Abstract
Purpose
This study aimed to isolate effective erythromycin-degrading fungi and determine the characteristics and pathway of degradation.
Methods
Erythromycin-degrading fungi were isolated from erythromycin-contaminated samples using a standard enrichment and isolation method. The degradation characteristics were investigated in mineral salt medium (MSM) with erythromycin as a sole carbon source. Key degradation intermediates were analyzed by high performance liquid chromatography–mass spectrometry (HPLC–MS) and used to deduce the erythromycin degradation pathway of strain RJJ-2.
Results
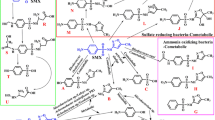
A novel erythromycin-degrading fungus RJJ-2, was isolated from a contaminated sample. Based on its morphology and internal transcribed spacer (ITS) sequence, the strain was 100% similar to P. oxalicum (MN759650) and named P. oxalicum RJJ-2. The strain RJJ-2 degraded 84.88% erythromycin after 96-h incubation at 35 °C and pH 6.0 in MSM with erythromycin (100 mg L−1) as the sole carbon source. Optimal degradation conditions for P. oxalicum RJJ-2 were 35 °C, and pH 6.0 with 0.1% ammonium sulfate supplementation. HPLC–MS analysis indicated that the main degradation intermediates were 3-depyranosyloxy erythromycin A, cladinose, desosamine, and 7,12-dyhydroxy-6-deoxyerythronolide B. It was inferred that the erythromycin was degraded to 3-depyranosyloxy erythromycin A by a glycoside hydrolase in the initial reaction.
Conclusion
This study demonstrated that P. oxalicum RJJ-2 is a novel erythromycin-degrading strain, which can provide a new eco-friendly and cost-effective approach for the disposal of erythromycin fermentation wastes and other hazardous chemicals.
Graphic Abstract







Similar content being viewed by others
Abbreviations
- PCR:
-
Polymerase chain reaction
- HPLC–MS:
-
High performance liquid chromatography–mass spectrometry
- AFWs:
-
Antibiotic fermentation wastes
- ITS:
-
Internal transcribed spacer
References
Chu, L., Chen, D., Wang, J., Yang, Z., Shen, Y.: Degradation of antibiotics and antibiotic resistance genes in erythromycin fermentation residues using radiation coupled with peroxymonosulfate oxidation. Waste Manage. 96, 190–197 (2019). https://doi.org/10.1016/j.wasman.2019.07.031
Klein, E.Y., Van Boeckel, T.P., Martinez, E.M., Pant, S., Gandra, S., Levin, S.A., Goossens, H., Laxminarayan, R.: Global increase and geographic convergence in antibiotic consumption between 2000 and 2015. Proc. Natl. Acad. Sci. USA 115(15), E3463–E3470 (2018)
Shen, Y., Chu, L., Zhuan, R., Xiang, X., Sun, H., Wang, J.: Degradation of antibiotics and antibiotic resistance genes in fermentation residues by ionizing radiation: a new insight into a sustainable management of antibiotic fermentative residuals. J Environ. Manage. 232, 171–178 (2019). https://doi.org/10.1016/j.jenvman.2018.11.050
Ma, D., Zhang, G., Areeprasert, C., Li, C., Shen, Y., Yoshikawa, K.: Characterization of NO emission in combustion of hydrothermally treated antibiotic mycelial residue. Chem. Eng. J. 284, 708–715 (2016)
Tang, R., Yuan, S., Chen, F., Zhan, X., Wang, W., Hu, Z.: Effects of roxarsone and sulfadiazine on biogas production and their degradation during anaerobic digestion. Int. Biodeterior. Biodegradation 140, 113–118 (2019). https://doi.org/10.1016/j.ibiod.2019.04.001
Sun, Q., Bai, Y., Zhao, C., Xiao, Y., Wen, D., Tang, X.: Aerobic biodegradation characteristics and metabolic products of quinoline by a Pseudomonas strain. Biores. Technol. 100(21), 5030–5036 (2009). https://doi.org/10.1016/j.biortech.2009.05.044
Liao, H., Zhao, Q., Cui, P., Chen, Z., Yu, Z., Geisen, S., Friman, V.P., Zhou, S.: Efficient reduction of antibiotic residues and associated resistance genes in tylosin antibiotic fermentation waste using hyperthermophilic composting. Environ. Int. 133, 105203 (2019). https://doi.org/10.1016/j.envint.2019.105203
Cizmek, L., Sabic, M., Mestrovic, E., Domanovac, M.: Biodegradation of Erythromycin with Environmental Microorganism Pseudomonas aeruginosa 3011. Chem. Biochem. Eng. Q. 29, 367–373 (2015)
Zhang, W., Qiu, L., Gong, A., Yuan, X.: Isolation and characterization of a high-efficiency erythromycin A-degrading Ochrobactrum sp. strain. Marine Pollut. Bull. 114(2), 896–902 (2017). https://doi.org/10.1016/j.marpolbul.2016.10.076
Kim, H.E., Park, K.R.: Purification and Characterization of an Esterase from Acinetobacter lwoffii I6C–1. Curr. Microbiol. 44(6), 401–405 (2002)
Wen, X., Jia, Y., Li, J.: Enzymatic degradation of tetracycline and oxytetracycline by crude manganese peroxidase prepared from Phanerochaete chrysosporium. J. Hazard. Mater. 177(1–3), 924–928 (2010). https://doi.org/10.1016/j.jhazmat.2010.01.005
Buchicchio, A., Bianco, G., Sofo, A., Masi, S., Caniani, D.: Biodegradation of carbamazepine and clarithromycin by Trichoderma harzianum and Pleurotus ostreatus investigated by liquid chromatography–high-resolution tandem mass spectrometry (FTICR MS-IRMPD). Sci. Total Environ. 557–558(Jul 1), 733–739 (2016)
Olicon-Hernandez, D.R., Camacho-Morales, R.L., Pozo, C., Gonzalez-Lopez, J., Aranda, E.: Evaluation of diclofenac biodegradation by the ascomycete fungus Penicillium oxalicum at flask and bench bioreactor scales. Sci. Total Environ. 662, 607–614 (2019). https://doi.org/10.1016/j.scitotenv.2019.01.248
Caniani, D., Caivano, M., Pascale, R., Bianco, G., Mancini, I.M., Masi, S., Mazzone, G., Firouzian, M., Rosso, D.: CO2 and N2O from water resource recovery facilities: evaluation of emissions from biological treatment, settling, disinfection, and receiving water body. Sci. Total Environ. 648, 1130–1140 (2019). https://doi.org/10.1016/j.scitotenv.2018.08.150
Birolli, W.G., Vacondio, B., Alvarenga, N., Seleghim, M.H.R., Porto, A.L.M.: Enantioselective biodegradation of the pyrethroid (+/-)-lambda-cyhalothrin by marine-derived fungi. Chemosphere 197, 651–660 (2018). https://doi.org/10.1016/j.chemosphere.2018.01.054
Ren, J., Fan, B., Huhetaoli, N., Niu, D., Gu, Y., Li, C.: Biodegradation of waste cooking oils by Klebsiella quasivariicola IUMR-B53 and characteristics of its oil-degrading enzyme. Waste Biomass Valoriz. (2020). https://doi.org/10.1007/s12649-020-01097-z
Zuo, S.S., Niu, D.Z., Ning, T.T., Zheng, M.L., Jiang, D., Xu, C.C.: Protein enrichment of sweet potato beverage residues mixed with peanut shells by Aspergillus oryzae and Bacillus subtilis using central composite design. Waste Biomass Valoriz. 9(5), 835–844 (2017). https://doi.org/10.1007/s12649-017-9844-x
Liu, M., Feng, P., Kakade, A., Yang, L., Chen, G., Yan, X., Ni, H., Liu, P., Kulshreshtha, S., Abomohra, A.E., Li, X.: Reducing residual antibiotic levels in animal feces using intestinal Escherichia coli with surface-displayed erythromycin esterase. J. Hazard. Mater. 388, 122032 (2020). https://doi.org/10.1016/j.jhazmat.2020.122032
Ding, T., Zhou, Y., Qin, J.J., Yang, L.J., Zhang, W.D., Shen, Y.H.: Chemical constituents from wetland soil fungus Penicillium oxalicum GY1. Fitoterapia 142, 104530 (2020). https://doi.org/10.1016/j.fitote.2020.104530
Li, F., Yu, D., Lin, X., Liu, D., Xia, H., Chen, S.: Biodegradation of poly(epsilon-caprolactone) (PCL) by a new Penicillium oxalicum strain DSYD05-1. World J. Microbiol. Biotechnol. 28(10), 2929–2935 (2012). https://doi.org/10.1007/s11274-012-1103-5
Pareek, N., Vivekanand, V., Dwivedi, P., Singh, R.P.: Penicillium oxalicum SAEM-51: a mutagenised strain for enhanced production of chitin deacetylase for bioconversion to chitosan. New Biotechnol. 28(2), 118–124 (2011). https://doi.org/10.1016/j.nbt.2010.09.009
Satti, S.M., Shah, Z., Luqman, A., Hasan, F., Osman, M., Shah, A.A.: Biodegradation of poly(3-hydroxybutyrate) and poly(3-hydroxybutyrate-co-3-hydroxyvalerate) by newly isolated Penicillium oxalicum SS2 in soil microcosms and partial characterization of extracellular depolymerase. Curr. Microbiol. (2020). https://doi.org/10.1007/s00284-020-01968-7
Tian, H., Ma, Y.J., Li, W.Y., Wang, J.W.: Efficient degradation of triclosan by an endophytic fungus Penicillium oxalicum B4. Environ. Sci. Pollut. Res. Int. 25(9), 8963–8975 (2018). https://doi.org/10.1007/s11356-017-1186-5
Fan, C., He, J.: Proliferation of antibiotic resistance genes in microbial consortia of sequencing batch reactors (SBRs) upon exposure to trace erythromycin or erythromycin-H2O. Water Res. 45(10), 1–3106 (2011)
Mao, F., Liu, C., He, M., Shi, W., Xue, G., Gao, P.: Isolation and identification of an erythromycin degradation bacterium and study on its biodegradation characteristics. Environ. Sci. Technol. 36(7), 9–12 (2013)
Akbar, S., Hasan, F., Nadhman, A., Khan, S., Shah, A.A.: Production and purification of poly (3-hydroxybutyrate-co-3-hydroxyvalerate) degrading enzyme from Streptomyces sp. AF-111. J. Polym. Environ. 21(4), 1109–1116 (2013)
Feng, W., Wei, Z., Song, J., Qin, Q., Yu, K., Li, G., Zhang, J., Wu, W., Yan, Y.: Hydrolysis of nicosulfuron under acidic environment caused by oxalate secretion of a novel Penicillium oxalicum strain YC-WM1. Sci. Rep. 7(1), 647 (2017). https://doi.org/10.1038/s41598-017-00228-2
Morar, M., Pengelly, K., Koteva, K., Wright, G.D.: Mechanism and diversity of the erythromycin esterase family of enzymes. Biochemistry 51(8), 1740–1751 (2012). https://doi.org/10.1021/bi201790u
Olicón-Hernández, D.R., Jesús, G.L., Elisabet, A.: Overview on the biochemical potential of filamentous fungi to degrade pharmaceutical compounds. Front. Microbiol. 8, 1792 (2017)
Hansen, L.H., Mauvais, P., Douthwaite, S.: The macrolide-ketolide antibiotic binding site is formed by structures in domains II and V of 23 S ribosomal RNA. Mol Microbiol 31, 623–631 (1999)
Acknowledgements
This study was financially supported by Yili Chuanning Biotechnology Co., Ltd. (Grant Nos. 2019K1238 and 2019K1237); Hebei Cixin Environmental Technology Co., Ltd. China. (Grant No. 2018K0948); Shaanxi Xintiandi Grass Industry Co., Ltd. China. (Grant No. 2018K0947); Research on the Development Strategy of Bioenergy and Geothermal Energy Engineering Science and Technology for 2035 (Grant No. 2019-ZCQ-04) and Science Foundation of Changzhou University (Grant No. ZMF18020316).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ren, J., Wang, Z., Deng, L. et al. Degradation of Erythromycin by a Novel Fungus, Penicillium oxalicum RJJ-2, and the Degradation Pathway. Waste Biomass Valor 12, 4513–4523 (2021). https://doi.org/10.1007/s12649-021-01343-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-021-01343-y