Abstract
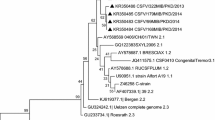
The present study focused on the detection and genetic characterisation of 5′ untranslated region (5′UTR) and E2 gene of classical swine fever virus (CSFV, family Flaviviridae, genus Pestivirus) from bovine population of the northeastern region of India. A total of 134 cattle serum samples were collected from organised cattle farms and were screened for CSFV antigen with a commercial antigen capture enzyme linked immunosorbent assay (Ag-ELISA) and reverse transcription-polymerase chain reaction (RT-PCR). A total of 10 samples were positive for CSFV antigen by ELISA, while all of them were positive in PCR for 5′UTR region. Full length E2 region of CSFV were successfully amplified from two positive samples and used for subsequent phylogenetic analysis and determination of protein 3D structure which showed similarity with reported CSFV isolate from Assam of sub-genogroup 2.1, with minor variations in protein structure.


Similar content being viewed by others
References
Ahuja A, Bhattacharjee U, Chakraborty AK, Karam A, Ghatak S, Puro K, Das S, Shakuntala I, Srivastava N, Ngachan SV, Sen A. Complete genome sequence of classical swine fever virus subgenogroup 2.1 from Assam, India. Genome Announc. 2015. https://doi.org/10.1128/genomea.01437-14.
Asfor AS, Wakeley PR, Drew TW, Paton DJ. Recombinant pestivirus E2 glycoproteins prevent viral attachment to permissive and non-permissive cells with different efficiency. Virus Res. 2014;189:147–57.
Barman N, Gupt R, Bora D, Kataria R, Tiwari A, Roychoudhury P. Molecular characterization of classical swine fever virus involved in the outbreak in Mizoram. Indian J Virol. 2010;21:76–81.
Bett B, Deka R, Padmakumar V, Rajasekhar M. Classical swine fever in northeast India: Prevention and control measures. Policy Brief. Nairobi: International Livestock Research Institute; 2012.
Chakraborty SV, Chakraborty S, Veeregoda B, Rathnamma D, Venkatesha M, Leena G, Veeresh H, Patil S. Molecular characterization and genogrouping of classical swine fever virus isolated from field outbreaks. Indian J Anim Sci. 2011;81:8.
Choori PPS, Rathnamma D, Sharada R, Chandranaik BM, Isloor S, Reddy GB, Geetha S, Rahman H. Prevalence of classical swine fever in Karnataka, India. Vet World. 2015;8:541–4.
Desai G, Sharma A, Kataria R, Barman N, Tiwari A. 5′-UTR-based phylogenetic analysis of classical swine fever virus isolates from India. Acta Virol. 2010;54:79–82.
Dräger C, Beer M, Blome S. Porcine complement regulatory protein CD46 and heparin sulfates are the major factors for classical swine fever virus attachment in vitro. Arch Virol. 2015;160:739–46.
Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39:783–91.
Geoghegan JL, Senior AM, Di Giallonardo F, Holmes EC. Virological factors that increase the transmissibility of emerging human viruses. Proc Natl Acad Sci USA. 2016. https://doi.org/10.1073/pnas.1521582113.
Holmes EC. Evolution and emergence of RNA viruses. Oxford: Oxford University Press; 2009.
Holmes EC, Zhang YZ. The evolution and emergence of hantaviruses. Curr Opin Virol. 2015. https://doi.org/10.1016/j.coviro.2014.12.007.
Hulst MM, Van Gennip HG, Vlot AC, Schooten E, de Smit AJ, Moormann RJ. Interaction of classical swine fever virus with membrane-associated heparan sulfate: role for virus replication in vivo and virulence. J Virol. 2001;75:9585–95.
Jackson AP, Charleston MA. A cophylogenetic perspective of RNA-virus evolution. Mol Biol Evol. 2004. https://doi.org/10.1093/molbev/msg232.
Kimura M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16:111–20.
Kitchen A, Shackelton LA, Holmes EC. Family level phylogenies reveal modes of macroevolution in RNA viruses. Proc Natl Acad Sci USA. 2011. https://doi.org/10.1073/pnas.1011090108.
Kumar S, Stecher G, Tamura K. MEGA7: molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Liang DL, Sainz IF, Ansari IH, Gil LH, Vassilev V, Donis RO. The envelope glycoprotein E2 is a determinant of cell culture tropism in ruminant pestiviruses. J Gen Virol. 2003;84:1269–74.
Maurer K, Krey T, Moennig V, Thiel HJ, Rümenapf T. CD46 is a cellular receptor for bovine viral diarrhea virus. J Virol. 2004;78:1792–9.
McGeoch DJ, Gatherer D. Integrating reptilian herpesviruses into the family herpesviridae. J Virol. 2005. https://doi.org/10.1128/JVI.79.2.725-731.2005.
Nandi S, Muthuchelvan D, Ahuja A, Bisht S, Chander V, Pandey AB, Singh RK. Prevalence of classical swine fever virus in India: a 6-year study (2004–2010). Transbound Emerg Dis. 2011;58:461–3.
Patil SS, Hemadri D, Shankar BP, Raghavendra AG, Veeresh H, Sindhoora B, Chandan S, Sreekala K, Gajendragad MR, Prabhudas K. Genetic typing of recent classical swine fever isolates from India. Vet Microbiol. 2010;141:367–73.
Postel A, Jha V, Schmeiser S, Becher P. First molecular identification and characterization of classical swine fever virus isolates from Nepal. Arch Virol. 2013;158:207–10.
Roy A, Kucukural A, Zhang Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc. 2010;5:725–38.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.
Sandvik T, Patonb David J, Lowingsb Paul J. Detection and identification of ruminant and porcine pestiviruses by nested amplification of 5′ untranslated cDNA regions. J Virol Methods. 1997;64:43–56.
Sarma DK, Mishra N, Vilcek S, Rajukumar K, Behera SP, Nema RK, Dubey P, Dubey SC. Phylogenetic analysis of recent classical swine fever virus (CSFV) isolates from Assam, India. Comp Immunol Microbiol Infect Dis. 2011;34:11–5.
Shimizu M, Kumagai T. Experimental infection of pregnant goats with swine fever virus. Vet Microbiol. 1989;20:207–14.
Thiel HJ, Stark R, Weiland E, Rumenapf T, Meyers G. Hog cholera virus: molecular composition of virions from a pestivirus. J Virol. 1991;65:4705–12.
Van Regenmortel MH, Fauquet C, Bishop D, Carstens E, Estes M, Lemon S, Maniloff J, Mayo M, McGeoch D, Pringle C. Virus taxonomy: classification and nomenclature of viruses. In: Seventh report of the international committee on taxonomy of viruses. Academic Press. 2000.
Villarreal LP, Defilippis VR, Gottlieb KA. Acute and persistent viral life strategies and their relationship to emerging diseases. Virology. 2000. https://doi.org/10.1006/viro.2000.0381.
Acknowledgements
This work was supported by a grant from Department of Biotechnology, Government of India, (DBT-NER/LIVS/11/2012dated24-04-2014, Project ‘‘Advance Animal Disease Diagnosis and Management Consortium’’). We thank all the persons involved in collection of samples from various farms.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chakraborty, A.K., Karam, A., Mukherjee, P. et al. Detection of classical swine fever virus E2 gene in cattle serum samples from cattle herds of Meghalaya. VirusDis. 29, 89–95 (2018). https://doi.org/10.1007/s13337-018-0433-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13337-018-0433-9