Abstract
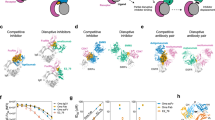
The engineering of affinity reagents has become a standard technology in modern drug development efforts. High-throughput screening of recombinant protein libraries has yielded numerous affinity reagents that are used in diagnostic or therapeutic applications. In our approach, we engineer intracellular affinity reagents by enhancing pre-existing intermolecular contacts to target functional epitopes in proteins. The designed affinity reagents allow a fast and specific interrogation of druggable sites in therapeutic relevant proteins.
Similar content being viewed by others
Literatur
Cohen P, Tcherpakov M (2010) Will the ubiquitin system furnish as many drug targets as protein kinases? Cell 143:686–693
Rolland T, Tasan M, Charloteaux B, et al. (2014) A proteome-scale map of the human interactome network. Cell 159:1212–1226
Argos P (1988) An investigation of protein subunit and domain interfaces. Protein Eng 2:101–113
Vajda S, Guarnieri F (2006) Characterization of protein-ligand interaction sites using experimental and computational methods. Curr Opin Drug Discov Devel 9:354–362
Tomlinson IM (2004) Next-generation protein drugs. Nat Biotechnol 22:521–522
Koide A, Abbatiello S, Rothgery L, et al. (2002) Probing protein conformational changes in living cells by using designer binding proteins: application to the estrogen receptor. Proc Natl Acad Sci USA 99:1253–1258
Binz HK, Amstutz P, Kohl A, et al. (2004) High-affinity binders selected from designed ankyrin repeat protein libraries. Nat Biotechnol 22:575–582
Ernst A, Avvakumov G, Tong J, et al. (2013) A strategy for modulation of enzymes in the ubiquitin system. Science 339:590–595
Liu F, Walters KJ (2010) Multitasking with ubiquitin through multivalent interactions. Trends Biochem Sci 35:352–360
Zhang D, Raasi S, Fushman D (2008) Affinity makes the difference: nonselective interaction of the UBA domain of Ubiquilin-1 with monomeric ubiquitin and polyubiquitin chains. J Mol Biol 377:162–180
Wiechmann S, Gartner A, Kniss A, et al. (2017) Site-specific inhibition of the SUMO-conjugating enzyme Ubc9 selectively impairs SUMO chain formation. J Biol Chem, DOI:10.1074/ibc.M117.794255
Author information
Authors and Affiliations
Corresponding author
Additional information
Svenja Wiechmann 2007–2013 Biologiestudium an der TU Darmstadt. Seit 2013 Doktorandin im Institut für Biochemie II, Goethe Universität Frankfurt a. M.
Andreas Ernst 1993–2000 Chemiestudium an der TU Darmstadt. 2000–2006 Promotion an der Universität Zürich, Schweiz, im Labor von Prof. Dr. Andreas Plückthun. 2007–2008 Postdoc in San Francisco, CA, USA, im Department Proteinengineering bei Genentech in der Arbeitsgruppe von Dr. S. Sidhu. 2008 Umzug mit Dr. S. Sidhu an die Universität Toronto, Kanada, an das Centre for Cellular and Biomolecular Research. Seit 2013 Gruppenleiter im Institut für Biochemie II, Goethe Universität Frankfurt a. M. und seit 2015 in Nebentätigkeit als Gruppenleiter am Fraunhofer-Institut für Molekular biologie und Angewandte Ökologie, Projektgruppe Translationale Medizin und Pharmakologie, Frankfurt a. M.
Rights and permissions
About this article
Cite this article
Wiechmann, S., Ernst, A. Engineering von intrazellulären Modulatoren. Biospektrum 23, 769–771 (2017). https://doi.org/10.1007/s12268-017-0870-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12268-017-0870-9