Abstract
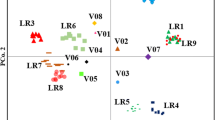
Genetic diversity was studied among 21 accessions of lentil using SSR markers and morphological traits in order to assess the diversification of Indian gene-pool of lentil through introgression of exotic genes and introduction of germplasm. Among these , 16 genotypes either had ‘Precoz’ gene, an Argentine line in their pedigree or genes from introduced lines from ICARDA. Sixty five SSR markers and eight phenotypic traits were used to analyse the level of genetic diversity in these genotypes. Forty three SSR markers (66 %) were polymorphic and generated a total of 177 alleles with an average of 4.1 alleles per SSR marker. Alleles per marker ranged from 2 to 6. The polymorphic information content ranged 0.33 to 0.80 with an average of 0.57, suggesting that SSR markers are highly polymorphic among the studied genotypes. Genetic dissimilarity based a dendrogram grouped these accessions into two main clusters (cluster I and cluster II) and it ranged 33 % to 71 %, suggesting high level of genetic diversity among the genotypes. First three components of PCA based morphological traits explained higher variance (95.6 %) compared to PCA components based on SSR markers (42.7 %) of total genetic variance. Thus, more diversity was observed for morphological traits and genotypes in each cluster and sub-cluster showed a range of variability for seed size, earliness, pods/plant and plant height. Molecular and phenotypic diversity analysis thus suggested that use of germplasm of exotic lines have diversified the genetic base of lentil germplasm in India. This diversified gene-pool will be very useful in the development of improved varieties of lentil in order to address the effect of climate change, to adapt in new cropping systems niches such as mixed cropping, relay cropping, etc. and to meet consumers’ preference.


Similar content being viewed by others
References
Abe J, Xu DH, Suzuki Y, Kanazawa A, Shimanmoto Y (2003) Soybean germplasm pools in Asia revealed by nuclear SSRs. Theor Appl Genet 106:445–453
Agrawal PK, Katiyar AK (2008) Validation of chickpea-STMS markers and DNA fingerprinting in lentil (Lens culinaris subsp. culinaris) cultivars of India. Indian J Genet 68:149–156
Andeden EE, Derya M, Baloch FS, Kilian B, Ozkan H (2013) Development of SSR Markers in Lentil. In: Plant and Animal Genome XXI. Jan 12-16 2013, San Diego, Canada (abstract)
Abdelnoor RV, Barros EG, Moreira (1995) Determination of genetic diversity within Brazilian soybean germplasm using random amplified polymorphic DNA techniques and comparative analysis with pedigree data. Braz J Genet 18:265–273
Asghar MJ, Abbas G, Shah TM (2010) Study of genetic diversity in some local and exotic lentil (Lens culinaris medik) genotypes. Pak J Bot 42:2681–2690
Botstein B, White RL, Skolnick M, Davis RW (1980) Molecular markers in plant genome analysis. Am J Hum Genet 32:314–331
Datta S, Tiwari S, Kaashyap M, Gupta PP, Choudhury PR, Kumari J, Kumar S (2011) Genetic similarity analysis in lentil using cross-genera legume sequence tagged microsatellite site markers. Crop Sci 51:2412–2422
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Erskine W (1997) Lessons for breeders from land races of lentil. Euphytica 93:107–112
Erskine W, Adham Y, Holly L (1989) Geographic distribution of variation in quantitative traits in a world of lentil collection. Euphytica 43:97–103
Erskine W, Chandra S, Chaudhary M, Malik IA, Sarker A, Sharma B, Tufail M, Tyagi MC (1998) A bottleneck in lentil: widening its genetic base in South Asia. Euphytica 101:207–211
Ferguson ME, Robertson LD, Ford-Lloyd BV, Newbury HJ, Maxted N (1998) Contrasting genetic variation amongst lentil landraces from different geographical origins. Euphytica 102:265–273
Gupta D, Sharma SK (2007) Widening the gene pool of cultivated lentils through introgression of alien chromatin from wild Lens subspecies. Plant Breed 126:58–61
Gupta M, Verma B, Kumar N, Chahota RK, Rathour R, Sharma SK, Bhatia S, Sharma TR (2012) Construction of intersubspecific molecular genetic map of lentil based on ISSR, RAPD and SSR markers. J Genet 91:279–287
Gupta PK, Balyan HS, Varshney RK, Gill KS (2013) Development and use of molecular markers for crop improvement. Plant Breed 132:431–432
Hamwieh A, Udupa SM, Choumane W, Sarker A, Dreyer F, Jung C, Baum M (2005) A genetic linkage map of Lens sp. based on microsatellite and AFLP markers and the localization of Fusarium vascular wilt resistance. Theor Appl Genet 77:839–843
Hamwieh A, Udupa SM, Sarkar A, Jung C, Baum M (2009) Development of new microsatellite markers and their application in the analysis of genetic diversity in lentils. Breed Sci 59:77–86
Jaccard P (1908) Novelle researchers sur la distribution florale. Bull Soc Vaud Sci Nat 44:270
Kaur S, Cogan NO, Pembleton LW, Shinozuka M, Savin KW, Materne M, Forster JW (2011) Transcriptome sequencing of lentil based on second-generation technology permits large-scale unigene assembly and SSR marker discovery. BMC Genomics 12:265–275
Kumar S, Gupta S, Chandra S, Singh BB (2004) How wide is the genetic base of pulse crops. In: Ali M, Singh BB, Kumar S, Dhar V (eds) Pulses in new perspective. ISPRD, Kanpur, pp 188–210
Ladizinski D, Braun D, Muehlbauer FJ (1984) The biological species of the genus Lens. Bot Gaz 145:235–261
Rahman MM, Sarker A, Kumar S, Ali A, Yadav NK, Rahman L (2009) Breeding for short season environments. In: Erskine W, Muehlbauer F, Sarker A, Sharma B (eds) The lentil: botany, production and uses. CAB International, Wallingford, pp 121–136
Reddy MRK, Rathour R, Kumar N, Kathoch P, Sharma TR (2009) Cross-genera legume SSR markers for analysis of genetic diversity in Lens species. Plant Breed 129:514–518
Rohlf FJ (1998) NTSYS-pc: numerical taxonomy and multivariate analysis system. Version 2.1. Exeter Publications, NY
Singh BB, Mishra SK, Sardana S, Dixit GP (2006) Lentil and pea. In: Dhillon BS, Saxena S, Agarwal A, Tyagi RK (eds) Plant genetic resources: food grain crops. Narosa Publishing House Pvt. Ltd., New Delhi, pp 240–254
Singh M, Rana MK, Kumar K, Bisht IS, Dutta M, Gautam NK, Sarker A, Bansal KC (2013) Broadening the genetic base of lentil cultivars through inter-sub-specific and interspecific crosses of Lens taxa. Plant Breed. doi:10.1111/pbr.12089
Udupa SM, Robertson LD, Weigand F, Baum M, Kahl G (1999) Allelic variation at (TAA)n microsatellite loci in a world collection of chickpea (Cicer arietinum L.) germplasm. Mol Gen Genet 261:354–363
Varshney RK, Close TJ, Singh NK, Hoisington DA, Cook DR (2009) Orphan legume crops enter the genomics era! Curr Opin Plant Biol 11:1–9
Varshney RK, Graner A, Sorrells ME (2005) Genic microsatellite markers in plants: features and applications. Trends Biotechnol 23:48–55
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kumar, J., Srivastva, E., Singh, M. et al. Diversification of indigenous gene- pool by using exotic germplasm in lentil (Lens culinaris Medikus subsp. culinaris). Physiol Mol Biol Plants 20, 125–132 (2014). https://doi.org/10.1007/s12298-013-0214-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-013-0214-2