Abstract
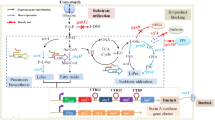
The effect of the response surface methodology was applied to the production of Caulobacter crescentus (strain NA1000) β-xylosidases using corn cob. The components of the medium that presented the greatest influence on the enzyme production were chosen for optimization including the concentration of the corn cob residue and temperature variation. Optimal concentrations were determined by a central composite rotational design and a combination of 3.5% (w/v) corn cob concentration and 27 °C temperature was found to be optimal. When C. crescentus was cultivated using the optimized conditions, a maximum activity of 393.36 U/mL of β-xylosidases was achieved in 24-h cultures with a yield of 95% in real test conditions compared to the predicted one. In parallel, there was an increase of 3.6 times in the production of intracellular xylanases when compared to cultures without statistical application. In the C. crescentus genome, 5 genes that encode β-xylosidases are present. In order to evaluate which of them would be induced in the optimized conditions, the quantitative expression (qPCR) of the xynB1–xynB5 genes was evaluated in the presence of 1 or 3.5% corn cob (w/v) and surprisingly, all showed a constitutive expression in relation to the control. Assays of Western Blot performed with a polyclonal antiserum against C. crescentus β-xylosidase II in the optimized condition also did not show mass variation of the referred protein. These data strongly suggest that post-transcriptional controls are operating in the induced condition to increase the activity of C. crescentus β-xylosidases I-V, but β-xylosidase II. To our knowledge, this is the first time that these data are reported in literature for a bacterial system.






Similar content being viewed by others
References
Bosetto, A., Justo, P.I., Zanardi, B., Venzon, S.S., Graciano, L., Santos, E.L., Simão, R.C.G.: Research progress concerning fungal and bacterial beta-xylosidases. Appl. Biochem. Biotechnol. 178, 766–795 (2016)
Corrêa, J.M., Christi, D., Della Torre, C.L., Henn, C., Da Conceição-Silva, J.L., Kadowaki, M.K., Simão, R.C.G.: High levels of & #x03B2;-xylosidase in Thermomyces lanuginosus: potential use for saccharification. Braz. J. Microbiol. 47, 680–690 (2016)
Marks, M.E., Castro-Rojas, C.M., Teiling, C., Du, L., Kapatral, V., Walunas, T.L., Crosson, S.: The genetic basis of laboratory adaptation in Caulobacter crescentus. J. Bacteriol. 192, 3678–3688 (2010)
Prade, R.A.: Xylanases: from biology to biotechnology. Biotechnol. Genet. Eng. Rev. 13, 101–131 (1996)
Sharma, H.K., Xu, C., Qin, W.: Biological pretreatment of lignocellulosic biomass for biofuels and bioproducts: an overview. Waste Biomass Valoriz. 10, 235–251 (2019)
Graciano, L., Corrêa, J.M., Gandra, R.F., Seixas, F.A.V., Kadowaki, M.K., Sampaio, S.C., da Conceição-Silva, Osaku, C.A., Simão, R.C.G.: The cloning, expression, purification, characterization and modeled structure of Caulobacter crescentus β-xylosidase I. World J. Microbiol. Biotechnol. 28, 2879–2888 (2012)
Corrêa, J.M., Graciano, L., Abrahão, J., Loth, E.A., Gandra, R.F., Kadowaki, M.K., Henn, C., Simão, R.C.G.: Expression and characterization of a GH39 β-xylosidase II from Caulobacter crescentus. Appl. Biochem. Biotechnol. 168, 2218–2229 (2012)
Justo, P.I., Corrêa, J.M., Maller, A., Kadowaki, M.K., Da Conceição-Silva, J.L., Gandra, R.F., Simão, R.C.G.: Analysis of the xynB5 gene encoding a multifunctional GH3-BglX β-glucosidase-β-Xylosidases-α-Arabinosidase member in Caulobacter crescentus. Antonie Van Leeuwenhoek 108, 993–1007 (2015)
Graciano, L., Corrêa, J.M., Vieira, F.G.N., Bosetto, A., Loth, E.A., Kadowaki, M.K., Gandra, R.F., Simão, R.C.G.: Cloning and expression of the xynA1 gene encoding a xylanase of the GH10 group in Caulobacter crescentus. Appl. Biochem. Biotechnol. 175, 3915–3929 (2015)
Li, Q., Wu, T., Qi, Z., Zhao, L., Pei, J., Tang, F.: Characterization of a novel thermostable and xylose-tolerant GH 39 β-xylosidase from Dictyoglomus thermophilum. BMC Biotechnol. 18(1), 29 (2019)
Zhang, S., Xie, J., Zhao, L., Pei, J., Su, E., Xiao, W., Wang, Z.: Cloning, overexpression and characterization of a thermostable β-xylosidase from Thermotoga petrophila and cooperated transformation of ginsenoside extract to ginsenoside 20(S)-Rg3 with a β-glucosidase. Bio-organ. Chem. 85, 159–167 (2019)
Evinger, M., Agabian, N.: Envelope-associated nucleoid from Caulobacter crescentus stalked and swarmer. J. Bacteriol. 132, 294–301 (1977)
Johnson, R.C., Ely, B.: Isolation of spontaneously derived mutants of Caulobacter crescentus. Genetics 86, 25–32 (1977)
Corrêa, J.M., Mingori, M.R., Gandra, R.F., Loth, E.A., Seixas, F.A.V., Simão, R.C.G.: Depletion of the xynB2 gene upregulates β-xylosidase expression in C. crescentus. Appl. Biochem. Biotechnol. 172, 1085–1097 (2014)
Miller, G.L.: Use of dinitrosalicylic acid reagent for determination of reducing sugars. Anal. Chem. 31, 426–428 (1959)
Bradford, M.M.: A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976)
Laemmli, U.K.: Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970)
Towbin, H., Staehelin, T., Gordon, J.: Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc. Natl. Acad. Sci. 76, 4350–4354 (1979)
Sambrook, J., Fritsch, E.F., Maniatis, T.: Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York (1989)
Bustin, S.A., Benes, V., Garson, J.A., Hellemans, J., Huggett, J., Kubista, M., Mueller, R., Nolan, T., Pfaffl, M.W., Shipley, G.L., Vandesompele, J., Wittwer, C.T.: The Miqe Guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009)
Hottes, A.K., Meewan, M., Yang, D., Arana, N., Romero, P., Mcadams, H.H., Stephens, C.: Transcriptional profiling of Caulobacter crescentus during growth on complex and minimal media. J. Bacteriol. 186, 1448–1461 (2004)
Marcolongo, L., La Cara, F., del Monaco, G., Paixão, S.M., Alves, L., Marques, I.P., Ionata, E.: A novel β-xylosidase from Anoxybacillus sp. 3 M towards an improved agro-industrial residues saccharification. Int. J. Biol. Macromol. 122, 1224–1234 (2019)
Vieria, F.G.N., Christ, D., Graciano, L., Corrêa, J.M., Kadowaki, M.K., da Conceição-Silva, J.L., Gandra, R.F., Maller, A., Polizeli, M.L.T.M., Simão, R.C.G.: Experimental design for optimization of β-xylosidase production by A. fumigatus isolated from the Atlantic Forest (Brazil). J. Adv. Biol. Biotechnol. 21(3), 1–16 (2019)
Neumann, A.P., Weimer, P.J., Suen, G.: A global analysis of gene expression in Fibrobacter succinogenes S85 grown on cellulose and soluble sugars at different growth rates. Biotechnol. Biofuels 11, 295 (2018)
Midorikawa, G.E.O., Correa, C.L., Noronha, E.F., Ferreira-Filho, E.X., Togawa, R.C., Costa, M.M.C., Silva-Junior, O.B., Grynberg, P., Miller, R.N.G.: Analysis of the transcriptome in Aspergillus tamarii during enzymatic degradation of sugarcane bagasse. Front. Bioeng. Biotechnol. 6(123), 1–17 (2018)
Santos, C.R., Polo, C.C., Corrêa, J.M., Simão, R.C.G., Seixas, F.A.V., Murakami, M.T.: Accessory domain changes accessibility and molecular topography of the catalytic interface in monomeric GH39 & #x03B2;-xylosidases. Acta Crystallogr. Sect. D 68, 1339–1345 (2012)
Acknowledgements
J.M. Corrêa was a PNPD/CAPES scholar. E.L. Santos was a CNPq scholar. R.C.G. Simão was a productivity scholar at Fundação Araucária.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Corrêa, J.M., dos Santos, E.L., Simões, M.R. et al. Optimization of C. crescentus β-Xylosidases and Expression of xynB1–5 Genes in Response to Agro-Industrial Waste. Waste Biomass Valor 11, 6169–6178 (2020). https://doi.org/10.1007/s12649-019-00881-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-019-00881-w