Abstract
Purpose
The aim of this work was to evaluate the use of chitin-rich mushroom (Agaricus bisporus) by-products chitin/glucan enriched fraction (M-Ch/G-F) as main carbon source for the production of chitinases by three different microorganisms (Trichoderma harzianum, Trichoderma atroviride and Bacillus licheniformis), in an attempt to obtain these enzymes using a cheap and abundant fermentation medium.
Methods
Microorganisms were grown in submerged fermentation using different media formulated with chitin powder (Chp), colloidal chitin (Chc) and M-Ch/G-F as the main carbon source, respectively. Enzyme productivity and secretion (secretome) was studied by electrophoretic and proteomic methods.
Results
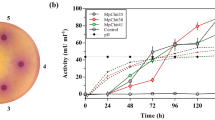
All microorganisms produced higher chitinase activity in a medium formulated with M-Ch/G-F as a carbon source compared to medium formulated with Chp or CHc. T. harzianum showed the highest chitinase productivity (261.5 mU L−1 per day). Chitinase production was monitored by electrophoretic and proteomic methods. Electrophoretic method allowed the detection of 28 different proteins—three different chitinases with 82, 50 and 31 kDa. Proteomic analysis could identify 161 different proteins: 60 of them hydrolases, and 80% having glycolytic activity—5 of them were chitinases. These results show that cultivation of T. harzianum in a cheap and abundant fermentation medium represents a good procedure for large scale production of chitinases.
Conclusions
Our results show that cultivation of T. harzianum in a culture medium formulated with M-Ch/G-F, a cheap and abundant fermentation medium, is a good procedure for large scale production of glycosidases, particularly chitinases within a relatively short cultivation period of 6 days.
Graphical Abstract





Similar content being viewed by others
Abbreviations
- M-Ch/G-F:
-
Mushroom chitin/glucan enriched fraction
- Mm:
-
Minimal medium
- GlcNAc:
-
N-acetylglucosamine
- Chp :
-
Chitin powder
- Chc :
-
Colloidal chitin
- PDB:
-
Potato dextrose broth
References
Shaikh, S.A., Deshpande, M.V.: Chitinolytic enzymes: their contribution to basic and applied research. World J. Microbiol. Biotechnol. 9, 468–475 (1993). https://doi.org/10.1007/BF00328035
Barboza-Corona, J.E., Reyes-Rios, D.M., Salcedo-Hernández, R., Bideshi, D.K.: Molecular and biochemical characterization of an endoquitinase (ChiA-HD73) from Bacillus thuringiensis subsp. Kurstaki HD-73. Mol. Biotechnol. 39, 27–29 (2008). https://doi.org/10.1007/s12033-007-9025-4
Tsujibo, H., Orikoshi, H., Shiotani, K., Hayashi, M., Umeda, J., Miyamoto, K., Imada, C., Okami, Y., Inamori, Y.: Characterization of chitinase C from marine bacterium Alteromonas sp. strain 0-7, and its corresponding gene and domain structure. Appl. Environ. Microbiol. 64, 472–478 (1998)
Fortuna-González, J.M.: Caracterización bioquímica y molecular de quitinasas en cepas mexicanas de B. thuringiensis. Doctoral thesis. Instituto Politécnico Nacional. Ciudad de México, México, (2010)
Sastoque, C.: Aislamiento y selección de microorganismos productores de quitinasas a partir de residuos de concha de camarón con potencial biocontrolador. Tesis para optar el título de microbiólogo industrial, agrícola y veterinario. Pontificia Universidad Javeriana. Bogotá, Colombia (2005)
Sahai, A.S., Manocha, M.S.: Chitinases of fungi and plants: Their involvement in morphogenesis and host-parasite interaction. Microbiol. 11, 317–338 (1993). https://doi.org/10.1111/j.1574-6976.1993.tb00004.x
Gessesse, A.: The use of Nug meal as a low cost substrate for the production of alkaline protease by the alkaliphilic Bacillus sp. AR-009 and some properties of the enzyme. Bioresour. Technol. 62, 59–61 (1997). https://doi.org/10.1016/S0960-8524(97)00059-X
Sneath, P.H.A., Mair, N.S., Sharpe, M.E., Holt, J.G.: Bergey’s Manual of Sytematic Bacteriology. vol. 2. Williams & Wilkins, Baltimore (1986)
Schallmey, M., Singh, A., Ward, O.P.: Developments in the use of Bacillus species for industrial production. Can. J. Microbiol. 50, 1–17 (2004). https://doi.org/10.1139/w03-076
Ramírez, M.G., Avelizapa, L.I., Avelizapa, N.G.R., Camarillo, R.C.: Colloidal chitin stained with Remazol Brilliant Blue R, a useful substrate to select chitinolytic microorganisms and to evaluate chitinases. J. Microbiol. Methods 56, 213–219 (2004). https://doi.org/10.1016/j.mimet.2003.10.011
Bautista, J.: CHAMPI-D. Project RTC-2015-4039-2 (2015)
Görs, S., Schumann, R., Häubner, N., Karsten, U.: Fungal and algal biomass in biofilms on artificial surfaces quantified by ergosterol and chlorophyll a as biomarkers. Int. Biodeterior. Biodegradation 60, 50–59 (2007). https://doi.org/10.1016/j.ibiod.2006.10.003
Miller, G.L.: Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 31, 426–428 (1959)
Bradford, M.M.: A rapid and sensitive method for the quantitation microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976). https://doi.org/10.1016/0003-2697(76)90527-3
Yamabhai, M., Emrat, S., Sukasem, S., Pesatcha, P., Jaruseranee, N., Buranabanyat, B.: Secretion of recombinant Bacillus hydrolytic enzymes using Escherichia coli expression systems. Biotechnol. J. 133, 50–57 (2008) https://doi.org/10.1016/j.jbiotec.2007.09.005
Carbonero-Aguilar, P. Estudio de la oxidación de proteínas en ratas con encefalopatía hepática: una aproximación Proteómica. Doctoral thesis. Universidad de Sevilla. Sevilla, Spain (2012)
Craig, R., Beavis, R.C.: A method for reducing the time required to match protein sequences with tandem mass spectra. Rapid Commun. Mass Spectrom. 17, 2310–2316 (2003). https://doi.org/10.1002/rcm.1198
Nesvizhskii, A.I., Keller, A., Kolker, E., Aebersold, R.: A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003). https://doi.org/10.1021/ac0341261
de Lima, F.B., Félix, C., Osório, N., Vitorino, A.A., Domingues, P., Correia, A., da Silva-Ribeiro, R.T., Esteves, A.C.: Secretome analysis of Trichoderma atroviride T17 biocontrol of Guignardia citricarpa. Biocontrol Sci. 99, 38–46 (2016). https://doi.org/10.1016/j.biocontrol.2017.01.003
Giese, E.,C., Corradi da Silva, M.I., Barbosa, A.M.: Fungal glucanase: production and application of β-1, 3 and β-1, 6-glucanases. Rev. Biotechnol. Cienc. Desenvol. 30, 97–104 (2003)
Mohamed, H.A.A., Wafaa, M.H., Attallah, A.G.: Genetic enhancement of Trichoderma viride to over produce different hydrolytic enzymes and their biocontrol potentially against root rot and white mold diseases in plants. Agric. Biol. J. N. Am. 1, 273–284 (2010)
Latgé, J.P.: Tasting the fungal cell wall. Cell. Microbiol. 12, 863–872 (2010). https://doi.org/10.1111/j.1462-5822.2010.01474.x
Rana, I.A., Loerz, H., Schaefer, W., Becker, D.: Over expression of chitinase and chitosanase genes from Trichoderma harzianum under constitutive and induced promoters in order to increase disease resistance in wheat (Triticum aestivium). Mol. Plant Breed. 3, 37–49 (2012). https://doi.org/10.5376/mpb.2012.03.0004
Gajera, H.P., Vakharia, D.N.: Production of lytic enzymes by Trichoderma isolated during in vitro antagonism with Aspergilus niger, the causal agent of collar rot of peanut. Braz. J. Microbiol. 43, 43–52 (2012). https://doi.org/10.1590/S1517-83822012000100005
Kubicek, C.P., Mach, R.I., Peterbauer, C.K., Lorito, M.: Trichoderma: from genes to biocontrol. Plant Pathol. J. 83, 11–23 (2001). http://www.jstor.org/stable/41998018
Herpoël-Gimbert, I., Margeot, A., Dolla, A., Jan, G., Mollé, D., Lignon, S., Mathis, H., Sigoillot, J.C., Monot, F., Asther, M.: Comparative secretome analyses of two Trichoderma reesei RUT-C30 and CL847 hypersecretory strains. Biotechnol. Biofuels 1, 1–12 (2008). https://doi.org/10.1186/1754-6834-1-18
Adav, S.S., Chao, L.T., Sze, S.K.: Quantitative secretomic analysis of Trichoderma reesei strains reveals enzymatic composition for lignocellulosic biomass degratation. Mol. Cell Proteomics. 11, 1–15 (2012). https://doi.org/10.1074/mcp.M111.012419
Bendtsen, J.D., Nielsen, H., Von Heijne, G., Brunak, S.: Improved prediction of signal peptides: signalP 3.0. J. Mol. Biol. 340, 783–795 (2004). https://doi.org/10.1016/j.jmb.2004.05.028
Acknowledgements
We are grateful to the Spanish Ministry of Science and Innovation for the financial support of this work (Project RTC-2015-4039-2), which has partial financial support from the FEDER funds of the European Union. J.A. del Campo was supported by Nicolás Monardes Program from Servicio Andaluz de Salud (SAS). The authors thank Paula Bautista for assistance in the preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Urbina-Salazar, A., Inca-Torres, A.R., Falcón-García, G. et al. Chitinase Production by Trichoderma harzianum Grown on a Chitin-Rich Mushroom Byproduct Formulated Medium. Waste Biomass Valor 10, 2915–2923 (2019). https://doi.org/10.1007/s12649-018-0328-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-018-0328-4