Abstract
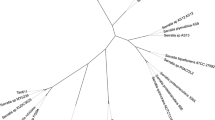
In this study, an indigenous novel hydrocarbonoclastic (kerosene and diesel degrading) and biosurfactant producing strain Fictibacillus phosphorivorans RP3 was identified. The characteristics of bacterial strain were ascertained through its unique morphological and biochemical attributes, 16S rRNA sequencing, and phylogenetic analysis. The degradation of hydrocarbons by F. phosphorivorans RP3 was observed at Day 7, Day 10 and Day 14 of the experimental duration. GC-FID chromatograms demonstrated a significant increase in hydrocarbon degradation (%) with progressing days (from 7 to 14). The bacterium exhibited capability to utilize and degrade n-hexadecane (used for primary screening) and petroleum hydrocarbons (kerosene and diesel; by ≥ 90%). With increase in the number of experimentation days, the optical density of the culture medium increased, whereas pH declined (became acidic) for both Kerosene and Diesel. Absence of resistance to routinely used antibiotics makes it an ideal candidate for future field application. The study is, thus, significant in view of toxicological implications of hydrocarbons and their degradation using environmentally safe techniques so as to maintain ecological and human health.





Similar content being viewed by others
References
Adebusoye SA, Ilori MO, Amund OO, Teniola OD, Olatope SO (2007) Microbial degradation of petroleum hydrocarbons in a polluted tropical stream. World J Microbiol Biotechnol 23:1149–1159. https://doi.org/10.1007/s11274-007-9345-3
Albert B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Molecular biology of the cell. Garland Science, New York
Al-Dhabaan FA (2019) Morphological, biochemical and molecular identification of petroleum hydrocarbons biodegradation bacteria isolated from oil polluted soil in Dhahran, Saudi Arabia. Saudi J Biol Sci 26:1247–1252. https://doi.org/10.1016/j.sjbs.2018.05.029
Asghar HN, Rafique HM, Zahir ZA, Khan MY, Akhtar MJ, Naveed M, Saleem M (2016) Petroleum hydrocarbons-contaminated soils: remediation approaches. In: Hakeem K, Akhtar J, Sabir M (eds) Soil science: agricultural and environmental prospectives. Springer International Publishing, Switzerland, pp 105–129
Aydin DC, Icgen B (2018) Gas chromatographic analyses of kerosene bioremediation displayed distinctive pattern of n-alkane degradation. Pet Sci Technol 36:1905–1912. https://doi.org/10.1080/10916466.2018.1517170
Ayed HB, Jemil N, Maalej H, Bayoudh A, Hmidet N, Nasri M (2015) Enhancement of solubilization and biodegradation of diesel oil by biosurfactant from Bacillus amyloliquefaciens An6. Int Biodeterior Biodegrad 99:8–14. https://doi.org/10.1016/j.ibiod.2014.12.009
Balan SS, Kumar CG, Jayalakshmi S (2017) Aneurinifactin, a new lipopeptide biosurfactant produced by a marine Aneurinibacillus aneurinilyticus SBP-11 isolated from Gulf of Mannar: purification, characterization and its biological evaluation. Microbiol Res 194:1–9. https://doi.org/10.1016/j.micres.2016.10.005
Banerjee S, Misra A, Chaudhury S, Dam B (2019) A Bacillus strain TCL isolated from Jharia coalmine with remarkable stress responses, chromium reduction capability and bioremediation potential. J Hazard Mater 367:215–223. https://doi.org/10.1016/j.jhazmat.2018.12.038
Bauer AW, Sherris JC, Turck M, Kirby WMM (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 45:493–496. https://doi.org/10.1093/ajcp/45.4_ts.493
Behnajady MA, Modirshahla N, Hamzavi R (2006) Kinetic study on photocatalytic degradation of CI Acid Yellow 23 by ZnO photocatalyst. J Hazard Mater 133:226–232. https://doi.org/10.1016/j.jhazmat.2005.10.022
Bertrand RL (2019) Lag phase is a dynamic, organized, adaptive, and evolvable period that prepares bacteria for cell division. J Bacteriol 201:e00697-e718. https://doi.org/10.1128/JB.00697-18
Cases I, de Lorenzo V (2005) Genetically modified organisms for the environment: stories of success and failure and what we have learned from them. Int Microbiol 8:213–222. https://doi.org/10.13039/50110000780
Chen Y, Wang W, Zhou D, Jing T, Li K, Zhao Y, Tang W, Qi D, Zhang M, Zang X, Luo Y (2020) Biodegradation of lignocellulosic agricultural residues by a newly isolated Fictibacillus sp. YS-26 improving carbon metabolic properties and functional diversity of the rhizosphere microbial community. Bioresour Technol 310:123381. https://doi.org/10.1016/j.biortech.2020.123381
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview. Biotechnol Res Int 2011:13. https://doi.org/10.4061/2011/941810
Das M, Bhattacharya A, Banu S, Kotoky J (2017) Enhanced biodegradation of anthracene by Bacillus cereus strain JMG-01 isolated from hydrocarbon contaminated soils. Soil Sediment Contam 26:510–525. https://doi.org/10.1080/15320383.2017.1357111
Dhiman K, Shirkot P (2015) Bioprospecting and molecular characterization of laccase producing bacteria from paper mills of Himachal Pradesh. Proc Natl Acad Sci India Sect B Biol Sci 85:1095–1103. https://doi.org/10.1007/s40011-015-0541-x
Dixit H, Lowry M, Mohsin U, Moond M, Kumar S, Chokriwal A, Gupta S (2018) Growth optimization and comparative analysis of diesel oil degradation potential of Bacillus sp. isolated from petroleum contaminated soil of Barmer, Rajasthan. Int J Recent Sci Res 9:24730–24737. https://doi.org/10.24327/ijrsr.2018.0903.1717
Dvořák P, Nikel PI, Damborský J, de Lorenzo V (2017) Bioremediation 3.0: Engineering pollutant-removing bacteria in the times of systemic biology. Biotechnol Adv 35:845–866. https://doi.org/10.1016/j.biotechadv.2017.08.001
García-Cruz NU, Sánchez-Avila JI, Valdés-Lozano D, Gold-Bouchot G, Aguirre-Macedo L (2018) Biodegradation of hexadecane using sediments from rivers and lagoons of the Southern Gulf of Mexico. Mar Pollut Bull 128:202–207. https://doi.org/10.1016/j.marpolbul.2018.01.026
Glaeser SP, Dott W, Busse HJ, Kämpfer P (2013) Fictibacillus phosphorivorans gen. nov., sp. nov. and proposal to reclassify Bacillus arsenicus, Bacillus barbaricus, Bacillus macauensis, Bacillus nanhaiensis, Bacillus rigui, Bacillus solisalsi and Bacillus gelatini in the genus Fictibacillus. Int J Syst Evol Microbiol 63:2934–2944. https://doi.org/10.1099/ijs.0.049171-0
Guerra AB, Oliveira JS, Silva-Portela RC, Araujo W, Carlos AC, Vasconcelos ATR et al (2018) Metagenome enrichment approach used for selection of oil-degrading bacteria consortia for drill cutting residue bioremediation. Environ Pollut 235:869–880. https://doi.org/10.1016/j.envpol.2018.01.014
Inakollu S, Hung HC, Shreve GS (2004) Biosurfactant enhancement of microbial degradation of various structural classes of hydrocarbon in mixed waste systems. Environ Eng Sci 21:463–469. https://doi.org/10.1089/109287504135846
Jia X, He Y, Huang L, Jiang D, Lu W (2019) n-Hexadecane and pyrene biodegradation and metabolization by Rhodococcus sp. T1 isolated from oil contaminated soil. Chinese J Chem Eng 27:411–417. https://doi.org/10.1016/j.cjche.2018.03.034
Kaushik P, Malik A (2009) Microbial decolourization of textile dyes through isolates obtained from contaminated sites. J Sci Ind Res 68:325–331
Khan MAI, Biswas B, Smith E, Naidu R, Megharaj M (2018) Toxicity assessment of fresh and weathered petroleum hydrocarbons in contaminated soil-a review. Chemosphere 212:755–767. https://doi.org/10.1016/j.chemosphere.2018.08.094
Kleindienst S, Seidel M, Ziervogel K, Grim S, Loftis K, Harrison S, et al. (2015) Chemical dispersants can suppress the activity of natural oil-degrading microorganisms. Proc Natl Acad Sci USA 112:14900–14905. https://doi.org/10.1073/pnas.1507380112
Krasowska A, Sigler K (2014) How microorganisms use hydrophobicity and what does this mean for human needs? Front Cell Infect Microbiol 4:112. https://doi.org/10.3389/fcimb.2014.00112
Kumar S, Teotia US, Singh Y (2017) Screening of poly vinyl chloride degrading bacteria from plastic contaminated area of Baddi. J Appl Pharm Res 5:34–37. https://doi.org/10.18231/2348-0335.2017.0005
Leahy JG, Colwell RR (1990) Microbial degradation of hydrocarbons in the environment. Microbiol Mol Biol Rev 54:305–315
Liu T, Wang F, Guo LP, Li XL, Yang XJ, Lin AJ (2012) Biodegradation of n-hexadecane by bacterial strains B1 and B2 isolated from petroleum-contaminated soil. Sci China Chem 55:1968–1975. https://doi.org/10.1007/s11426-012-4618-6
Marchut-Mikolajczyk O, Drożdżyński P, Struszczyk-Świta K (2020) Biodegradation of slop oil by endophytic Bacillus cereus EN18 coupled with lipase from Rhizomucor miehei (Palatase®). Chemosphere 250:126203. https://doi.org/10.1016/j.chemosphere.2020.126203
Margesin R, Moertelmaier C, Mair J (2013) Low-temperature biodegradation of petroleum hydrocarbons (n-alkanes, phenol, anthracene, pyrene) by four actinobacterial strains. Int Biodeterior Biodegrad 84:185–191. https://doi.org/10.1016/j.ibiod.2012.05.004
Martin-Sanchez PM, Becker R, Gorbushina AA, Toepel J (2018) An improved test for the evaluation of hydrocarbon degradation capacities of diesel-contaminating microorganisms. Int Biodeterior Biodegrad 129:89–94. https://doi.org/10.1016/j.ibiod.2018.01.00
Mishra S, Singh SN (2012) Microbial degradation of n-hexadecane in mineral salt medium as mediated by degradative enzymes. Bioresour Technol 111:148–154. https://doi.org/10.1016/j.biortech.2012.02.049
Mojarad M, Alemzadeh A, Ghoreishi G, Javaheri M (2016) Kerosene biodegradation ability and characterization of bacteria isolated from oil-polluted soil and water. J Environ Chem Eng 4:4323–4329. https://doi.org/10.1016/j.jece.2016.09.035
Morales-Guzmán G, Ferrera-Cerrato R, Rivera-Cruz MC, Torres-Bustillos LG, Arteaga-Garibay RI, Mendoza-López MR, Esquivel-Cote R, Alarcón A (2017) Diesel degradation by emulsifying bacteria isolated from soils polluted with weathered petroleum hydrocarbons. Appl Soil Ecol 121:127–134. https://doi.org/10.1016/j.apsoil.2017.10.003
Neris JB, Olivares DMM, Velasco FG, Luzardo FHM, Correia LO, González LN (2019) HHRISK: a code for assessment of human health risk due to environmental chemical pollution. Ecotox Environ Safe 170:538–547. https://doi.org/10.1016/j.ecoenv.2018.12.017
Nie H, Nie M, Xiao T, Wang Y, Tian X (2017) Hexadecane degradation of Pseudomonas aeruginosa NY3 promoted by glutaric acid. Sci Total Environ 575:1423–1428. https://doi.org/10.1016/j.scitotenv.2016.09.223
Pemmaraju SC, Sharma D, Singh N, Panwar R, Cameotra SS, Pruthi V (2012) Production of microbial surfactants from oily sludge-contaminated soil by Bacillus subtilis DSVP23. Appl Biochem Biotechnol 167:1119–1131. https://doi.org/10.1007/s12010-012-9613-z
Płaza GA, Jangid K, Łukasik K, Nałęcz-Jawecki G, Berry CJ, Brigmon RL (2008) Reduction of petroleum hydrocarbons and toxicity in refinery wastewater by bioremediation. Bull Environ Contam Toxicol 81(4):329–333. https://doi.org/10.1007/s00128-008-9411-z
Poornachander RM, Anitha Y, Satyaprasad K (2016) Abilities of Bacillus cereus CPOU13 in biodegradation of polycyclic aromatic hydrocarbons (PAHs). J Pharm Chem Biol Sci 4:54–64
Rezaei M, Habibi-Yangjeh A (2013) Simple and large scale refluxing method for preparation of Ce-doped ZnO nanostructures as highly efficient photocatalyst. Appl Surf Sci 265:591–596. https://doi.org/10.1016/j.apsusc.2012.11.053
Ron EZ, Rosenberg E (2014) Enhanced bioremediation of oil spills in the sea. Curr Opin Biotechnol 27:191–194. https://doi.org/10.1016/j.copbio.2014.02.004
Sawhney R, Bhojia AK (2011) Congo Red (Azo dye) decolourization by local isolate VT-II inhabiting dye-effluent exposed soil. Int J Environ Sci 1:1261–1267
Shahi A, Aydin S, Ince B, Ince O (2016) Reconstruction of bacterial community structure and variation for enhanced petroleum hydrocarbons degradation through biostimulation of oil contaminated soil. Chem Eng J 306:60–66. https://doi.org/10.1016/j.cej.2016.07.016
Shen T, Pi Y, Bao M, Xu N, Liab Y, Lub J (2015) Biodegradation of different petroleum hydrocarbons by free and immobilized microbial consortia. Environ Sci 17:2022–2033. https://doi.org/10.1039/C5EM00318K
Shiri Z, Kermanshahi RK, Soudi MR, Farajzadeh D (2014) Isolation and characterization of an n-hexadecane degrading Acinetobacter baumannii KSS1060 from a petrochemical wastewater treatment plant. Int J Environ Sci Technol 12:455–464. https://doi.org/10.1007/s13762-014-0702-0
Siegmund I, Wagner F (1991) New method for detecting rhamnolipids excreted by Pseudomonas species during growth on mineral agar. Biotechnol Tech 5:265–268. https://doi.org/10.1007/BF02438660
Varjani SJ (2017) Microbial degradation of petroleum hydrocarbons. Bioresour Technol 223:277–286. https://doi.org/10.1016/j.biortech.2016.10.037
Whitman WB, Goodfellow M, Kämpfer P, Busse H-J, Trujillo ME, Ludwig W, Suzuki K-I (eds) (2012) Bergey’s Manual of Systematic Bacteriology, vol 5, 2nd edn. Parts A and B, Springer-Verlag, New York
Xu X, Liu W, Tian S, Wang W, Qi Q, Jiang P, Gao X, Li F, Li H, Yu H (2018) Petroleum hydrocarbon-degrading bacteria for the remediation of oil pollution under aerobic conditions: a perspective analysis. Front Microbiol 9:2885. https://doi.org/10.3389/fmicb.2018.02885
Xue J, Yu Y, Bai Y, Wang L, Wu Y (2015) Marine oil-degrading microorganisms and biodegradation process of petroleum hydrocarbon in marine environments: a review. Curr Microbiol 71:220–228. https://doi.org/10.1007/s00284-015-0825-7
Youssef NH, Duncan KE, Nagle DP, Savage KN, Knapp RM, Mcinerney MJ (2004) Comparison of methods to detect biosurfactant production by diverse microorganisms. J Microbiol Method 56:339–347. https://doi.org/10.1016/j.mimet.2003.11.001
Zhang X, Xu D, Zhu C, Lundaa T, Scherr KE (2012) Isolation and identification of biosurfactant producing and crude oil degrading Pseudomonas aeruginosa strains. Chem Eng J 209:138–146. https://doi.org/10.1016/j.cej.2012.07.110
Zhong H, Wang Z, Liu Z, Liu Y, Yu M, Zeng G (2016) Degradation of hexadecane by Pseudomonas aeruginosa with the mediation of surfactants: relation between hexadecane solubilization and bioavailability. Int Biodeterior Biodegrad 115:141–145. https://doi.org/10.1016/j.ibiod.2016.08.008
Acknowledgements
RP is thankful to University Grants Commission (UGC), India, for research fellowship.
Author information
Authors and Affiliations
Contributions
RP and BK formulated the hypothesis, RP carried out work of the study; RP and BK prepared the first draft of the manuscript; RP, PS and SR analysed and edited the manuscript; HPS, DRB and RKK supervised and reviewed the work; HPS, DRB and RKK proofread the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that they have no conflict of interest in the publication.
Rights and permissions
About this article
Cite this article
Pandey, R., Sharma, P., Rathee, S. et al. Isolation and characterization of a novel hydrocarbonoclastic and biosurfactant producing bacterial strain: Fictibacillus phosphorivorans RP3. 3 Biotech 11, 105 (2021). https://doi.org/10.1007/s13205-021-02655-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-021-02655-5