Abstract
Anti-severe acute respiratory syndrome coronavirus 2 (anti-SARS-CoV-2) nucleoprotein (NP) antibodies were isolated from pig sera using human SARS-CoV-2 NP-immobilized magnetic beads. The binding properties of the isolated antibodies against SARS-CoV-2 NP were tested via flow cytometry using SARS-CoV-2 NP-immobilized magnetic beads. A competitive immunoassay was developed for detecting SARS-CoV-2 NP as well as SARS-CoV-2 in the culture fluid using magnetic beads with immobilized anti-SARS-CoV-2 NP antibodies. Selectivity tests were carried out during the competitive immunoassay for SARS-CoV, MERS-CoV, and CoV strain 229E in the culture fluid.
Similar content being viewed by others
1 Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a beta-corona virus, and four protein types (spike proteins, membrane proteins, envelop proteins, and nucleoproteins (NPs)) have been used for detecting this virus. Among these, NPs have been most frequently used for developing vaccines and the SARS-CoV-2 antigen test for in vitro diagnosis of coronavirus disease 2019 (COVID-19) [1, 2]. Recently, the antibodies against human SARS-CoV-2 nucleoprotein (NP) from pig sera were used for detecting human SARS-CoV-2 in the culture fluid. This detection of human SARS-CoV-2 was possible because (1) a certain portion of virus particles was broken to release NP into the culture fluid [3] and (2) the NPs of humans and pigs presented highly homologous amino acid sequences (> 40%) during BLAST analysis. In a previous report, anti-SARS-CoV-2 NP antibodies were used to detect SARS-CoV-2 and other coronaviruses such as MERS-CoV and CoV strain 229E using the surface plasmon resonance biosensor [4].
Sandwich-type immunoassays have been used to detect antigens in samples using immobilized antibodies. In general, the response from the binding of antigens to the immobilized antibodies was generated in proportion to the antigen concentration in the samples [5,6,7]. In the case of competitive immunoassays, antigens in the sample compete for their binding sites with a fixed concentration of antigen-like signaling molecules. The signal from the indirect immunoassays usually decreased as the antigen concentration in the samples increased because the antigen-like signaling molecules were released from the immobilized antibodies. Such indirect immunoassays have been used to evaluate whether a certain target molecule was included in the sample, such as narcotics [8, 9] and insecticides [10,11,12].
In the present study, we presented a competitive immunoassay to detect SARS-CoV-2 using isolated anti-SARS-CoV-2 NP antibodies from pig sera, which exhibited a binding affinity to SARS-CoV-2 NP via immunoassays with immobilized antibodies and flow cytometry with magnetic beads of immobilized SARS-CoV-2 NP. The competitive immunoassay was configured by mixing magnetic beads with immobilized SARS-CoV-2 NPs and known concentrations of the isolated antibodies. The competitive assay overcame the Hook effect at high concentrations of SARS-CoV-2 and the narrow dynamic detection range of the standard rapid test for the SARS-CoV-2 Ag test. Eventually, a selectivity test was carried out for the competitive immunoassay for SARS-CoV, MERS-CoV, and CoV strain 229E in the culture fluid.
2 Results and Discussion
2.1 Isolation of Anti-SARS-CoV-2 NP Antibody from Pig Sera
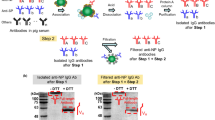
Anti-SARS-CoV-2 NP antibodies were isolated from pig sera using human SARS-CoV-2 NP-immobilized magnetic beads. As previously reported, SARS-CoV-2 NPs were immobilized on the magnetic beads via covalent bonding between the amino groups of SARS-CoV-2 NP and tosyl group present on the magnetic beads. As illustrated in Fig. 1(a), the isolation was carried out through the following steps: (1) incubation of pig sera for the binding of anti-SARS-CoV-2 NP antibodies to the magnetic beads, (2) acid dissociation of the bound proteins including anti-SARS-CoV-2 NP antibodies on the magnetic beads, and (3) protein-A purification of antibody (IgG) fraction among the dissociated proteins.
The isolated fraction was analyzed using SDS-PAGE, as illustrated in Fig. 1(b). To assess the antibody fraction after protein-A purification, the protein band corresponding to IgG (150 kDa) was observed without the treatment with the reducing agent dithiothreitol (DTT). After treatment with DTT for the reduction of disulfide bonds in IgG, the protein bands of the light chain (~ 25 kDa) and heavy chain (~ 50 kDa) of IgG were observed. These results indicated that the isolated proteins from pig sera corresponded to the IgG fraction. The isolation yield was estimated to be 2.91% ± 0.7% because 156 μg protein was obtained from 1 mL of pig sera before protein-A purification (n = 5). Eventually, the yield of antibody isolation was estimated to be 0.24% ± 0.05% because 12.3 μg of antibodies (IgG) was isolated from 1 mL of pig sera after protein-A purification (n = 5).
2.2 Binding Properties of Anti-SARS-CoV-2 NP Antibody
As previously reported, the binding properties of anti-SARS-CoV-2 NP antibodies were tested using immobilized SARS-CoV-2 NP proteins on a microplate and on magnetic beads. Initially, SARS-CoV-2 NP protein was immobilized on the microplate through hydrophilic and hydrophobic interactions, and then the isolated anti-SARS-CoV-2 NP antibodies were added to the microplate. As illustrated in Fig. 2(a), the amount of bound antibodies to the microplate was estimated using anti-pig IgG antibodies labeled with HRP and a chromogenic reaction with 3,3′,5,5′-tetramethylbenzidine (TMB). Eventually, the OD value was measured at 450 nm after quenching with sulfuric acid. As a negative control, a bovine serum albumin (BSA)-blocked microplate was used. As illustrated in Fig. 2(b), the OD value was increased in the antibody concentration range of 0.14–10.0 μg/mL (n = 3). In the negative control group, the OD value remained at the baseline level (n = 3). These results indicated that the isolated anti-SARS-CoV-2 NP antibodies from pig sera could bind specifically to the immobilized SARS-CoV-2 NP proteins.
The quantitative binding of anti-SARS-CoV-2 NP antibodies was analyzed via flow cytometry with SARS-CoV-2 NP-immobilized magnetic beads. As depicted in Fig. 3(a), the assay was carried out by mixing SARS-CoV-2 NP immobilized magnetic beads and samples at different concentrations of anti-SARS-CoV-2 NP antibodies. For the quantitative analysis of the bound anti-SARS-CoV-2 NP antibodies, the secondary antibody was reacted against pig-IgG labeled with fluorescein (λex = 488 nm, λem = 508 nm). Using the flow cytometry of magnetic beads, the fluorescence intensity from these beads was measured for different concentrations of isolated antibodies, as illustrated in Fig. 3(b). From the dose–response curve of flow cytometry (Fig. 3(c)), the anti-SARS-CoV-2 NP antibodies could be quantified at a concentration range of 0.03–22.2 μg/mL (n = 3). Considering the four parameter logistic equation [13], the binding constant (KD) of the isolated antibody against the immobilized SARS-CoV-2 NP on the magnetic beads was estimated to be 1.15 nM (n = 3).
Quantitative binding of anti-SARS-CoV-2 NP antibodies using flow cytometry with the isolated anti-SARS-CoV-2 NP on magnetic beads. a Configuration of the quantitative binding test. b Results of flow cytometry with anti-SARS-CoV-2 NP antibodies bound to SARS-CoV-2 NP on the magnetic beads. c Standard curve for quantitative binding of anti-SARS-CoV-2 NP antibodies using flow cytometry
2.3 Competitive Immunoassay of SARS-CoV-2
As previously reported, the isolated anti-SARS-CoV-2 NP antibodies from pig sera could be used to detect SARS-CoV-2 NP as well as SARS-CoV-2 in the culture fluid. In the present study, an assay was configured based on the competitive immunoassay for the quantitative analysis of SARS-CoV-2 NP as well as SARS-CoV-2 using magnetic beads with immobilized SARS-CoV-2 NP. As illustrated in Fig. 4(a), the magnetic beads and a known concentration of anti-SARS-CoV-2 NP antibodies were mixed with the samples. When the sample had SARS-CoV-2 NP (or SARS-CoV-2), the isolated antibodies were bound to both magnetic beads and SARS-CoV-2 NP (or SARS-CoV-2). Eventually, secondary antibodies against pig-IgG labeled with fluorescein were used to quantify the bound antibodies on the magnetic beads. In the negative samples lacking SARS-CoV-2 NP (or SARS-CoV-2), all the isolated antibodies were bound to the magnetic beads. Therefore, when the secondary antibodies against pig-IgG labeled with fluorescein were treated, a remarkably higher fluorescence signal was observed from the magnetic beads. Therefore, the fluorescence signal decreased with an increase in the SARS-CoV-2 NP (or SARS-CoV-2) in the sample, and the positiveness of the sample could be evaluated by considering the difference between signals of the positive and negative samples.
Competitive immunoassay using the isolated anti-SARS-CoV-2 NP antibodies on magnetic beads. a Configuration of competitive immunoassay. b Results of flow cytometry for quantitative analysis of SARS-CoV-2 NP using competitive immunoassay. c Standard curve for quantitative analysis of SARS-CoV-2 NP using competitive immunoassay
First, SARS-CoV-2 NP was quantified in samples using a competitive assay. As illustrated in Fig. 4(b), PBS without SARS-CoV-2 NP was used as a negative control. When samples with SARS-CoV-2 NP were applied to the competitive assay, the fluorescence signal gradually decreased in the concentration range of 0.12–10.0 μg/mL and the signal was maintained at a baseline level above the concentration of 1 µg/mL (Fig. 4(c)). These results indicate that the competitive assay could be used for the quantitative analysis of SARS-CoV-2 NP.
Second, SARS-CoV-2 in the culture fluid was evaluated via a competitive assay. The quantification of SARS-CoV-2 was carried out using a rapid test for SARS-CoV-2 from SD Biosensors (Seoul, Korea) [14]. As illustrated in Fig. 5(a), the signal band was observed for the samples with diluted SARS-CoV-2 culture fluid, and the control bands indicated the validity of the test. According to the manufacturer’s manual [14], the cutoff value for determining the positivity of COVID-19 was established to be the dilution factor of 925, corresponding to a culture fluid fraction of 0.11%. As illustrated in Fig. 5(b), PBS without diluted SARS-CoV-2 culture fluid was used as a negative control for both the rapid test and the competitive assay. In the case of the rapid test, a linear detection range was observed in the dilution fraction range of 0.15%–1%. For the higher dilution fraction, the signal was observed to decrease, which indicated the Hook effect of the immunoassay. For the lower dilution fraction, the signal remained at the baseline level. When the diluted samples of SARS-CoV-2 culture fluid were applied to the competitive assay, the fluorescence signal gradually decreased the dilution fraction of 0.078%–5%, and the quantitative analysis was determined to be possible in the whole range of used dilution fractions (n = 6). These results indicated that (1) the competitive assay had a considerably wider detection range than that observed in the conventional rapid test and (2) the competitive assay could be used even for the dilution fraction range where the Hook effect occurred (in the range of high dilution fraction) and the detection limit was attained (in the range of low dilution fraction).
Competitive immunoassay of SARS-CoV-2. a Results of a rapid test for the diagnosis of SARS-CoV-2 antigen test using diluted culture fluid with SARS-CoV-2. b Comparison of the standard curve for quantitative analysis of SARS-CoV-2 using rapid test and competitive immunoassay. c Selectivity test for competitive immunoassay of SARS-CoV, MERS-CoV, and CoV strain 229E
Third, the selectivity of the competitive assay was tested against SARS-CoV-2 and other types of coronaviruses of SARS-CoV, MERS-CoV, and CoV strain 229E. The culture fluid of CoV strain 229E was used as a negative control. To compare the assay signals, normalized fluorescence intensity was used in the dilution fraction range of 0.078%–5%. As shown in Fig. 5(c), the coronaviruses SARS-CoV-2, SARS-CoV, and MERS-CoV revealed quantitative response according to the dilution fraction of each culture fluid. Such response indicated that the isolated anti-SARS-CoV-2 NP antibodies had a binding affinity against NP proteins from the corresponding viruses. In particular, the signals of the competitive assay from SARS-CoV-2 and SARS-CoV were quite similar because the homology of amino acid sequences in both NPs was estimated to be more than 90% (Table 1) from BLAST analysis, whereas the homology in MERS-CoV and CoV strain 229E was 48% (Table 2) and 29% (Table 3), respectively. These results indicated that a competitive assay based on magnetic beads with immobilized NPs could be used for the quantitative analysis of SARS-CoV-2 and other types of coronaviruses of SARS-CoV, MERS-CoV, and CoV strain 229E.
3 Conclusion
Anti-SARS-CoV-2 NP antibodies were isolated from pig sera using SARS-CoV-2 NP-immobilized magnetic beads, via the three steps discussed in the Results and Discussion section. Finally, the yield of isolated antibody was estimated to be 0.24% ± 0.05% because 12.3 μg of antibodies (IgG) was isolated from 1 mL of pig sera after protein-A purification (n = 5). From the dose–response curve of flow cytometry, the quantification of anti-SARS-CoV-2 NP antibodies was possible at 0.03–22.2 μg/mL (n = 3), and the binding constant (KD) of the isolated antibody against SARS-CoV-2 NP on the magnetic beads was estimated to be 1.15 nM (n = 3). A configuration based on the competitive immunoassay was developed for the quantitative analysis of SARS-CoV-2 NP as well as SARS-CoV-2 using magnetic beads with immobilized SARS-CoV-2 NP. When diluted samples of SARS-CoV-2 culture fluid were applied to the competitive assay, the fluorescence signal gradually decreased the dilution fraction of 0.078%–5%, and the quantitative analysis was carried out for the entire range of used dilution fractions. These results indicated that (1) the competitive assay had a far wider detection range than the conventional rapid test and (2) the competitive assay could be used even for the dilution fraction range where the Hook effect occurred (in the range of high dilution fraction) and the detection limit was attained (in the range of low dilution fraction). Using the selectivity test for SARS-CoV-2 and other kinds of coronaviruses such as SARS-CoV, MERS-CoV and CoV strain 229E, a competitive assay based on the magnetic beads with immobilized SARS-CoV-2 NP was used to quantify these viruses.
4 Materials and Methods
4.1 Materials
Anti-pig IgG labeled with horseradish peroxidase (ab102135) or fluorescein (ab9611) was purchased from Abcam (Cambridge, UK). The tosyl-activated magnetic beads (diameter of 2.8 µm, 14203) were purchased from Invitrogen Co. (Carlsbad, CA, USA). BSA (A2153) and other chemicals were purchased from Sigma-Aldrich Korea (Seoul, Korea). The bicinchoninic acid (BCA) assay kit (23227), pig sera (porcine sera, 26250084), and TMB reagent with H2O2 solution (34021) were purchased from Thermo Fisher Scientific (Waltham, MA, USA). Maxibinding microplates (32296) were purchased from SPL Life Science (Pocheon, Korea). Culture fluids of SARS-CoV-2 (0810587CFHI), MERS-CoV (0810575CFHI), CoV strain 229E (0810229CFHI), and SARS-CoV (NATSARS-ST) were purchased from Zeptometrix (Buffalo, NY, USA). The His-tagged SARS-CoV-2 nucleoprotein (SARS-CoV-2 NP) was supplied by Genbody Inc. (Cheonan, Korea) and Optolane Inc. (Seongnam, Korea).
4.2 Purification of Anti-SARS-CoV-2 Antibodies
The antibodies were isolated from pig sera using the following procedure. Initially, the pig sera (1 mL) were added to magnetic beads coated with SARS-CoV-2 NP (10 mg). Thereafter, the reaction tube was mixed continuously with a wheel-rotating mixer (RT-10, Daehan Scientific Co, Wonju, Korea) for 1 h. The beads were prepared according to the manufacturer’s instructions. Next, the proteins bound to the beads were dissociated by treatment with 1 mL of 0.1 M glycine–HCl buffer (pH 2.7) for 10 s. Furthermore, Tris–HCl buffer (1 M, pH 10.5, 50 μL) was added for neutralization. The SARS-CoV-2 NP antibodies were eventually purified using a protein-A column without unwanted proteins. The product was finally concentrated with Amicon centrifugal filtration (molecular weight cutoff: 3 kDa). The yield of antibodies was determined as the amount of protein isolated from a unit volume (1 mL) of pig sera. The concentration of protein was estimated using a BCA assay kit from Thermo Fisher Scientific (Waltham, MA, USA).
4.3 The Binding Affinity Analysis of Anti-SARS-CoV-2 Antibodies
Immunoassays using microplates (Fig. 2) and flow cytometry (Fig. 3) were performed to analyze the binding affinity of the isolated antibodies. For the microplate assay, the proteins (SARS-CoV-2 NP) were immobilized on a Maxibinding plate for 2 h at 100 μg/mL. Thereafter, each well was blocked with BSA (5 mg/mL, 1 h). The SARS-CoV-2 antibodies were added to each well at various concentrations, followed by a secondary antibody reaction (100 ng/mL, 1 h for both antibodies). After washing once with PBST and twice with PBS, the colorimetric reaction was performed using the TMB reagent with H2O2 (1:1 retio, 100 μL) for 3 min. The optical density was measured at a wavelength of 450 nm after quenching with 2 M sulfuric acid (100 μL) using a microplate reader (Versamax Microplate reader) from Molecular Devices Co. (San Jose, CA, USA).
For the assay based on flow cytometry, 0.1 mg of the magnetic beads immobilized with SARS-CoV-2 NP was utilized. Anti-SARS-CoV-2 NP antibodies were incubated for 1 h at various concentrations, followed by the reaction of anti-pig(FITC) antibody solution (5 μg/mL, 1 h). After the aforementioned washing step, the fluorescence signals were obtained using FACSCalibur (BD, NJ, USA).
4.4 Competitive Assay of Anti-SARS-CoV-2 Antibodies
The magnetic beads (0.1 mg) were aliquoted into each vial. Then various concentrations of SARS-CoV-NP (100 μL) and fixed concentration of anti-SARS-CoV-2 antibody (12.5 μg/mL, 100 μL) were added simultaneously. After 1 h of washing, anti-pig(FITC) antibody solution (12.5 μg/mL, 100 μL) was incubated for 1 h. The fluorescence signals were obtained using FACSCalibur (BD, NJ, USA).
4.5 Standard Curve Fitting
The results were fitted with the four parameter logistic equation [15, 16] as described below:
where a and d are the maximum and minimum signals (a.u.), respectively; c is the concentration of SARS-CoV-2 NP (pM), and b is the Hill’s slope of the curve.
References
Spencer, K.-A., Osorio, F.A., Hiscox, J.A.: Recombinant viral proteins for use in diagnostic ELISAs to detect virus infection. Vaccine. 25, 5653–5659 (2007). https://doi.org/10.1016/j.vaccine.2007.02.053
Feng, W., Newbigging, A.M., Le, C., Pang, B., Peng, H., Cao, Y., Wu, J., Abbas, G., Song, J., Wang, D.-B., Cui, M., Tao, J., Tyrrell, D.L., Zhang, X.-E., Zhang, H., Le, X.C.: Molecular diagnosis of COVID-19: challenges and research needs. Anal. Chem. 92, 10196–10209 (2020). https://doi.org/10.1021/acs.analchem.0c02060
Li, Y.-H., Li, J., Liu, X.-E., Wang, L., Li, T., Zhou, Y.-H., Zhuang, H.: Detection of the nucleocapsid protein of severe acute respiratory syndrome coronavirus in serum: comparison with results of other viral markers. J. Virol. Methods. 130, 45–50 (2005). https://doi.org/10.1016/j.jviromet.2005.06.001
Bong, J.-H., Kim, T.-H., Jung, J.-Y., Lee, S.-J., Sung, J.S., Lee, C., Kang, M.-J., Kim, H.O., Pyun, J.-C.: Pig sera-derived anti-SARS-CoV-2 antibodies in surface plasmon resonance biosensors. Biochip. J. 14, 358–368 (2020). https://doi.org/10.1007/s13206-020-4404-z
Bulemo, P.M., Kim, I.D.: Recent advances in ABO(3) perovskites: their gas-sensing performance as resistive-type gas sensors. J. Korean Ceram. Soc. 57, 24–39 (2020). https://doi.org/10.1007/s43207-019-00003-1
Kim, H.R., Bong, J.H., Jung, J., Sung, J.S., Kang, M.J., Park, J.G., Pyun, J.C.: An on-chip chemiluminescent immunoassay for bacterial detection using in situ-synthesized cadmium sulfide nanowires with passivation layers. Biochip. J. 14, 11 (2020). https://doi.org/10.1007/s13206-020-4305-1
Pham, X.H., Hahm, E., Huynh, K.H., Son, B.S., Kim, H.M., Jun, B.H.: Sensitive colorimetric detection of prostate specific antigen using a peroxidase-mimicking anti-PSA antibody coated Au Nanoparticle. Biochip. J. (2020). https://doi.org/10.1007/s13206-019-4204-5
Tran, R.J., Sly, K.L., Conboy, J.C.: Revealing the kinetic advantage of a competitive small-molecule immunoassay by direct detection. Anal. Chem. (2020). https://doi.org/10.1021/acs.analchem.0c02286
Singh, S., Mishra, P., Banga, I., Parmar, S.A., Tripathi, P.P., Gandhi, S.: Chemiluminescence based immunoassay for the detection of heroin and its metabolites. Bioimpacts. 8, 53–58 (2018). https://doi.org/10.15171/bi.2018.07
Cheng, N., Shi, Q., Zhu, C., Li, S., Lin, Y., Du, D.: Pt–Ni(OH)2 nanosheets amplified two-way lateral flow immunoassays with smartphone readout for quantification of pesticides. Biosens. Bioelectron. 142, 111498 (2019). https://doi.org/10.1016/j.bios.2019.111498
He, J., Tao, X., Wang, K., Ding, G., Li, J., Li, Q.X., Gee, S.J., Hammock, B.D., Xu, T.: One-step immunoassay for the insecticide carbaryl using a chicken single-chain variable fragment (scFv) fused to alkaline phosphatase. Anal. Biochem. 572, 9–15 (2019). https://doi.org/10.1016/j.ab.2019.02.022
Wang, Y., Li, Z., Barnych, B., Huo, J., Wan, D., Vasylieva, N., Xu, J., Li, P., Liu, B., Zhang, C., Hammock, B.D.: Investigation of the small size of nanobodies for a sensitive fluorescence polarization immunoassay for small molecules: 3-Phenoxybenzoic Acid, an exposure biomarker of pyrethroid insecticides as a model. J. Agric. Food Chem. 67, 11536–11541 (2019). https://doi.org/10.1021/acs.jafc.9b04621
Tijssen, P.: Practice and theory of enzyme immunoassays. Elsevier, Amsterdam (1985)
Masdor, N.A., Altintas, Z., Tothill, I.E.: Sensitive detection of Campylobacter jejuni using nanoparticles enhanced QCM sensor. Biosens. Bioelectron. 78, 328–336 (2016). https://doi.org/10.1016/j.bios.2015.11.033
Hasani, A., Do, H.H., Tekalgne, M., Hong, S.H., Jang, H.W., Kim, S.Y.: Recent progress of two-dimensional materials and metal-organic framework-based taste sensors. J. Korean Ceram. Soc. 57, 353–367 (2020). https://doi.org/10.1007/s43207-020-00047-8
Acknowledgments
This work was supported by the National Research Foundation of Korea [grant number: NRF-2020R1A2B5B01002187 and NRF-2020R1A5A101913111.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bong, JH., Kim, TH., Jung, J. et al. Competitive Immunoassay of SARS-CoV-2 Using Pig Sera-Derived Anti-SARS-CoV-2 Antibodies. BioChip J 15, 100–108 (2021). https://doi.org/10.1007/s13206-021-00011-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-021-00011-6