controls and four CF patients, finding that a panel of three discriminant EBC metabolites, some of which had been previously detected by other methods, differentiated these two classes with excellent cross-validated accuracy.

ᅟ
Similar content being viewed by others
Introduction
Exhaled breath condensate (EBC) consists of aerosolized epithelial lining fluid (ELF) containing volatile and nonvolatile compounds trapped and diluted by water vapor condensation [1, 2]. Increasingly, EBC is being used as a noninvasive means for probing pathophysiological processes occurring within the lung [1–3]. For pulmonary diseases, such as asthma, chronic obstructive pulmonary disease (COPD), and cystic fibrosis (CF), the chemical composition of EBC is systematically altered [3, 4]. Targeted and untargeted breath metabolomics approaches are therefore useful for EBC profiling in an effort to identify markers of airway inflammation, characterize pulmonary disease states, and yield a better understanding of disease pathophysiology [5–12].
Mass spectrometry (MS) and nuclear magnetic resonance (NMR) spectroscopy are typically utilized in EBC metabolomics experiments owing to their sensitivity and/or selectivity [10, 13–16]. In general, MS provides greater sensitivity than NMR [13], lending itself to the detection of ELF metabolites that are 103- to 104-fold diluted in EBC [17]. MS-based approaches often rely on gas- or liquid-chromatography (GC, LC) separations, requiring run times in the tens of min. Direct infusion (DI) electrospray ionization (ESI) or atmospheric pressure chemical ionization (APCI) methods can be alternatively used to speed up analysis [18], but often suffer from limited peak capacity, the inability to distinguish overlapping compounds in crowded spectra, and ionization suppression.
The combination of DI methods with ion mobility spectrometry (IMS) is generally appealing for its ability to simplify spectra, increase signal to noise ratio by eliminating chemical noise, produce cleaner MS/MS data even when precursor ion co-selection occurs, and separate closely related compounds such as isobars on a millisecond timescale. Various IMS techniques are currently available, including drift tube IMS (DTIMS), differential mobility spectrometry (DMS), and traveling wave ion mobility spectrometry (TWIMS), each offering varying degrees of separation power and ion focusing capabilities [19]. In combination with MS, these mobility techniques have found applications in proteomics [20], glycomics [21], clinical analysis [22], and metabolomics [23].
Direct analysis in real time (DART), first reported by Cody et al. [24], is an open air, direct sampling plasma ionization technique capable of high-throughput analysis of solids, liquids, or gases. It has been coupled to stand alone DTIMS [25, 26], and TWIMS-MS [27], but not to stand alone DMS or DMS-MS. In DART-MS, a heated gas stream of metastable atomic or molecular species, typically He or N2, is directed at a sample placed within the ionization region. Thermally desorbed analytes are then ionized via a number of gas-phase mechanisms [28–30], and subsequently suctioned into the mass spectrometer. Fluid dynamics play a critical role in DART-MS, determining not only the extent of ion transmission efficiency [31], but also its reproducibility [32]. To stabilize detrimental fluid dynamic effects, the transmission mode (TM)-DART geometry, in which a sample is directly deposited on a stainless steel mesh in a flow-through fashion, has been proposed [33]. This approach has been shown to be superior to probe DART approaches in terms of precision [34].
Building on previous results where two LC-MS methods were compared with DI-TWIMS-MS to quantify expected EBC glucose levels in healthy controls and patients with CF [35], we here describe TM-DART-TWIMS time-of-flight (TOF) MS as a feasible approach for rapid, high-throughput untargeted EBC metabolomics studies in CF, with the goal of expanding the toolbox available for exposing metabolic alterations associated with CF disease pathology and aiding in assessing CF therapy effectiveness [36]. The EBC metabolome coverage yielded by DART is compared with ESI and APCI approaches in DI mode, and metabolomics experiments performed on EBC samples from healthy controls and CF patients are presented.
Materials and Methods
Chemicals
Omnisolv LC-MS grade acetonitrile was purchased from EMD (Billerica, MA, USA). Ultrapure water with 18.2 MΩ cm resistivity (Thermo Scientific Barnstead Nanopure UV ultrapure water system, Marietta, OH, USA) was used in all sample preparation protocols.
Exhaled Breath Condensate (EBC) Sample Collection and Preparation
Samples were collected by the CF Discovery Core, part of the Emory+Children’s Center for CF and Airways Disease Research, under the guidelines approved by the Emory University Institutional Review Boards (approval number IRB00000372) and the Georgia Institute of Technology.
EBC for Method Development and Comparison
EBC from a healthy volunteer was collected using an R-Tube collector (Respiratory Research, Inc., Austin, TX, USA), pooled to obtain a single, homogenous sample for method development, and kept frozen at −80 °C until processed. Prior to analysis, EBC was thawed, and 2-mL aliquots of the sample were placed into separate vials, stored at −80 °C for a minimum of 2 h, and freeze-dried overnight using a VirTis Genesis 25EL lyophilizer (SP Industries, Stone Ridge, NY, USA) according to the program detailed in Supplementary Table S-1. The lyophilized residues were reconstituted without derivatization in 100 μL of acetonitrile:water 80:20 v/v (concentration factor = 20) and recombined to make a single, homogenous EBC concentrate. This concentrate was then comparatively analyzed by both TM-DART-TWIMS-MS and DI ESCi-TWIMS-MS. Three technical replicates were performed for each method.
EBC of Healthy Controls and Cystic Fibrosis Patients
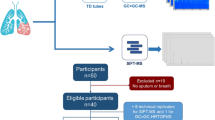
EBC samples collected from five healthy controls and four patients diagnosed with CF were processed individually according to the procedure described above. These samples were analyzed by TM-DART-TWIMS-MS in the negative ion mode to test the applicability of this technique to CF metabolomics investigations. Two technical replicates were performed for each sample.
Transmission-Mode Direct Analysis in Real-Time (TM-DART)
A DART SVP 100 ion source and transmission module (IonSense, Saugus, MA, USA) were used to conduct TM-DART experiments at a 1 L min–1 He gas flow rate. The discharge gas was heated to either 250 °C (positive mode) or 300 °C (negative mode). A custom-built flange and gas-ion separator tube (GIST), connected to a Vacuubrand 2C diaphragm pump (Vacuubrand, Wertheim, Germany), were used to couple the DART ion source to the mass spectrometer in order to reduce the amount of gas flowing into the atmospheric pressure inlet. The exit grid voltage was set at 300 V for both ion polarities. Individual 1-cm stainless steel mesh discs were placed within the 10-position transmission module, and 4 μL of sample (e.g., solvent blank or EBC concentrate) were deposited in the center of the exposed mesh area and allowed to dry for approximately 5 min. A blank mesh was used between sample runs to minimize cross-contamination. An automated software method was used to introduce the transmission module into the DART ionization region at 10 mm s–1, and each position was held within the DART ionization region for 2 min. To avoid disturbing the DART ionizing gas stream, a 1 s hold time was used when the transmission module was advanced to the next position. These experiments generated a transient peak-shaped chronogram with a typical FWHM of approximately 6 s for positive ion mode and between 1 and 2 s for negative ion mode.
Direct Infusion Electrospray Chemical Ionization (DI-ESCi)
An ESCi multi-mode ion source (Waters Corporation, Manchester, UK) was used for high-speed switching between ESI and APCI within the same analytical run. Polarity specific ion source parameters were as follows: capillary voltage: 3 kV(+)/2.2 kV(–); corona current: 15 μA(+)/20 μA(–). All other source parameters were as follows: sampling cone voltage 35 V, source temperature 120 °C, desolvation temperature 250 °C, and nitrogen desolvation gas flow rate 650 L h–1. Samples were introduced into the ESCi source using direct infusion at a flow rate of 2 μL min–1, and each run was acquired for 2 min.
Traveling Wave Ion Mobility Spectrometry Time-of-Flight Mass Spectrometry (TWIMS-TOF MS)
TWIMS-TOF MS was performed on a Synapt G2 High Definition Mass Spectrometry system (Waters Corporation, Manchester, UK), a hybrid quadrupole-ion mobility-TOF mass spectrometer with a typical resolving power of 20,000 m/∆m (FWHM), and mass accuracy of 9 ppm at m/z 554.2615. Initial TWIMS-TOF MS experiments (data not shown) were performed using a control EBC concentrate sample to optimize ion source parameters for maximum signal-to-noise ratio, and TWIMS parameters for ion separation, in both positive and negative ion modes. Optimized TWIMS parameters were as follows: wave height ramped between 10 and 40 V, wave velocity ramped between 200 and 800 m s–1 in positive ion mode, and between 400 and 1000 m s–1 in negative ion mode, IMS gas flow rate 95 mL min-1 in positive ion mode and 40 mL min-1 in negative ion mode, and a helium gas flow rate 180 mL min–1. The mass spectrometer was calibrated across the m/z 50–1000 range using a 0.5 mM sodium formate solution prepared in 90:10 (v/v) 2-propanol:water. An option within the MS acquisition method was selected to add a drift time function, which would contain mobility total ion current chronogram data for each data file. Raw data were examined using MassLynx ver. 4.1 (Waters Corp., Milford, MA, USA).
Mass Spectral Feature Extraction
Mass spectral features were extracted from TWIMS-TOF MS data as (drift time, m/z) “compounds” using Progenesis QI ver. 2.0 (Nonlinear Dynamics, Waters Corp.). Although originally designed for mining chromatographic data, the acquired TWIMS-TOF MS data could be similarly processed by importing the drift time function of each raw data file generated by the TWIMS-MS system. The feature extraction workflow consisted of mass detection followed by drift time alignment, peak picking, integration, and deconvolution to group together adducts derived from the same compound. In all cases, EBC sample data were curated against corresponding solvent blank data to determine EBC-specific features, filtering out signals with peak areas less than or equal to two times of those present in solvent blanks.
Multivariate Analysis for CF Metabolomics Investigations
The resulting matrix of TM-DART-TWIMS-MS spectral features obtained from five healthy controls and four CF patients was normalized after blank correction. Subsequently, m/z values of all extracted features were searched in the Human Metabolome Database (HMDB) [37] with a mass error window of 10 mDa, and only those that had candidates with endogenous human and/or microbial origins were retained. The remaining features were utilized to build a model for class discrimination via orthogonal partial least squares-discriminant analysis (oPLS-DA) [38] (MATLAB, R2015a, The MathWorks, Natick, MA, USA with PLS-Toolbox, ver. 8.0, Eigenvector Research, Inc., Manson, WA, USA). Reverse interval PLS-DA (iPLS-DA) was applied to autoscaled data to select an optimum panel of discriminant features and number of latent variables (LVs) that maximized the classification accuracy. The iPLS-DA interval size was set to 1 and the maximum number of LVs set to 6. Leave-one-out cross-validation (LOOCV) was used for oPLS-DA model building.
Results and Discussion
TM-DART-TWIMS-TOF MS Optimization
Parameters to be optimized in TM-DART typically include source-to-sample distance and DART gas temperature [33]. Previous work had shown that the greatest sensitivity was obtained when the DART gas outlet and the sample were in close proximity to one another [34]. A similar effect was observed in these experiments and, therefore, this parameter was not investigated further, with all TM-DART experiments using a DART-to-sample distance of ≤1 mm, the minimum possible. The plasma gas temperature was optimized in negative ion mode, having an important effect on the number of detected EBC spectral features, as illustrated in Supplementary Figure S-1. A compromise between sensitivity and number of detected features was achieved by using a high enough temperature for fast analyte desorption while still minimizing thermal ion fragmentation. The average number of detected features (after blank correction) increased when the gas temperature was augmented from 250 to 300 °C, with more reproducible results at 300 °C; beyond 300 °C the number of detected features decreased, and therefore 300 °C was used in all subsequent experiments.
Comparison of TM-DART and DI- ESCi MS for EBC Analysis
TM-DART and DI-ESCi were compared to investigate if unique and/or complementary EBC metabolome coverage was produced from these different methods. Both positive and negative ion mass spectra of EBC from a healthy volunteer were acquired with both techniques, yielding different mass spectral patterns.
Figure 1 shows 2D drift time versus m/z plots in positive and negative ion modes corresponding to the compounds detected after data mining and blank correction. TM-DART (Figure 1a and b) resulted in the detection of only singly charged EBC metabolites for both ion polarities, most of them grouped in compact trend lines. DI-ESI produced a comparatively higher number of compounds (Figure 1c and d) than DART (~23- and 83-fold higher for positive and negative ion modes, respectively), most likely associated with this technique’s characteristic production of both singly and multiply charged ions, as well as the ability to ionize very polar and nonvolatile species. The ESCi APCI mode (Figure 1e and f) yielded ~3- and 4-fold more EBC-specific features than DART, but ~7- and 21-fold less than ESI, in positive and negative ion modes, respectively.
Drift time versus m/z plots for datasets obtained with different techniques from a healthy volunteer’s EBC. Data were blank-corrected, de-isotoped, and corrected for multiple adducts. (a) TM-DART (+): 106 compounds; (b) TM-DART (–): 31 compounds; (c) ESI (+): 2449 compounds; (d) ESI (–): 2559 compounds; (e) APCI (+): 357 compounds; and (f) APCI (–) 122 compounds
Despite the fact that DART makes use of ionization mechanisms that predominantly follow APCI-like pathways [28, 30], the 2D drift time versus m/z plots of TM-DART appeared different from those of DI-APCI for both ion polarities. As indicated above, the ESCi APCI function produced more ions than DART in both ion modes, producing a few multiply charged ions in positive ion mode (Figure 1e), and predominantly singly charged ions in negative ion mode (Figure 1f). DART only produced singly charged species in both positive and negative ionization modes (Figure 1a and b). These differences may arise from the way in which the specific dual ESCi ion source operates, i.e., the desolvation parameters (temperature and nitrogen gas flow rate) used in ESCi were unchanged during the alternating switching between ESI and APCI modes. Therefore, analytes may not be exposed to true APCI-like conditions where they are fully desolvated and vaporized at a higher temperature than that utilized in the dual ion source.
The data shown in Figure 1 were further analyzed to quantitatively determine the number of unique and overlapping features between DART, ESI, and APCI, as shown in Figure 2. For the three ionization methods, a total of 2686 and 2619 EBC-specific compounds were detected for positive and negative ion modes, respectively, evidencing the capability of EBC metabolomics approaches to profile airway secretions. Not surprisingly, ESI accounted for a large number of species, detecting 91% of the total compounds in positive ion mode, and 98% in negative ion mode. In contrast, APCI produced about 13% and 5%, and DART detected about 4% and 1% of the total number of compounds in positive and negative ion modes, respectively. Many of the EBC signals produced using DART were removed after blank (background) correction, and thus the number of DART-detected compounds was much lower, overall.
It is noted that there are numerous unique compounds produced by only one ionization method (Figure 2). The proportions of unique compounds to total compounds detected by any of the three ionization methods are 78% and 90% for DART positive and negative ion modes, 91% and 96% for ESI positive and negative ion modes, and 40% and 26% for APCI positive and negative ion modes, respectively. The proportions of compounds detected by APCI overlapping with those detected by ESI reached 57% and 74% for positive and negative ion modes, respectively. However, the compounds detected by DART had a much smaller overlap with the other two ionization methods, with 10% for both positive and negative ESI, and 13% and 0% with positive and negative mode APCI, respectively. These results suggest that although DART produced a lower number of features; these were rather unique and may be an important complement to exhaled breath condensate metabolomics experiments performed by either ESI or APCI.
To explore the signal abundances observed for the three different ionization methods, scatter plots of average peak areas of overlapping compounds were created (Supplementary Figures S-2 to S-11). In general, for compounds overlapping between ESI and APCI, signal intensities were higher in ESI than in APCI for both positive and negative ion modes (Supplementary Figures S-2 to S-6; S-9 and S-10) with a median increase from APCI to ESI of 26-fold for positive ion mode and 64-fold for negative ion mode. For compounds overlapping between ESI and DART, there was no obvious trend in the signal intensities (Supplementary Figures S-7 and S-11). For overlapping compounds between APCI and DART in positive ion mode, signal intensities were higher in DART than in APCI, with a median increase of 6-fold from APCI to DART (Supplementary Figure S-8).
Relative standard deviations (RSDs) of overlapping compounds in Figure 2 were calculated in order to compare the precision of each method. The results are shown as box plots in Supplementary Figure S-12. In general, ESI had a higher precision than APCI, which had a higher precision than DART. For overlapping compounds between ESI and APCI in positive ion mode, the median RSDs were 7% and 12%, respectively; and 6% and 15% in negative ion mode, respectively. For overlapping compounds between ESI and DART, the median RSD was 8% for both ionization methods in positive ion mode; and 9% and 17% in negative ion mode, respectively. The median RSDs of overlapping compounds between APCI and DART in positive ion mode were 7% and 10%, respectively.
Regarding analysis time, DART experiments presented the advantage that they did not require rinsing the tubing that connected the pump propelling the liquid samples to the ion source as in DI-ESCi, which was translated into higher-throughput sample analysis. In addition, the fact that no wetted tubing is involved in DART, also made this technique much more resistant to memory effects, minimizing carryover.
CF Sample Analysis and Multivariate Classification
The ability of TM-DART-TWIMS-TOF MS to rapidly acquire metabolic profiles from EBC was tested to investigate if the metabolic differences between four CF patients and five healthy individuals could be rapidly established using this approach. Negative ion mode was chosen because the EBC pH in CF patients is known to be lower than in healthy controls [39], suggesting that acidic metabolites may be important discriminant species. Overall, metabolic profile acquisition required less than 20 min per sample following lyophilization, considering ~5 min for sample reconstitution, 5–10 min for sample deposition and drying, and ~4 min per sample for replicate DART analysis. This last step is ~5–15 times faster than a typical 10–30 min LC-MS run. Representative data for two EBC samples from a selected CF patient and a healthy subject are shown in Figures 3a and b, respectively, depicting mass spectra in the range of m/z 50–400. Some differences could be readily observed. A feature with m/z 128.0382 (marked with an asterisk) for example, showed much higher abundance in the CF patient sample than in the healthy control sample. A total of 29 metabolic features were extracted from the acquired TM-DART metabolic profiles. Of these features, 11 had candidates with endogenous human and/or microbial origins in HMDB, which were subjected to multivariate classification.
Negative ion TM-DART-TWIMS-TOF mass spectrum from (a) a sample from a CF patient, and (b) a sample from a healthy control subject (inset shows a zoomed in view of the m/z 120–140 range). The asterisk indicates the spectral peak at m/z 128.0382. (c) Extracted ion mobility chronograms for the best three discriminant features from the CF patient sample illustrated in (a)
Figure 4a and b show the results of multivariate classification between the four samples from CF patients and the five samples from healthy controls. A set of three metabolic features and two LVs was selected through the reverse iPLS-DA feature selection process. The resulting oPLS-DA model (Figure 4a) yielded 100% cross-validated accuracy, sensitivity, and specificity. This model captured 70.7% and 96.9% of the X- and Y-block variances, respectively.
oPLS-DA model for discrimination of CF patient samples (red circles) from healthy control samples (black squares). (a) Cross-validated prediction plot using the three discriminant metabolic feature panel obtained from iPLS-DA variable selection. (b) oPLS-DA calibration scores plot for (a). The model consisted of 2 LVs with 70.7% and 96.9% total captured X- and Y-block variances, respectively. The accuracy, sensitivity, and specificity were all 100%
Figure 3c shows the TM-DART extracted ion mobility chronograms for the three discriminant metabolic features used in the oPLS-DA model, which vary in relative abundances. Metabolites in the 3-feature panel were tentatively identified (Table 1) based on accurate mass measurements and database searches. Different metabolites were separated in drift time in TWIMS based on their m/z and structure types. Feature #1 had a mean fold increase of 17.4 from healthy control to CF samples, and was tentatively identified as pyroglutamic acid or other of its structural isomers. Interestingly, in a metabolomics study by Joseloff et al. [40], pyroglutamic acid was identified as an important metabolite responsible for distinguishing 31 CF from 31 non-CF serum samples from children, with a CF to non-CF fold change of 1.2. A recent study by our team also identified this metabolite as increased in CF patients before an acute pulmonary exacerbation [41]. Feature #2 had tentative identifications matching short chain carboxylic acids (dimethylmalonic acid, 2-acetolactate, glutaric acid, monoethyl malonic acid, ethylmalonic acid, and methylsuccinic acid, amino acids/peptides (glycyl-glycine and asparagine), and intermediates in amino acid (N-carbamoylsarcosine) or uracil (ureidopropionic acid) metabolic pathways. Feature #3 had a mean fold increase of 14.2 from healthy control to CF samples. It was potentially assigned to several hydroxyeicosatetraenoic acids (HETEs) (5-, 8-, 9-, 11-, 12-, 15-, 16-, 17-, 18-, 19-, and 20-HETE) and epoxyeicosatrienoic acids (EETs) (5,6-, 8,9-, 11,12-, and 14,15-EET) involved in arachidonic acid metabolism. Interestingly, an increased arachidonic acid ratio has been reported in bronchial phospholipids in CF patients compared with normal controls, suggesting abnormal arachidonic acid metabolism in CF patients [42].
Conclusions
TWIMS-TOF MS for CF untargeted EBC metabolomics studies is demonstrated as a potential high-throughput alternative to conventional LC-MS-based methods typically used in present investigations. It was found that the EBC metabolome coverage provided by DART ionization is an important complement to ESI and APCI-based methods. The metabolites detected by DART can provide biochemical information pertinent in metabolomics applications for studying pathophysiologic processes occurring within the lung. The analysis of a small set of EBC samples from a cohort of healthy individuals and CF patients shows the initial use of DART in metabolomics studies. Multivariate analyses of the resulting TM-DART-TWIMS-TOF MS datasets can successfully discriminate between EBC samples from CF patients and healthy controls.
References
Hunt, J.: Exhaled breath condensate: an evolving tool for noninvasive evaluation of lung disease. J. Allergy Clin. Immunol. 110, 28–34 (2002)
Mutlu, G.M., Garey, K.W., Robbins, R.A., Danziger, L.H., Rubinstein, I.: Collection and analysis of exhaled breath condensate in humans. Am. J. Respir. Crit. Care Med. 164, 731–737 (2001)
Montuschi, P., Barnes, P.J.: Analysis of exhaled breath condensate for monitoring airway inflammation. Trends Pharmacol. Sci. 23, 232–237 (2002)
Grob, N.M., Aytekin, M., Dweik, R.A.: Biomarkers in exhaled breath condensate: a review of collection, processing, and analysis. J. Breath Res. 2, 18 (2008)
Carraro, S., Giordano, G., Reniero, F., Carpi, D., Stocchero, M., Sterk, P.J., Baraldi, E.: Asthma severity in childhood and metabolomic profiling of breath condensate. Allergy 68, 110–117 (2013)
Carraro, S., Rezzi, S., Reniero, F., Heberger, K., Giordano, G., Zanconato, S., Guillou, C., Baraldi, E.: Metabolomics applied to exhaled breath condensate in childhood asthma. Am. J. Respir. Crit. Care Med. 175, 986–990 (2007)
Esther, C., Jasin, H.M., Collins, L.B., Swenberg, J.A., Boysen, G.: A mass spectrometric method to simultaneously measure a biomarker and dilution marker in exhaled breath condensate. Rapid Commun. Mass Spectrom. 22, 701–705 (2008)
Montuschi, P.: Exhaled breath condensate analysis in patients with COPD. Clin. Chim. Acta 356, 22–34 (2005)
Montuschi, P., Paris, D., Melck, D., Lucidi, V., Ciabattoni, G., Raia, V., Calabrese, C., Bush, A., Barnes, P.J., Motta, A.: NMR spectroscopy metabolomic profiling of exhaled breath condensate in patients with stable and unstable cystic fibrosis. Thorax 67, 222–228 (2012)
Nobakht, M.G.B.F., Aliannejad, R., Rezaei-Tavirani, M., Taheri, S., Oskouie, A.A.: The metabolomics of airway diseases, including COPD, asthma, and cystic fibrosis. Biomarkers 20, 5–16 (2015)
Snowden, S., Dahlen, S.E., Wheelock, C.E.: Application of metabolomics approaches to the study of respiratory diseases. Bioanalysis 4, 2265–2290 (2012)
Wheelock, C.E., Goss, V.M., Balgoma, D., Nicholas, B., Brandsma, J., Skipp, P.J., Snowden, S., Burg, D., D’Amico, A., Horvath, I., Chaiboonchoe, A., Ahmed, H., Ballereau, S., Rossios, C., Chung, K.F., Montuschi, P., Fowler, S.J., Adcock, I.M., Postle, A.D., Dahlen, S.E., Rowe, A., Sterk, P.J., Auffray, C., Djukanovic, R., Grp, U.B.S.: Application of ‘omics technologies to biomarker discovery in inflammatory lung diseases. Eur. Resp. J. 42, 802–825 (2013)
Dettmer, K., Aronov, P.A., Hammock, B.D.: Mass spectrometry-based metabolomics. Mass Spectrom. Rev. 26, 51–78 (2007)
Lenz, E.M., Wilson, I.D.: Analytical strategies in metabonomics. J. Proteome Res. 6, 443–458 (2007)
Sofia, M., Maniscalco, M., de Laurentiis, G., Paris, D., Melck, D., Motta, A.: Exploring airway diseases by NMR-based metabonomics: a review of application to exhaled breath condensate. J. Biomed. Biotechnol. 7, 2011: 403260 (2011)
Wishart, D.S.: Quantitative metabolomics using NMR. TrAC Trends Anal. Chem. 27, 228–237 (2008)
Effros, R.M., Hoagland, K.W., Bosbous, M., Castillo, D., Foss, B., Dunning, M., Gare, M., Lin, W., Sun, F.: Dilution of respiratory solutes in exhaled condensates. Am. J. Respir. Crit. Care Med. 165, 663–669 (2002)
Draper, J., Lloyd, A.J., Goodacre, R., Beckmann, M.: Flow infusion electrospray ionisation mass spectrometry for high throughput, non-targeted metabolite fingerprinting: a review. Metabolomics 9, S4–S29 (2013)
May, J.C., McLean, J.A.: Ion mobility-mass spectrometry: time-dispersive instrumentation. Anal. Chem. 87, 1422–1436 (2015)
McLean, J.A., Ruotolo, B.T., Gillig, K.J., Russell, D.H.: Ion mobility-mass spectrometry: a new paradigm for proteomics. Int. J. Mass Spectrom. 240, 301–315 (2005)
Taraszka, J.A., Counterman, A.E., Clemmer, D.E.: Gas-phase separations of complex tryptic peptide mixtures. Fresenius J. Anal. Chem. 369, 234–245 (2001)
Chouinard, C.D., Wei, M.S., Beekman, C.R., Kemperman, R.H.J., Yost, R.A.: Ion mobility in clinical analysis: current progress and future perspectives. Clin. Chem. 62, 124–133 (2016)
Paglia, G., Williams, J.P., Menikarachchi, L., Thompson, J.W., Tyldesley-Worster, R., Halldorsson, S., Rolfsson, O., Moseley, A., Grant, D., Langridge, J., Palsson, B.O., Astarita, G.: Ion mobility derived collision cross-sections to support metabolomics applications. Anal. Chem. 86, 3985–3993 (2014)
Cody, R.B., Laramee, J.A., Durst, H.D.: Versatile new ion source for the analysis of materials in open air under ambient conditions. Anal. Chem. 77, 2297–2302 (2005)
Harris, G.A., Kwasnik, M., Fernandez, F.M.: Direct analysis in real time coupled to multiplexed drift tube ion mobility spectrometry for detecting toxic chemicals. Anal. Chem. 83, 1908–1915 (2011)
Keelor, J.D., Dwivedi, P., Fernandez, F.M.: An effective approach for coupling direct analysis in real time with atmospheric pressure drift tube ion mobility spectrometry. J. Am. Soc. Mass Spectrom. 25, 1538–1548 (2014)
Rasanen, R.M., Dwivedi, P., Fernandez, F.M., Kauppila, T.J.: Desorption atmospheric pressure photoionization and direct analysis in real time coupled with traveling wave ion mobility mass spectrometry. Rapid Commun. Mass Spectrom. 28, 2325–2336 (2014)
McEwen, C.N., Larsen, B.S.: Ionization mechanisms related to negative ion APPI, APCI, and DART. J. Am. Soc. Mass Spectrom. 20, 1518–1521 (2009)
Song, L.G., Dykstra, A.B., Yao, H.F., Bartmess, J.E.: Ionization mechanism of negative ion-direct analysis in real time: a comparative study with negative ion-atmospheric pressure photoionization. J. Am. Soc. Mass Spectrom. 20, 42–50 (2009)
Song, L.G., Gibson, S.C., Bhandari, D., Cook, K.D., Bartmess, J.E.: Ionization mechanism of positive-ion direct analysis in real time: a transient microenvironment concept. Anal. Chem. 81, 10080–10088 (2009)
Harris, G.A., Fernandez, F.M.: Simulations and experimental investigation of atmospheric transport in an ambient metastable-induced chemical ionization source. Anal. Chem. 81, 322–329 (2009)
Harris, G.A., Falcone, C.E., Fernandez, F.M.: Sensitivity “hot spots” in the direct analysis in real time mass spectrometry of nerve agent simulants. J. Am. Soc. Mass Spectrom. 23, 153–161 (2012)
Perez, J.J., Harris, G.A., Chipuk, J.E., Brodbelt, J.S., Green, M.D., Hampton, C.Y., Fernandez, F.M.: Transmission-mode direct analysis in real time and desorption electrospray ionization mass spectrometry of insecticide-treated bednets for malaria control. Analyst 135, 712–719 (2010)
Jones, C., Fernandez, F.M.: Transmission mode direct analysis in real time mass spectrometry for fast untargeted metabolic fingerprinting. Rapid Commun. Mass Spectrom. 27, 1311–1318 (2013)
Monge, M.E., Perez, J.J., Dwivedi, P., Zhou, M.S., McCarty, N.A., Stecenko, A.A., Fernandez, F.M.: Ion mobility and liquid chromatography/mass spectrometry strategies for exhaled breath condensate glucose quantitation in cystic fibrosis studies. Rapid Commun. Mass Spectrom. 27, 2263–2271 (2013)
Wetmore, D.R., Joseloff, E., Pilewski, J., Lee, D.P., Lawton, K.A., Mitchell, M.W., Milburn, M.V., Ryals, J.A., Guo, L.: Metabolomic profiling reveals biochemical pathways and biomarkers associated with pathogenesis in cystic fibrosis cells. J. Biol. Chem. 285, 30516–30522 (2010)
Wishart, D.S., Jewison, T., Guo, A.C., Wilson, M., Knox, C., Liu, Y., Djoumbou, Y., Mandal, R., Aziat, F., Dong, E., Bouatra, S., Sinelnikov, I., Arndt, D., Xia, J., Liu, P., Yallou, F., Bjorndahl, T., Perez-Pineiro, R., Eisner, R., Allen, F., Neveu, V., Greiner, R., Scalbert, A.: HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Res. 41, D801–D807 (2013)
Trygg, J., Wold, S.L.: Orthogonal projections to latent structures (O-PLS). J. Chemom. 16, 119–128 (2002)
Ojoo, J.C., Mulrennan, S.A., Kastelik, J.A., Morice, A.H., Redington, A.E.: Exhaled breath condensate pH and exhaled nitric oxide in allergic asthma and in cystic fibrosis. Thorax 60, 22–26 (2005)
Joseloff, E., Sha, W., Bell, S.C., Wetmore, D.R., Lawton, K.A., Milburn, M.V., Ryals, J.A., Guo, L., Muhlebach, M.S.: Serum metabolomics indicate altered cellular energy metabolism in children with cystic fibrosis. Pediatr. Pulmonol. 49, 463–472 (2014)
Zang, X., Monge, M.E., McCarty, N.A., Stecenko, A.A., Fernandez, F.M.: Feasibility of early detection of cystic fibrosis acute pulmonary exacerbations by exhaled breath condensate metabolomics: a pilot study. J. Proteome Res. 16, 550–558 (2017)
Gilljam, H., Strandvik, B., Ellin, A., Wiman, L.G.: Increased mole fraction of arachidonic acid in bronchial phospholipids in patients with cystic fibrosis. Scand. J. Clin. Lab. Invest. 46, 511–518 (1986)
Acknowledgements
This work was supported by an Emory University and Children’s Healthcare of Atlanta Pediatric Research Center Pilot Project to A.A.S. and F.M.F. The authors also thank Amanda Castle and Veranika Pazharskaya at Emory University for invaluable help in collecting EBC samples. They are also thankful for access to MS instrumentation acquired through the NSF/NASA Center for Chemical Evolution. M.E.M. is currently a Research Staff member from CONICET (Consejo Nacional de Investigaciones Científicas y Técnicas, Argentina).
Author information
Authors and Affiliations
Corresponding author
Additional information
Xiaoling Zang and José J. Pérez contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 19343 kb)
Rights and permissions
About this article
Cite this article
Zang, X., Pérez, J.J., Jones, C.M. et al. Comparison of Ambient and Atmospheric Pressure Ion Sources for Cystic Fibrosis Exhaled Breath Condensate Ion Mobility-Mass Spectrometry Metabolomics. J. Am. Soc. Mass Spectrom. 28, 1489–1496 (2017). https://doi.org/10.1007/s13361-017-1660-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-017-1660-9