Abstract
Purpose
Pharmaceutical technology transfer is one of the components of pharmaceutical innovation. Currently, a gap exists in pharmaceutical technology transfer from academia to industry. This study aims to develop an objective model to identify valuable pharmaceutical technologies for transferring in order to drive pharmaceutical innovation.
Methods
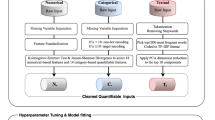
We created a support vector machine classifier model using the data of pharmaceutical patents held by universities to predict the licensing outcomes of those patents. We collected data on 369 United States (US) pharmaceutical patents, using 142 licensed patents as the positive samples and 227 unlicensed patents as the negative samples. We also collected the licensing data of the patents, and the distinguished patent features were selected for model training and generation. Upon optimization, the machine learning model was evaluated using different scoring methods.
Results
Our support vector machine-based model achieved a fairly good performance of 82.50% in precision and 88.89% in specificity.
Conclusions
To the best of our knowledge, our study is the first to apply the machine learning approach to predict the licensing outcomes for pharmaceutical patent valuation and technology transfer. Our work is a good alternative to the current patent valuation methods available in the market, and it could be further developed for practical use in real business contexts.


Similar content being viewed by others
References
Ni J, Shao R, Ung COL, Wang Y, Hu Y, Cai Y. Valuation of pharmaceutical patents: a comprehensive analytical framework based on technological, commercial, and legal factors. J Pharm Innov. 2015;10(3):281–5.
Butler D. Translational research: crossing the valley of death. Nature News. 2008;453(7197):840–2.
Henderson R, Jaffe AB, Trajtenberg M. Universities as a source of commercial technology: a detailed analysis of university patenting, 1965–1988. Rev Econ Stat. 1988;80(1):119–27.
Fernandez JM, Stein RM, Lo AW. Commercializing biomedical research through securitization techniques. Nat Biotechnol. 2012;30:964–75.
Ruckman K, McCarthy I. Why do some patents get licensed while others do not? Ind Corp Change. 2017;26(4):667–88.
Vapnik VN. An overview of statistical learning theory. IEEE Trans Neural Netw. 1999;10(5):988–99.
Zhang Y, Yang Y, Zhang H, Jiang X, Xu B, Xue Y, et al. Prediction of novel pre-microRNAs with high accuracy through boosting and SVM. Bioinformatics. 2011;27(10):1436–7.
Walt SVD, Colbert SC, Varoquaux G. The NumPy array: a structure for efficient numerical computation. Comput Sci Eng. 2011;13(2):22–30.
Oliphant T, SciPy TE. Open source scientific tools for Python. Comput Sci Eng. 2007;9:10–20.
Racine J. Gnuplot 4.0: a portable interactive plotting utility. J Appl Econ. 2006;21(1):133–41.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: machine learning in Python. J Mach Learn Res. 2011;12:2825–30.
Chang CC, Lin CJ. LIBSVM: a library for support vector machines. ACM Trans Intell Syst Technol. 2011;2(3):27.
Cervantes M. Academic patenting: how universities and public research organizations are using their intellectual property to boost research and spur innovative start-ups. WIPO Small And Medium-sized Enterprises Documents 2003. http://www.wipo.int/sme/en/documents/academic_patenting.html. Accessed 1 April 2018.
Salazar A, Hackney R, Howells J. The strategic impact of internet technology in biotechnology and pharmaceutical firms: insights from a knowledge management perspective. Info Technol Manag. 2003;4(2):289–301.
Laursen K, Leone MI, Torrisi S. Technological exploration through licensing: new insights from the licensee’s point of view. Ind Corp Change. 2010;19(3):871–97.
Leone MI, Reichstein T. Licensing-in fosters rapid invention! The effect of the grant-back clause and technological unfamiliarity. Strat Mgmt J. 2012;33(8):965–85.
Neuhäusler P, Frietsch R. Patent families as macro level patent value indicators: applying weights to account for market differences. Scientometrics. 2013;96(1):27–49.
Smith DKW. A new methodology for citation dependent patent evaluations. Carleton University. 2014. (Electronic, M.Sc. thesis) http://curve.carleton.ca/system/files/theses/31557.pdf. Accessed 1 Oct 2017.
Sakakibara M. An empirical analysis of pricing in patent licensing contracts. Ind Corp Chang. 2010;19(3):927–45.
Pai PF, Hong WC, Change PT, Chen CT. The application of support vector machines to forecast tourist arrivals in Barbados: an empirical study. Int J Manag. 2006;23(2):375.
Tseng CY, Chen MS. Incremental SVM model for spam detection on dynamic email social networks. IEEE CSE Int Conf. 2009;4:128–35.
Hua S, Sun Z. Support vector machine approach for protein subcellular localization prediction. Bioinformatics. 2001;17(8):721–8.
Yu CS, Lin CJ, Hwang JK. Predicting subcellular localization of proteins for Gram-negative bacteria by support vector machines based on n-peptide compositions. Protein Sci. 2004;13(5):1402–6.
Powers DM. Evaluation: from precision, recall and F-measure to ROC, 2011 informedness, markedness and correlation. J Mach Learn Tech. 2011;2(1):37–63.
Campos TC. The idea of patents vs. the idea of university. New Bioethnol. 2015;21(2):164–76.
Funding
This study was funded by the grant MYRG2015-00145-ICMS-QRCM from the University of Macau.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Research Involving Human Participants or Animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Not applicable.
Rights and permissions
About this article
Cite this article
Lin, HH., Ouyang, D. & Hu, Y. Intelligent Classifier: a Tool to Impel Drug Technology Transfer from Academia to Industry. J Pharm Innov 14, 28–34 (2019). https://doi.org/10.1007/s12247-018-9332-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12247-018-9332-2