Abstract
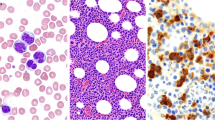
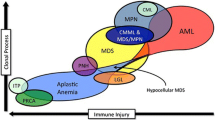
The MDS/MPN overlap syndromes are recently evolved entities that have been quite difficult to define since their discovery. They have overlapping features with other myeloid neoplasms such as MDS and MPN, which further complicates the task of their diagnosis. The unravelling of their molecular pathogenesis by recent diagnostic innovations was of paramount significance in understanding the mechanism of these syndromes. The identification of the major genetic pathways implicated in their pathogenesis not only will help in their diagnosis, but also will enable development of targeted molecular therapy as well as prognostic markers. This review discus the basic molecular aberrations in MDS/MPN overlap syndromes and their possible future implications.


Similar content being viewed by others
References
Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Thiele J (2017) WHO classification of tumours of haematopoietic and lymphoid tissue, Revised 4th edn. IARC, Lyon, p p10
Tiu RV, Gondek LP, O’Keefe CL et al (2011) Prognostic impact of SNP array karyotypingin myelodysplastic syndromes and related myeloid malignancies. Blood 117(17):4552–4560
Cazzola M, Malcovati L, Invernizzi R (2011) Myelodysplastic/myeloproliferative neoplasms. Hematology Am Soc Hematol Educ Program 2011:264–272
Delhommeau F, Pisani DF, James C et al (2006) Oncogenic mechanisms in myeloproliferative disorders. Cell Mol Life Sci 63(24):2939–2953
James C, Ugo V, Le Couédic JP et al (2005) A unique clonal JAK2 mutation leading to constitutive signalling causes polycthemia vera. Nature 434(7037):1144–1148
Cools J, DeAngelo DJ, Gotlib J et al (2003) A tyrosine kinase created by fusion of the PDGFRA and FIP1L1 genes is a therapeutic target of imatinib in idiopathic hypereosinophilic syndrome. N Engl J Med 348(13):1201–1214
Chase A, Bryant C, Score J, Cross NC (2003) Ponatinib as targeted therapy for FGR1 fusions associated with the 8p11 myeloproliferative syndrome. Haematologica 98(1):103–106
Lierman E, Selleslag D, Smits S, Bilet J, Vandenberghe P (2012) Ruxolitinib inhibits transforming JAK2 fusion proteins in vitro and induces complete remission in t(8;9)(p22;p24)/PCM1-JAK2-positive chronic eosinophilic leukemia. Blood 120(7):1529–1531
Chase A, Bryant C, Score J et al (2013) Ruxolitinib as potential targeted therapy for patients with JAK2 rearrangements. Haematologica 98(3):404–408
Haferlach T, Nagata Y, Grossmann V, Okuno Y, Bacher U, Nagae G et al (2014) Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia 28:241–247
Kohlmann A, Grossmann V, Nadarajah N, Haferlach T (2013) Next generation sequencing—feasibility and practicality in haematology. Br J Haematol 160:736–753
Wang J, Liu Y, Li Z et al (2010) Endogenous oncogenic NRAS mutation promotes aberrant GM-CSF signaling in granulocytic/monocytic precursors in a murine model of chronic myelomonocyticleukemia. Blood 116(26):5991–6002
Itzykson R, Kosmider O, Renneville A, Gelsi-Boyer V, Meggendorfer M, Morabito M et al (2013) Prognostic score including gene mutations in chronic myelomonocytic leukemia. J Clin Oncol 31:2428–2436
Padron E, Painter JS, Kunigal S, Mailloux AW, McGraw K, McDaniel JM et al (2013) GM-CSF-dependent pSTAT5 sensitivity is a feature with therapeutic potential in chronic myelomonocyticleukemia. Blood 121:5068–5077
Braun BS, Tuveson DA, Kong N, Le DT, Kogan SC, Rozmus J et al (2004) Somatic activation of oncogenic Kras in hematopoietic cells initiates a rapidly fatal myeloproliferative disorder. Proc Natl Acad Sci USA 101:597–602
Chan IT, Kutok JL, Williams IR, Cohen S, Kelly L, Shigematsu H et al (2004) Conditional expression of oncogenic K-RAS from its endogenous promoter induces a myeloproliferative disease. J Clin Invest 113:528–538
Van Meter ME, Diaz-Flores E, Archard JA, Passegue E, Irish JM, Kotecha N et al (2007) K-RASG12D expression induces hyperproliferation and aberrant signaling in primary hematopoietic stem/progenitor cells. Blood 109:3945–3952
Cross NC (2011) Genetic and epigenetic complexity in myeloproliferative neoplasms. Hematology Am Soc Hematol Educ Program 2011:208–214
Oh ST, Gotlib J (2010) JAK2 V617F and beyond: role of genetics and aberrant signaling in the pathogenesis of myeloproliferative neoplasms. Expert Rev Hematol 3:323–337
Szpurka H, Gondek LP, Mohan SR, Hsi ED, Theil KS, Maciejewski JP (2009) UPD1p indicates the presence of MPL W515L mutation in RARS-T, a mechanism analogous to UPD9p and JAK2 V617F mutation. Leukemia 23:610–614
Grand FH, Hidalgo-Curtis CE, Ernst T, Zoi K, Zoi C, McGuire C et al (2009) Frequent CBL mutations associated with 11q acquired uniparental disomy in myeloproliferative neoplasms. Blood 113:6182–6192
Itzykson R, Kosmider O, Renneville A, Morabito M, Preudhomme C, Berthon C et al (2013) Clonal architecture of chronic myelomonocyticleukemias. Blood 121:2186–2198
Valent P, Arock M, Akin C, Sperr WR, Reiter A, Sotlar K et al (2010) The classification of systemic mastocytosis should include mast cell leukemia (MCL) and systemic mastocytosis with a clonal hematologic non-mast cell lineage disease (SM-AHNMD). Blood 116:850–851
Machherndl-Spandl S, Sega W, Bosmuller H, Germing U, Gruber C, Nachtkamp K et al (2014) Prognostic impact of blast cell counts in dysplastic bone marrow disorders (MDS and CMML I) with concomitant fibrosis. Ann Hematol 93:57–64
Sotlar K, Marafioti T, Griesser H, Theil J, Aepinus C, Jaussi R et al (2000) Detection of c-kit mutation Asp 816 to Val in microdissected bone marrow infiltrates in a case of systemic mastocytosis associated with chronic myelomonocytic leukaemia. Mol Pathol 53:188–193
Grand FH, Iqbal S, Zhang L, Russell NH, Chase A, Cross NC (2007) A constitutively active SPTBN1-FLT3 fusion in atypical chronic myeloid leukemia is sensitive to tyrosine kinase inhibitors and immunotherapy. Exp Hematol 35:1723–1727
Walz C, Erben P, Ritter M, Bloor A, Metzgeroth G, Telford N et al (2011) Response of ETV6-FLT3-positive myeloid/lymphoid neoplasm with eosinophilia to inhibitors of FMS-like tyrosine kinase 3. Blood 118:2239–2242
Wang SA, Hasserjian RP, Fox PS, Rogers HJ, Geyer JT, Chabot- Richards D et al (2014) Atypical chronic myeloid leukemia is clinically distinct from unclassifiable myelodysplastic/myeloproliferative neoplasms. Blood 123:2645–2651
Pardanani A, Lasho TL, Laborde RR, Elliott M, Hanson CA, Knudson RA et al (2013) CSF3R T618I is a highly prevalent and specific mutation in chronic neutrophilic leukemia. Leukemia 27:1870–1873
Maxson JE, Gotlib J, Pollyea DA, Fleischman AG, Agarwal A, Eide CA et al (2013) Oncogenic CSF3R mutations in chronic neutrophilic leukemia and atypical CML. N Engl J Med 368:1781–1790
Piazza R, Valletta S, Winkelmann N, Redaelli S, Spinelli R, Pirola A et al (2013) Recurrent SETBP1 mutations in atypical chronic myeloid leukemia. Nat Genet 45:18–24
Minakuchi M, Kakazu N, Gorrin-Rivas MJ, Abe T, Copeland TD, Ueda K, Adachi Y (2001) Identification and characterization of SEB, a novel protein that binds to the acute undifferentiated leukemia-associated protein SET. Eur J Biochem 268:1340–1351
Chen HC, Cheng SC (2012) Functional roles of protein splicing factors. Biosci Rep 32:345–359
Visconte V, Avishai N, Mahfouz R, Tabarroki A, Cowen J, Sharghi-Moshtaghin R et al (2014) Distinct iron architecture in SF3B1-mutant myelodysplastic syndrome patients is linked to an SLC25A37 splice variant with a retained intron. Leukemia 29(1):188–195
Ernst T, Chase A, Zoi K, Waghorn K, Hidalgo-Curtis C, Score J et al (2010) Transcription factor mutations in myelodysplastic/myeloproliferative neoplasms. Haematologica 95:1473–1480
Kon A, Shih LY, Minamino M, Sanada M, Shiraishi Y, Nagata Y et al (2013) Recurrent mutations in multiple components of the cohesin complex in myeloid neoplasms. Nat Genet 45:1232–1237
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A et al (2010) Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18:553–567
Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV et al (2010) Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet 42:722–726
Abdel-Wahab O, Adli M, LaFaveLM Gao J, Hricik T, Shih AH et al (2012) ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 22:180–193
Patnaik MM, Padron E, Laborde RR et al (2013) Mayo prognostic model for WHO-Defined chronic myelomonocytic leukemia-ASXL1 and spliceosome component mutations and outcomes. Leukemia 27(7):1504–1510
Niemeyer CM, Kratz CP (2008) Paediatric myelodysplastic syndromes and juvenile myelomonocytic leukaemia: molecular classification and treatment options. Br J Haematol 140(6):610–624
Loh ML (2011) Recent advances in the pathogenesis and treatment of juvenile myelomonocytic leukaemia. Br J Haematol 152(6):677–687
Shannon KM, O’Connell P, Martin GA et al (1994) Loss of the normal NF1 allele from the bone marrow of children with type 1 neurofibromatosis and malignant myeloid disorders. N Engl J Med 330(9):597–601
Flotho C, Valcamonica S, Mach-Pascual S, Schmahl G, Corral L, Ritterbach J et al (1999) RAS mutations and clonality analysis in children with juvenile myelomonocytic leukemia (JMML). Leukemia 13(1):32–37
Tartaglia M, Niemeyer CM, Fragale A et al (2003) Somatic mutations in PTPN11 in juvenile myelomonocytic leukemia, myelodysplastic syndromes and acute myeloid leukemia. Nat Genet 34(2):148–150
Loh ML, Vattikuti S, Schubbert S et al (2004) Mutations in PTPN11 implicate the SHP-2 phosphatase in leukemogenesis. Blood 103(6):2325–2331
Pérez B, Mechinaud F, Galambrun C et al (2010) Germline mutations of the CBL gene define a new genetic syndrome with predisposition to juvenile myelomonocytic leukaemia. J Med Genet 47(10):686–691
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pati, H., Kundil Veetil, K. Myelodysplastic Syndrome/Myeloproliferative Neoplasm (MDS/MPN) Overlap Syndromes: Molecular Pathogenetic Mechanisms and Their Implications. Indian J Hematol Blood Transfus 35, 3–11 (2019). https://doi.org/10.1007/s12288-019-01084-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12288-019-01084-y