Abstract
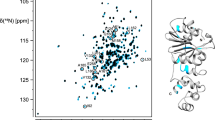
Ribonuclase P (RNase P) is an essential metallo-endonuclease that catalyzes 5′ precursor-tRNA (ptRNA) processing and exists as an RNA-based enzyme in bacteria, archaea, and eukaryotes. In bacteria, a large catalytic RNA and a small protein component assemble to recognize and accurately cleave ptRNA and tRNA-like molecular scaffolds. Substrate recognition of ptRNA by bacterial RNase P requires RNA–RNA shape complementarity, intermolecular base pairing, and a dynamic protein–ptRNA binding interface. To gain insight into the binding specificity and dynamics of the bacterial protein–ptRNA interface, we report the backbone and side chain 1H, 13C, and 15N resonance assignments of the hyperthermophilic Thermatoga maritima RNase P protein in solution at 318 K. Our data confirm the formation of a stable RNA recognition motif (RRM) with intrinsic heterogeneity at both the N- and C-terminus of the protein, consistent with available structural information. Comprehensive resonance assignments of the bacterial RNase P protein serve as an important first step in understanding how coupled RNA binding and protein–RNA conformational changes give rise to ribonucleoprotein function.


Similar content being viewed by others
References
Buck AH, Dalby AB, Poole AW, Kazantsev AV, Pace NR (2005a) Protein activation of a ribozyme: the role of bacterial RNase P protein. EMBO J 24:3360–3368
Buck AH, Kazantsev AV, Dalby AB, Pace NR (2005b) Structural perspective on the activation of RNase P RNA by protein. Nat Struct Mol Biol 12(11):958
Christian EL, Smith KM, Perera N, Harris ME (2006) The P4 metal binding site in RNase P RNA affects active site metal affinity through substrate positioning. RNA 12:1463–1467
Crary SM, Niranjanakumari S, Fierke CA (1998) The protein component of Bacillus subtilis Ribonuclease P increases catalytic efficiency by enhancing interactions with the 5′ leader sequence of pre-tRNAAsp. Biochemistry 37:9409–9416
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Goddard TD, Kneller DG. SPARKY 3. University of California, San Francisco. http://www.cgl.ucsf.edu/home/sparky/
Gopalan V, Jarrous N, Krasilnikov AS (2017) Chance and necessity in the evolution of RNase P. RNA 24(1):1–5 https://doi.org/10.1261/rna.063107.117
Guenther UP, Yandek LE, Niland CN, Campbell FE, Anderson D, Anderson VE, Harris ME, Jankowsky E (2013) Hidden specificity in an apparently nonspecific RNA-binding protein. Nature 502:385–388
Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S (1983) The RNA moiety of Ribonuclease P is the catalytic subunit of the enzyme. Cell 35:849–857
Hartmann RK, Gossringer M, Spath B, Fischer S, Marchfelder A (2009) The making of tRNAs and more—RNase P and tRNase Z. Prog Mol Biol Transl Sci 85:319–368
Hsieh J, Fierke CA (2009) Conformational change in the Bacillus subtilis RNase P holoenzyme–pre-tRNA complex enhances substrate affinity and limits cleavage rate. RNA 15:1565–1577
Kazantsev AV, Pace NR (2006) Bacterial RNase P: a new view of an ancient enzyme. Nat Rev Microbiol 4:729–740
Kazantsev AV, Krivenko AA, Harrington DJ, Holbrook SR, Adams PD, Pace NR (2005) Crystal structure of a bacterial Ribonuclease P RNA. Proc Natl Acad Sci USA 102:13392–13397
Keller RLJ (2004) The Computer Aided Resonance Assignment Tutorial. Cantina, The Swiss Federal Institute of Technology Zürich, Zürich
Klemm BP, Wu N, Chen Y, Liu X, Kaitany KJ, Howard MJ, Fierke CA (2016) The diversity of Ribonuclease P: protein and RNA catalysts with analogous biological functions. Biomolecules 6:27
Koutmou KS, Zahler NH, Kurz JC, Campbell FE, Harris ME, Fierke CA (2010) Protein-precursor tRNA contact leads to sequence-specific recognition of 5′ leaders by bacterial Ribonuclease P. J Mol Biol 396:195–208
Krasilnikov AS, Yang X, Pan T, Mondragón A (2003) Crystal structure of the specificity domain of Ribonuclease P. Nature 421:760–764
Krasilnikov AS, Xiao Y, Pan T, Mondragon A (2004) Basis for structural diversity in homologous RNAs. Science 306:104–107
Krivenko AA, Kazantsev AV, Adamidi C, Harrington DJ, Pace NR (2002) Expression, purification, crystallization and preliminary diffraction analysis of RNase P protein from Thermotoga maritima. Acta Crystallogr Sect D 58:1234–1236
Kurz JC, Niranjanakumari S, Fierke CA (1998) Protein component of Bacillus subtilis RNase P specifically enhances the affinity for precursor-tRNAAsp. Biochemistry 37:2393–2400
Lemieux B, Laterreur N, Perederina A, Noel JF, Dubois ML, Krasilnikov AS, Wellinger RJ (2016) Active Yeast Telomerase shares subunits with Ribonucleoproteins RNase P and RNase MRP. Cell 165:1171–1181
Lin HC, Zhao J, Niland CN, Tran B, Jankowsky E, Harris ME (2016) Analysis of the RNA binding specificity landscape of C5 protein reveals structure and sequence preferences that direct RNase P specificity. Cell Chem Biol 23:1271–1281
Liu F, Altman S (2010) Ribonuclease P. Springer, New York
Liu X, Chen Y, Fierke CA (2017) Inner-sphere coordination of divalent metal ion with nucleobase in catalytic RNA. J Am Chem Soc. https://doi.org/10.1021/jacs.7b08755
Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wuthrich K (1998) Recommendations for the presentation of NMR structures of proteins and nucleic acids. IUPAC-IUBMB-IUPAB Inter-Union Task Group on the Standardization of Data Bases of Protein and Nucleic Acid Structures Determined by NMR Spectroscopy. J Biomol NMR 12:1–23
Martin WJ, Reiter NJ (2017) Structural roles of noncoding RNAs in the heart of enzymatic complexes. Biochemistry 56:3–13
Mondragon A (2013) Structural studies of RNase P. Ann Rev Biophys 42:537–557
Niland CN, Zhao J, Lin HC, Anderson DR, Jankowsky E, Harris ME (2016) Determination of the specificity landscape for Ribonuclease P processing of precursor tRNA 5′ leader sequences. ACS Chem Biol 11:2285–2292
Niland CN, Anderson DR, Jankowsky E, Harris ME (2017) The contribution of the C5 protein subunit of Escherichia coli Ribonuclease P to specificity for precursor tRNA is modulated by proximal 5′ leader sequences. RNA 23:1502–1511
Niranjanakumari S, Stams T, Crary SM, Christianson DW, Fierke CA (1998) Protein component of the ribozyme Ribonuclease P alters substrate recognition by directly contacting precursor tRNA. Proc Natl Acad Sci USA 95:15212–15217
Paul R, Lazarev D, Altman S (2001) Characterization of RNase P from Thermotoga maritima. Nucleic Acids Res 29:880–885
Reiter NJ, Osterman A, Torres-Larios A, Swinger KK, Pan T, Mondragon A (2010) Structure of a bacterial Ribonuclease P holoenzyme in complex with tRNA. Nature 468:784–789
Reiter NJ, Osterman AK, Mondragon A (2012) The bacterial Ribonuclease P holoenzyme requires specific, conserved residues for efficient catalysis and substrate positioning. Nucleic Acids Res 40(20):10384–10393
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241
Stams T, Niranjanakumari S, Fierke CA, Christianson DW (1998) Ribonuclease P protein structure: evolutionary origins in the translational apparatus. Science 280:752–755
Sun L, Campbell FE, Zahler NH, Harris ME (2006) Evidence that substrate-specific effects of C5 protein lead to uniformity in binding and catalysis by RNase P. EMBO J 25:3998–4007
Sun L, Campbell FE, Yandek LE, Harris ME (2010) Binding of C5 protein to P RNA enhances the rate constant for catalysis for P RNA processing of Pre-tRNAs lacking a consensus G(+ 1)/C(+ 72) pair. J Mol Biol 395:1019–1037
Torres-Larios A, Swinger KK, Krasilnikov AS, Pan T, Mondragon A (2005) Crystal structure of the RNA component of bacterial Ribonuclease P. Nature 437:584–587
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J Biomol NMR 6:135–140
Acknowledgements
This work was supported in part by grants for NMR instrumentation from the NSF (0922862), NIH (S10 RR025677) and Vanderbilt University. BPB acknowledges support from the NIGMS (T32GM007347). NJR acknowledges support from Vanderbilt University and Marquette University, as well as funding from the American Heart Association (14GRNT20380334) and the NIH (GM120572). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zeng, D., Brown, B.P., Voehler, M.W. et al. NMR resonance assignments of RNase P protein from Thermotoga maritima. Biomol NMR Assign 12, 183–187 (2018). https://doi.org/10.1007/s12104-018-9806-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-018-9806-7