Abstract
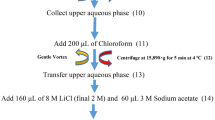
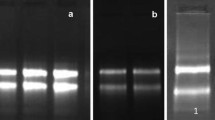
Isolation of high-quality RNA from coffee is challenging because of high level of polysaccharides, polyphenols and other secondary metabolites. In the present study, a rapid and efficient RNA extraction protocol from different tissues of coffee was optimized. Sufficiently high quality and quantity (225.6–454.8 µg/g) of RNA was obtained by using the optimized protocol. The presence of two distinct bands of 28S rRNA and 18S rRNA in agarose gel proved the intactness of the RNA samples. The average spectrophotometric values of the isolated RNA ranged from 1.96 to 2.02 (A260/280) and 1.95 to 2.14 (A260/230), indicating the high quality of RNA devoid of polyphenols, polysaccharides and protein contamination. In the optimized protocol, addition of PVPP to the extraction buffer and a brief incubation of samples at 65 °C and subsequent purification with potassium acetate resulted in good-quality RNA isolation. The suitability of RNA for downstream processing was confirmed by PCR amplification with cytochrome c oxidase gene-specific primers. The amplification of a single 392 bp fragment using cDNA and 1.5 kb fragment using genomic DNA samples confirmed the absence of DNA contamination. The present protocol is rapid and yielded good quality and quantity of RNA suitable for functional genomics studies.


Similar content being viewed by others
References
Ainsworth C (1994) Isolation of RNA from floral tissue of Rumex acetosa (Sorrel). Plant Mol Biol Rep 12(3):198–203
Asif M, Trivedi P, Solomos T, Tucker M (2006) Isolation of high-quality RNA from apple (Malus domestica) fruit. J Agric Food Chem 54:5227–5229
Choudhary SB, Kumar M, Chowdhury I, Singh RK, Pandey SP, Sharma HK, Karmakar PG (2016) An efficient and cost effective method of RNA extraction from mucilage, phenol and secondary metabolite rich bark tissue of tossa jute (C. olitorius) actively developing phloem fiber. 3 Biotech 6(1):100. https://doi.org/10.1007/s13205-016-0415-9
de Paula MFB, Sagio SA, Lazzari F, Barreto HG, Paiva LV, Antonio C Jr (2012) Efficiency of RNA extraction protocols in different types of coffee plant tissues. Coffee Sci, Lavras, pp 284–293
Deepa K, Sheeja TK, Santhi R, Sasikumar B, Cyriac A, Deepesh PV, Prasath D (2014) A simple and efficient protocol for isolation of high quality functional RNA from different tissues of turmeric (Curcuma longa L). Physiol Mol Biol Plants 20(2):263–271. https://doi.org/10.1007/s12298-013-0218-y
Graham GC (1993) A method for extraction of total RNA from Pinus radiata and other conifers. Plant Mol Biol Rep 11(1):32–37
Logemann J, Schell J, Willmitzer L (1987) Improved method for the isolation of RNA from plant tissues. Anal Biochem 163:16–20
Manning K (1991) Isolation of nucleic acids from plants by differential solvent precipitation. Anal Biochem 195(1):45–50
Mishra MK, Slater A (2012) Recent advances in the genetic transformation of coffee. Hindawi Publishing Corporation. Biotechnol Res Int. https://doi.org/10.1155/2012/580857
Razafinarivo NJ, Guyol R, Davis AP, Couturon E, Hamon S, Crouzillat D, Rigoreau M, Tranchant CD, Poncet V, Kochko ADE, Rakotomalala JJ, Hamon P (2013) Genetic structure and diversity of coffee (Coffea) across Africa and the Indian Ocean islands revealed using microsatellites. Ann Bot 111:229–248
Sah SK, Kaur G, Kaur A (2014) Rapid and reliable method of high-quality RNA extraction from diverse plants. Am J Plant Sci 5:3129–3139
Tan SC, Yiap BC (2009) DNA, RNA, and protein extraction: the past and the present. J Biomed Biotechnol. https://doi.org/10.1155/2009/574398
Vasanthaiah HKN, Katam R, Sheikh MB (2008) Efficient protocol for isolation of functional RNA from different grape tissue rich in polyphenols and polysaccharides for gene expression studies. Electron J Biotechnol 11(3):1–8
Zarei A, Zamani Z, Mousavi A, Fatahi R, Alavijeh MK, Dehsara B, Salami SA (2012) An effective protocol for isolation of high-quality RNA from pomegranate seeds. Asian Aust J Plant Sci Biotechnol 6:32–37
Acknowledgements
The authors thank Dr. Y. Raghuramulu, Director of Research, Central Coffee Research Institute, India, for providing laboratory facilities and encouragement. Funding support from Coffee Board, Govt. of India, is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
The corresponding author designed the experiment and wrote the manuscript. The first and second authors carried out the experiments and participated in manuscript preparation. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Ethical standards
The experiment complied with the ethical standards.
Rights and permissions
About this article
Cite this article
Huded, A.K.C., Jingade, P. & Mishra, M.K. A rapid and efficient SDS-based RNA isolation protocol from different tissues of coffee. 3 Biotech 8, 183 (2018). https://doi.org/10.1007/s13205-018-1209-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-018-1209-z