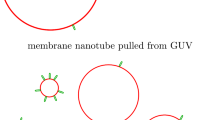
Abstract
The conformational free energy landscape of a system is a fundamental thermodynamic quantity of importance particularly in the study of soft matter and biological systems, in which the entropic contributions play a dominant role. While computational methods to delineate the free energy landscape are routinely used to analyze the relative stability of conformational states, to determine phase boundaries, and to compute ligand-receptor binding energies its use in problems involving the cell membrane is limited. Here, we present an overview of four different free energy methods to study morphological transitions in bilayer membranes, induced either by the action of curvature remodeling proteins or due to the application of external forces. Using a triangulated surface as a model for the cell membrane and using the framework of dynamical triangulation Monte Carlo, we have focused on the methods of Widom insertion, thermodynamic integration, Bennett acceptance scheme, and umbrella sampling and weighted histogram analysis. We have demonstrated how these methods can be employed in a variety of problems involving the cell membrane. Specifically, we have shown that the chemical potential, computed using Widom insertion, and the relative free energies, computed using thermodynamic integration and Bennett acceptance method, are excellent measures to study the transition from curvature sensing to curvature inducing behavior of membrane associated proteins. The umbrella sampling and WHAM analysis has been used to study the thermodynamics of tether formation in cell membranes and the quantitative predictions of the computational model are in excellent agreement with experimental measurements. Furthermore, we also present a method based on WHAM and thermodynamic integration to handle problems related to end-point-catastrophe that are common in most free energy methods.






Similar content being viewed by others
Notes
Calculated as (nanocarrier radius + length of the antibody + length of the receptor + \(d_{0}\)).
References
Israelachvili, J.N.: Intermolecular and Surface Forces, 3rd edn. Academic Press, Boston (2011)
Escribá, P.V., González-Ros, J.M., Goñi, F.M., Kinnunen, P.K.J., Vigh, L., Sánchez-Magraner, L., Fernández, A.M., Busquets, X., Horváth, I., Barceló-Coblijn, G.: Membranes: a meeting point for lipids, proteins and therapies. J. Cell. Mol. Med. 12(3), 829 (2008). doi:10.1111/j.1582-4934.2008.00281.x
Singer, S.J., Nicolson, G.L.: The fluid mosaic model of the structure of cell membranes. Science 175(4023), 720 (1972). http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=1972Sci...175.720S&link_type=EJOURNAL
Edidin, M.: Lipids on the frontier: a century of cell-membrane bilayers. Nat. Rev. Mol. Cell Biol. 4(5), 414 (2003). doi:10.1038/nrm1102. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=12728275&retmode=ref&cmd=prlinks
Engelman, D.M.: Membranes are more mosaic than fluid. Nat. Cell Biol. 438(7068), 578 (2005). doi:10.1038/nature04394. http://www.nature.com/doifinder/10.1038/nature04394
Conner, S.D., Schmid, S.L.: Regulated portals of entry into the cell. Nature 422(6927), 37 (2003). doi:10.1038/nature01451. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=2003Natur.422...37C&link_type=ABSTRACT
Doherty, G.J., McMahon, H.T.: Mechanisms of endocytosis. Annu. Rev. Biochem. 78(1), 857 (2009). doi:10.1146/annurev.biochem.78.081307.110540
Ewers, H., Helenius, A.: Lipid-mediated endocytosis. Cold Spring Harb. Perspect. Biol. 3(8), a004721 (2011). doi:10.1101/cshperspect.a004721
Canton, I., Battaglia, G.: Endocytosis at the nanoscale. Chem. Soc. Rev. 41(7), 2718 (2012). doi:10.1039/c2cs15309b
Kholodenko, B.N.: Cell-signalling dynamics in time and space. Nature 7(3), 165 (2006). doi:10.1038/nrm1838
Sorkin, A., von Zastrow, M.: Endocytosis and signalling: intertwining molecular networks. Nat. Rev. Mol. Cell Biol. 10(9), 609 (2009). doi:10.1038/nrm2748
Sheetz, M.P.: Cell control by membrane-cytoskeleton adhesion. Nat. Rev. Mol. Cell Biol. 2(5), 392 (2001). doi:10.1038/35073095
Ananthakrishnan, R., Ehrlicher, A.: The forces behind cell movement. Int. J. Biol. Sci. 3(5), 303 (2007). http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=17589565&retmode=ref&cmd=prlinks
Keren, K.: Cell motility: the integrating role of the plasma membrane. Eur. Biophys. J. 40(9), 1013 (2011). doi:10.1007/s00249-011-0741-0
Chaikin, P.M., Lubensky, T.C.: Principles of Condensed Matter Physics. Cambridge University Press, Cambridge (2000). http://books.google.co.in/books?id=P9YjNjzr9OIC
Frenkel, D., Smit, B.: Understanding Molecular Simulation : From Algorithms to Applications, 2nd edn. Academic Press, New York (2001). http://www.worldcat.org/isbn/0122673514
Seifert, U.: Configurations of fluid membranes and vesicles. Adv. Phys. 46(1), 13 (1997). http://gateway.webofknowledge.com/gateway/Gateway.cgi?GWVersion=2&SrcAuth=mekentosj&SrcApp=Papers&DestLinkType=FullRecord&DestApp=WOS&KeyUT=A1997WE91800002
Tieleman, D.P., Marrink, S.J., Berendsen, H.J.: A computer perspective of membranes: molecular dynamics studies of lipid bilayer systems. Biochim. Biophys. Acta (BBA) Rev. Biomembr. 1331(3), 235 (1997). http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=9512654&retmode=ref&cmd=prlinks
Venturoli, M., Maddalena Sperotto, M., Kranenburg, M., Smit, B.: Mesoscopic models of biological membranes. Phys. Rep. 437(1), 1 (2006). doi:10.1016/j.physrep.2006.07.006. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=2006PhR...437....1V&link_type=ABSTRACT
Ayton, G.S., Voth, G.A.: Multiscale simulation of protein mediated membrane remodeling. Semin. Cell Dev. Biol. 21(4), 357 (2010). doi:10.1016/j.semcdb.2009.11.011. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=19922811&retmode=ref&cmd=prlinks
Shinoda, W., DeVane, R., Klein, M.L.: Computer simulation studies of self-assembling macromolecules. Curr. Opin. Struct. Biol. 22(2), 175 (2012). doi:10.1016/j.sbi.2012.01.011
Bradley, R.P., Radhakrishnan R.: Coarse-grained models for protein-cell membrane interactions. Polymers 5(3), 890 (2013). doi:10.3390/polym5030890. http://www.mdpi.com/2073-4360/5/3/890/
Ramakrishnan, N., Sunil Kumar, P.B., Radhakrishnan, R.: Mesoscale computational studies of membrane bilayer remodeling by curvature-inducing proteins. Phys. Rep. 543(1), 1 (2014). doi:10.1016/j.physrep.2014.05.001
Deserno, M.: Fluid lipid membranes: from differential geometry to curvature stresses. Chem. Phys. Lipids (2014). doi:10.1016/j.chemphyslip.2014.05.001. http://linkinghub.elsevier.com/retrieve/pii/S000930841400053X
Canham, P.B.: The minimum energy of bending as a possible explanation of the biconcave shape of the human red blood cell. J. Theor. Biol. 26(1), 61 (1970)
Helfrich, W.: Elastic properties of lipid bilayers: theory and possible experiments. Z. Naturforsch. C 28, 693 (1973). http://www.ncbi.nlm.nih.gov/pubmed/4273690
Diz-Muñoz, A., Fletcher, D.A., Weiner, O.D.: Use the force: membrane tension as an organizer of cell shape and motility. Trends Cell Biol. 23(2), 47 (2013). doi:10.1016/j.tcb.2012.09.006
Shi, Z., Baumgart, T.: Membrane tension and peripheral protein density mediate membrane shape transitions. Nat. Commun. 6, 5974 (2015). doi:10.1038/ncomms6974. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=25569184&retmode=ref&cmd=prlinks
Deserno, M.: Fluid lipid membranes: from differential geometry to curvature stresses. Chem. Phys. Lipids 185, 11 (2015). doi:10.1016/j.chemphyslip.2014.05.001. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=24835737&retmode=ref&cmd=prlinks
do Carmo, M.P.: Differential Geometry of Curves and Surfaces. Prentice Hall, New Jersey (1976)
Lipowsky, R.: Spontaneous tubulation of membranes and vesicles reveals membrane tension generated by spontaneous curvature. Faraday Discuss. 161, 305 (2013). doi:10.1039/c2fd20105d. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=2013FaDi.161.305L&link_type=EJOURNAL
Schnur, J.M.: Lipid tubules: a paradigm for molecularly engineered structures. Science 262(5140), 1669 (1993). doi:10.1126/science.262.5140.1669. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=1993Sci...262.1669S&link_type=ABSTRACT
Kohyama, T., Kroll, D.M., Gompper, G.: Budding of crystalline domains in fluid membranes. Phys. Rev. E 68(6), 061905 (2003). doi:10.1103/PhysRevE.68.061905
Sunil Kumar, P.B., Gompper, G., Lipowsky, R.: Modulated phases in multicomponent fluid membranes. Phys. Rev. E 60(4 Pt B), 4610 (1999). doi:10.1103/PhysRevE.60.4610. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=1999PhRvE.60.4610K&link_type=ABSTRACT
Nelson, D.R., Piran, T.: Statistical mechanics of membranes and surfaces. World Sci. (2004). http://books.google.com/books?id=FbcMqgNrVjcC&pg=PA323&dq=intitle:Statistical+mechanics+of+membranes+and+surfaces&hl=&cd=1&source=gbs_api
Ramakrishnan, N., Sunil Kumar, P.B., Ipsen, J.H.: Monte Carlo simulations of fluid vesicles with in-plane orientational ordering. Phys. Rev. E 81(4), 041922 (2010). doi:10.1103/PhysRevE.81.041922
Agrawal, N.J., Nukpezah, J., Radhakrishnan, R.: Minimal mesoscale model for protein-mediated vesiculation in Clathrin-dependent endocytosis. PLoS Comput. Biol. 6(9), e1000926 (2010). doi:10.1371/journal.pcbi.1000926.s008
Ramanan, V., Agrawal, N.J., Liu, J., Engles, S., Toy, R., Radhakrishnan, R.: Systems biology and physical biology of clathrin-mediated endocytosis. Integr. Biol. 3(8), 803 (2011). doi:10.1039/c1ib00036e. http://www.ncbi.nlm.nih.gov/pubmed/21792431
Liu, J., Tourdot, R.W., Ramanan, V., Agrawal, N.J., Radhakrishanan, R.: Mesoscale simulations of curvature-inducing protein partitioning on lipid bilayer membranes in the presence of mean curvature fields. Mol. Phys. 110(11–12), 1127 (2012). doi:10.1080/00268976.2012.664661
Tourdot, R.W., Ramakrishnan, N., Radhakrishnan R.: Defining the free-energy landscape of curvature-inducing proteins on membrane bilayers. Phys. Rev. E 90, 022717 (2014). http://journals.aps.org/pre/abstract/10.1103/PhysRevE.90.022717
Zhao, Y., Liu, J., Yang, C., Capraro, B.R., Baumgart, T., Bradley, R.P., Ramakrishnan, N., Xu, X., Radhakrishnan, R., Svitkina, T., Guo, W.: Exo70 generates membrane curvature for morphogenesis and cell migration. Dev. Cell 26(3), 266 (2013). doi:10.1016/j.devcel.2013.07.007. http://linkinghub.elsevier.com/retrieve/pii/S1534580713004152
Tourdot, R.W., Bradley, R.P., Ramakrishnan, N., Radhakrishnan, R.: Multiscale computational models in physical systems biology of intracellular trafficking. IET Syst. Biol. 8(5), 198 (2014). doi:10.1049/iet-syb.2013.0057. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=25257021&retmode=ref&cmd=prlinks
Ramakrishnan, N., Radhakrishnan, R.: Phenomenology based multiscale models as tools to understand cell mand organelle morphologies. In: Iglič, Aleš, Kulkarni, Chandrashekhar V, Rappolt, Michael (eds.), Academic Press, pp. 129–175 (2015) doi:10.1016/bs.adplan.2015.06.004. http://www.sciencedirect.com/science/article/pii/S1554451615000320
Metropolis, N., Rosenbluth, A.W., Rosenbluth, M.N., Teller, A.H., Teller, E.: Equation of state calculations by fast computing machines. J. Chem. Phys. 21(6), 1087 (1953). doi:10.1063/1.1699114. http://link.aip.org/link/?JCP/21/1087/1
Widom, B.: Some topics in the theory of fluids. J. Chem. Phys. 39(11), 2808 (1963). doi:10.1063/1.1734110. http://link.aip.org/link/JCPSA6/v39/i11/p2808/s1&Agg=doi
Bennett, C.H.: Efficient estimation of free-energy differences from Monte-Carlo data. J. Comput. Phys. 22(2), 245 (1976). doi:10.1016/0021-9991(76)90078-4. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=1976JCoPh.22.245B&link_type=ABSTRACT
Roux, B.: The calculation of the potential of mean force using computer simulations. Comput. Phys. Commun. 91(1), 275 (1995). http://www.sciencedirect.com/science/article/pii/001046559500053I
Ramakrishnan, N., Eckmann, D.M., Ayyaswamy, P.S., Weaver,Valerie M., Radhakrishnan, R.: Subcellular membrane mechanotyping using local estimates of cell membrane excess area (Unpublished data)
Souaille, M., Roux, B.: Extension to the weighted histogram analysis method: combining umbrella sampling with free energy calculations. Comput. Phys. Commun. 135(1), 40 (2001). doi:10.1016/S0010-4655(00)00215-0. http://adsabs.harvard.edu/cgi-bin/nph-data_query?bibcode=2001CoPhC.135...40S&link_type=ABSTRACT
Liu, J., Weller, G.E., Zern, B., Ayyaswamy, P.S., Eckmann, D.M., Muzykantov, V.R., Radhakrishnan, R.: Computational model for nanocarrier binding to endothelium validated using in vivo, in vitro, and atomic force microscopy experiments. Proc. Natl. Acad. Sci. USA 107(38), 16530 (2010). doi:10.1073/pnas.1006611107/-/DCSupplemental. http://www.pnas.org/content/107/38/16530.short
Liu, J., Agrawal, N.J., Calderon, A., Ayyaswamy, P.S., Eckmann, D.M., Radhakrishnan, R.: Multivalent binding of nanocarrier to endothelial cells under shear flow. Biophys. J. 101(2), 319 (2011). doi:10.1016/j.bpj.2011.05.063. http://linkinghub.elsevier.com/retrieve/pii/S0006349511006680
Acknowledgments
This work was supported in part by the US National Science Foundation Grants DMR-1120901, and CBET-1244507. The research leading to these results has received funding from the European Commission Grant FP7-ICT-2011-9-600841, US NIH U01-EB016027, and NIH 1U54CA193417. Computational resources were provided in part by the National Partnership for Advanced Computational Infrastructure under Grant No. MCB060006 from XSEDE.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ramakrishnan, N., Tourdot, R.W. & Radhakrishnan, R. Thermodynamic free energy methods to investigate shape transitions in bilayer membranes. Int J Adv Eng Sci Appl Math 8, 88–100 (2016). https://doi.org/10.1007/s12572-015-0159-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12572-015-0159-5