Abstract
Background
Uncovering molecular pathogenesis and mechanisms of small cell lung cancer (SCLC) will contribute to SCLC therapy. Multiple studies demonstrated that miR-451a acts as an anti-tumor miRNA in non-small cell lung cancer. However, the mechanism of miR-451a in SCLC was ambiguous.
Objective
We aimed to explore the function of miR-451a in SCLC and decipher the underlying mechanisms.
Methods
TargetScan and dual-luciferase reporter assays were used to analyze the target genes of miR-451a. Cell counting kit-8 and colony formation assays were performed to assess the roles of miR-451a on cell growth. Gene set enrichment analysis (GSEA) was utilized to enrich biological pathways. Western blot was used to measure protein expression.
Results
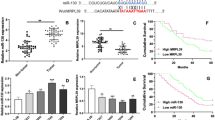
MiR-451a expression was reduced dramatically in SCLC tissues and cell lines (NCI-H1688 and NCI-H446). Helicase, Lymphoid Specific (HELLS) was proved to be a target gene of miR-451a. In addition, cell proliferation assays in SCLC cells transfected with miR-451a mimic and/or HELLS revealed that miR-451a inhibited cell proliferation via targeting HELLS. Moreover, the roles of miR-451a/HELLS in expression of key proteins in mTOR and apoptosis signaling pathways suggested that miR-451a inactivated mTOR and activated apoptosis signaling pathway via directly silencing HELLS.
Conclusions
Our study indicated that miR-451a hinders SCLC cell proliferation in vitro through regulating mTOR and apoptosis signaling pathways via silencing HELLS, suggesting that miR-451a could be a promising tumor suppressor in SCLC. And there is a potential for miR-451a to be a drug target and biomarker for SCLC.





Similar content being viewed by others
References
Chen DQ, Yu C, Zhang XF, Liu ZF, Wang R, Jiang M, Chen H, Yan F, Tao M, Chen LB et al (2018) HDAC3-mediated silencing of miR-451 decreases chemosensitivity of patients with metastatic castration-resistant prostate cancer by targeting NEDD9. Ther Adv Med Oncol 10:1758835918783132. https://doi.org/10.1177/1758835918783132
Christensen LL, Holm A, Rantala J, Kallioniemi O, Rasmussen MH, Ostenfeld MS, Dagnaes-Hansen F, Oster B, Schepeler T, Tobiasen H et al (2014) Functional screening identifies miRNAs influencing apoptosis and proliferation in colorectal cancer. PLoS One 9:e96767. https://doi.org/10.1371/journal.pone.0096767
Dou C, Zhou Z, Xu Q, Liu Z, Zeng Y, Wang Y, Li Q, Wang L, Yang W, Liu Q et al (2019) Hypoxia-induced TUFT1 promotes the growth and metastasis of hepatocellular carcinoma by activating the Ca(2+)/PI3K/AKT pathway. Oncogene 38:1239–1255. https://doi.org/10.1038/s41388-018-0505-8
Goto A, Tanaka M, Yoshida M, Umakoshi M, Nanjo H, Shiraishi K, Saito M, Kohno T, Kuriyama S, Konno H et al (2017) The low expression of miR-451 predicts a worse prognosis in non-small cell lung cancer cases. PLoS One 12:e0181270. https://doi.org/10.1371/journal.pone.0181270
Huang JY, Zhang K, Chen DQ, Chen J, Feng B, Song H, Chen Y, Zhu Z, Lu L, De W et al (2015) MicroRNA-451: epithelial-mesenchymal transition inhibitor and prognostic biomarker of hepatocelluar carcinoma. Oncotarget 6:18613–18630. https://doi.org/10.18632/oncotarget.4317
Minna E, Romeo P, Dugo M, De Cecco L, Todoerti K, Pilotti S, Perrone F, Seregni E, Agnelli L, Neri A et al (2016) miR-451a is underexpressed and targets AKT/mTOR pathway in papillary thyroid carcinoma. Oncotarget 7:12731–12747. https://doi.org/10.18632/oncotarget.7262
Su R, Gong JN, Chen MT, Song L, Shen C, Zhang XH, Yin XL, Ning HM, Liu B, Wang F et al (2016) c-Myc suppresses miR-451 dash, verticalYWTAZ/AKT axis via recruiting HDAC3 in acute myeloid leukemia. Oncotarget 7:77430–77443. https://doi.org/10.18632/oncotarget.12679
Sun Y, Huang YH, Huang FY, Mei WL, Liu Q, Wang CC, Lin YY, Huang C, Li YN, Dai HF et al (2018) 3’-epi-12beta-hydroxyfroside, a new cardenolide, induces cytoprotective autophagy via blocking the Hsp90/Akt/mTOR axis in lung cancer cells. Theranostics 8:2044–2060. https://doi.org/10.7150/thno.23304
Wang W, Zhang L, Wang Y, Ding Y, Chen T, Wang Y, Wang H, Li Y, Duan K, Chen S et al (2017) Involvement of miR-451 in resistance to paclitaxel by regulating YWHAZ in breast cancer. Cell Death Dis 8:e3071. https://doi.org/10.1038/cddis.2017.460
Adams BD, Kasinski AL, Slack FJ (2014) Aberrant regulation and function of microRNAs in cancer. Curr Biol 24:R762–R776. https://doi.org/10.1016/j.cub.2014.06.043
Albanell J, Dalmases A, Rovira A, Rojo F (2007) mTOR signalling in human cancer. Clin Transl Oncol 9:484–493. https://doi.org/10.1007/s12094-007-0092-6
Baltimore D, Boldin MP, O’Connell RM, Rao DS, Taganov KD (2008) MicroRNAs: new regulators of immune cell development and function. Nat Immunol 9:839–845. https://doi.org/10.1038/ni.f.209
Blandin Knight S, Crosbie PA, Balata H, Chudziak J, Hussell T, Dive C (2017) Progress and prospects of early detection in lung cancer. Open Biol 7:170070. https://doi.org/10.1098/rsob.170070
Cheng D, Xu Y, Sun C, He Z (2016) MicroRNA-451 sensitizes lung cancer cells to cisplatin through regulation of Mcl-1. Mol Cell Biochem 423:85–91. https://doi.org/10.1007/s11010-016-2827-6
Dong Y, Wang G (2020) Knockdown of lncRNA SNHG12 suppresses cell proliferation, migration and invasion in breast cancer by sponging miR-451a. Int J Clin Exp Pathol 13:393–402
Feng RM, Zong YN, Cao SM, Xu RH (2019) Current cancer situation in China: good or bad news from the 2018 Global Cancer Statistics? Cancer Commun (Lond) 39:22. https://doi.org/10.1186/s40880-019-0368-6
Filkowski JN, Ilnytskyy Y, Tamminga J, Koturbash I, Golubov A, Bagnyukova T, Pogribny IP, Kovalchuk O (2010) Hypomethylation and genome instability in the germline of exposed parents and their progeny is associated with altered miRNA expression. Carcinogenesis 31:1110–1115. https://doi.org/10.1093/carcin/bgp300
Geiman TM, Muegge K (2000) Lsh, an SNF2/helicase family member, is required for proliferation of mature T lymphocytes. Proc Natl Acad Sci USA 97:4772–4777. https://doi.org/10.1073/pnas.97.9.4772
Ilnytskyy Y, Zemp FJ, Koturbash I, Kovalchuk O (2008) Altered microRNA expression patterns in irradiated hematopoietic tissues suggest a sex-specific protective mechanism. Biochem Biophys Res Commun 377:41–45. https://doi.org/10.1016/j.bbrc.2008.09.080
Karachaliou N, Pilotto S, Lazzari C, Bria E, de Marinis F, Rosell R (2016) Cellular and molecular biology of small cell lung cancer: an overview. Transl Lung Cancer Res 5:2–15. https://doi.org/10.3978/j.issn.2218-6751.2016.01.02
Liu X, Hou X, Zhou Y, Li Q, Kong F, Yan S, Lei S, Xiong L, He J (2019) Downregulation of the helicase lymphoid-specific (HELLS) gene impairs cell proliferation and induces cell cycle arrest in colorectal cancer cells. Onco Targets Ther 12:10153–10163. https://doi.org/10.2147/OTT.S223668
Li L, Peng M, Xue W, Fan Z, Wang T, Lian J, Zhai Y, Lian W, Qin D, Zhao J (2018) Integrated analysis of dysregulated long non-coding RNAs/microRNAs/mRNAs in metastasis of lung adenocarcinoma. J Transl Med 16:372. https://doi.org/10.1186/s12967-018-1732-z
Manning BD, Cantley LC (2007) AKT/PKB signaling: navigating downstream. Cell 129:1261–1274. https://doi.org/10.1016/j.cell.2007.06.009
Mundi PS, Sachdev J, McCourt C, Kalinsky K (2016) AKT in cancer: new molecular insights and advances in drug development. Br J Clin Pharmacol 82:943–956. https://doi.org/10.1111/bcp.13021
Petrek H, Yu AM (2019) MicroRNAs in non-small cell lung cancer: gene regulation, impact on cancer cellular processes, and therapeutic potential. Pharmacol Res Perspect 7:e00528. https://doi.org/10.1002/prp2.528
Pietanza MC, Byers LA, Minna JD, Rudin CM (2015) Small cell lung cancer: will recent progress lead to improved outcomes? Clin Cancer Res 21:2244–2255. https://doi.org/10.1158/1078-0432.CCR-14-2958
Saxton RA, Sabatini DM (2017) mTOR signaling in growth, metabolism, and disease. Cell 168:960–976. https://doi.org/10.1016/j.cell.2017.02.004
Shen YY, Cui JY, Yuan J, Wang X (2018) MiR-451a suppressed cell migration and invasion in non-small cell lung cancer through targeting ATF2. Eur Rev Med Pharmacol Sci 22:5554–5561. https://doi.org/10.26355/eurrev_201809_15818
Streleckiene G, Inciuraite R, Juzenas S, Salteniene V, Steponaitiene R, Gyvyte U, Kiudelis G, Leja M, Ruzgys P, Satkauskas S et al (2020) miR-20b and miR-451a are involved in gastric carcinogenesis through the PI3K/AKT/mTOR signaling pathway: data from gastric cancer patients, cell lines and Ins-Gas mouse model. Int J Mol Sci 21:877. https://doi.org/10.3390/ijms21030877
Sun S, Hang T, Zhang B, Zhu L, Wu Y, Lv X, Huang Q, Yao H (2019) miRNA-708 functions as a tumor suppressor in colorectal cancer by targeting ZEB1 through Akt/mTOR signaling pathway. Am J Transl Res 11:5338–5356
Su Z, Zhao J, Rong Z, Geng W, Wang Z (2015) MiR-451, a potential prognostic biomarker and tumor suppressor for gastric cancer. Int J Clin Exp Pathol 8:9154–9160
Tian T, Li X, Zhang J (2019) mTOR signaling in cancer and mTOR inhibitors in solid tumor targeting therapy. Int J Mol Sci 20:755. https://doi.org/10.3390/ijms20030755
Uchida A, Seki N, Mizuno K, Yamada Y, Misono S, Sanada H, Kikkawa N, Kumamoto T, Suetsugu T, Inoue H (2019) Regulation of KIF2A by antitumor miR-451a inhibits cancer cell aggressiveness features in lung squamous cell carcinoma. Cancers (Basel) 11:258. https://doi.org/10.3390/cancers11020258
Wang R, Wang ZX, Yang JS, Pan X, De W, Chen LB (2011) MicroRNA-451 functions as a tumor suppressor in human non-small cell lung cancer by targeting ras-related protein 14 (RAB14). Oncogene 30:2644–2658. https://doi.org/10.1038/onc.2010.642
Xiao D, Huang J, Pan Y, Li H, Fu C, Mao C, Cheng Y, Shi Y, Chen L, Jiang Y et al (2017) Chromatin remodeling factor LSH is upregulated by the LRP6-GSK3beta-E2F1 Axis linking reversely with survival in Gliomas. Theranostics 7:132–143. https://doi.org/10.7150/thno.17032
Yamada Y, Arai T, Sugawara S, Okato A, Kato M, Kojima S, Yamazaki K, Naya Y, Ichikawa T, Seki N (2018) Impact of novel oncogenic pathways regulated by antitumor miR-451a in renal cell carcinoma. Cancer Sci 109:1239–1253. https://doi.org/10.1111/cas.13526
Yano M, Ouchida M, Shigematsu H, Tanaka N, Ichimura K, Kobayashi K, Inaki Y, Toyooka S, Tsukuda K, Shimizu N et al (2004) Tumor-specific exon creation of the HELLS/SMARCA6 gene in non-small cell lung cancer. Int J Cancer 112:8–13. https://doi.org/10.1002/ijc.20407
Yin P, Peng R, Peng H, Yao L, Sun Y, Wen L, Wu T, Zhou J, Zhang Z (2015) MiR-451 suppresses cell proliferation and metastasis in A549 lung cancer cells. Mol Biotechnol 57:1–11. https://doi.org/10.1007/s12033-014-9796-3
Zeng T, Peng L, Chao C, Fu B, Wang G, Wang Y, Zhu X (2014) miR-451 inhibits invasion and proliferation of bladder cancer by regulating EMT. Int J Clin Exp Pathol 7:7653–7662
Zhang G, Dong Z, Prager BC, Kim LJ, Wu Q, Gimple RC, Wang X, Bao S, Hamerlik P, Rich JN (2019) Chromatin remodeler HELLS maintains glioma stem cells through E2F3 and MYC. JCI Insight 4:e126140. https://doi.org/10.1172/jci.insight.126140
Zhao K, Wang L, Li T, Zhu M, Zhang C, Chen L, Zhao P, Zhou H, Yu S, Yang X (2017) The role of miR-451 in the switching between proliferation and migration in malignant glioma cells: AMPK signaling, mTOR modulation and Rac1 activation required. Int J Oncol 50:1989–1999. https://doi.org/10.3892/ijo.2017.3973
Zhu W, Li LL, Songyang Y, Shi Z, Li D (2020) Identification and validation of HELLS (Helicase, Lymphoid-Specific) and ICAM1 (Intercellular adhesion molecule 1) as potential diagnostic biomarkers of lung cancer. PeerJ 8:e8731. https://doi.org/10.7717/peerj.8731
Zocchi L, Mehta A, Wu SC, Wu J, Gu Y, Wang J, Suh S, Spitale RC, Benavente CA (2020a) Chromatin remodeling protein HELLS is critical for retinoblastoma tumor initiation and progression. Oncogenesis 9:25. https://doi.org/10.1038/s41389-020-0210-7
Zocchi L, Wu SC, Benavente CA (2020b) Heavenly HELLS? A potential new therapeutic target for retinoblastoma. Oncoscience 7:23–25. https://doi.org/10.18632/oncoscience.502
Author information
Authors and Affiliations
Contributions
Conceptualization: Jiyun Cui and Dianjie Lin; Methodology: Jiyun Cui and Jing Wang; Formal analysis and investigation: Yuyao Shen; Writing - original draft preparation: Jiyun Cui, Jing Wang and Yuyao Shen; Writing - review and editing: Dianjie Lin; Approval of the manuscript: All authors.
Corresponding author
Ethics declarations
Conflict of Interest
Jiyun Cui, Jing Wang, Yuyao Shen and Dianjie Lin declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cui, J., Wang, J., Shen, Y. et al. Suppression of HELLS by miR-451a represses mTOR pathway to hinder aggressiveness of SCLC. Genes Genom 43, 105–114 (2021). https://doi.org/10.1007/s13258-020-01028-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-01028-1