Abstract
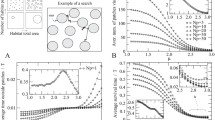
Animal movement, whether for foraging, mate-seeking, predator avoidance, dispersal, or migration, is a fundamental aspect of ecology that shapes spatial abundance distributions, genetic compositions, and dynamics of populations. A variety of movement models have been used for predicting the effects of natural or human-caused landscape changes, invading species, or other disturbances on local ecology. Here we introduce the flow network—a general modeling framework for population dynamics and movement in a metapopulation representing a network of habitat sites (nodes). Based on the principles of physical transport phenomena such as fluid flow through pipes (Pouiselle’s Law) and analogously, the flow of electric current across a circuit (Ohm’s Law), the flow network provides a novel way of modeling movement, where flow rates are functions of relative node pressures and the resistance to movement between them. Flow networks offer the flexibility of incorporating abiotic and biotic conditions that affect either pressures, resistance, or both. To illustrate an application of the flow network, we present a theoretical invasion scenario. We consider the effects of spatial structure on the speed of invasion by varying the spatial regularity of node arrangement. In the context of invasion, we model management actions targeting nodes or edges, and consider the effects on speed of invasion, node occupation, and total abundance. The flow network approach offers the flexibility to incorporate spatial heterogeneity in both rates of flow and site pressures and offers an intuitive approach to connecting population dynamics and landscape features to model movement.




Similar content being viewed by others
References
Adriaensen F, Chardon J, De Blust G, Swinnen E, Villalba S, Gulinck H, Matthysen E (2003) The application of least-cost modelling as a functional landscape model. Landscape Urban Plan 64(4):233–247
Amarasekare P (2004) The role of density-dependent dispersal in source–sink dynamics. J Theor Biol 226(2):159–168
Amos JN, Bennett AF, Mac Nally R, Newell G, Pavlova A, Radford JQ, Thomson JR, White M, Sunnucks P (2012) Predicting landscape-genetic consequences of habitat loss, fragmentation and mobility for multiple species of woodland birds. PLoS One 7(2):e30,888
Armsworth PR (2002) Recruitment limitation, population regulation, and larval connectivity in reef fish metapopulations. Ecology 83(4):1092–1104
Barrios JM, Verstraeten WW, Maes P, Aerts JM, Farifteh J, Coppin P (2012) Using the gravity model to estimate the spatial spread of vector-borne diseases. Int J Environ Res Publ Health 9(12):4346–4364
Bascompte J, Solé RV (1996) Habitat fragmentation and extinction thresholds in spatially explicit models. J Anim Ecol 65(4):465–473
Bossenbroek JM, Kraft CE, Nekola JC (2001) Prediction of long-distance dispersal using gravity models: zebra mussel invasion of inland lakes. Ecol Appl 11(6):1778–1788
Carroll C, Miquelle DG (2006) Spatial viability analysis of amur tiger panthera tigris altaica in the russian far east: the role of protected areas and landscape matrix in population persistencerstudio. J Appl Ecol 43(6):1056–1068. https://doi.org/10.1111/j.1365-2664.2006.01237.x
Dingle H (2014) Migration: the biology of life on the move. Oxford University Press, USA
Dingle H, Drake VA (2007) What is migration? Bioscience 57(2):113–121
Elliot NB, Cushman SA, Macdonald DW, Loveridge AJ (2014) The devil is in the dispersers: predictions of landscape connectivity change with demography. J Appl Ecol 51(5):1169–1178
Facon B, David P (2006) Metapopulation dynamics and biological invasions: a spatially explicit model applied to a freshwater snail. Am Nat 168:769–783. https://doi.org/10.1086/508669
Geritz SA, Gyllenberg M, Ondráċek P (2009) Evolution of density-dependent dispersal in a structured metapopulation. Math Biosci 219(2):142–148
Gilarranz LJ, Bascompte J (2012) Spatial network structure and metapopulation persistence. J Theor Biol 297:11–16
Grilli J, Barabás G, Allesina S (2015) Metapopulation persistence in random fragmented landscapes. PLoS Comput Biol 11(5): e1004,251
Hanski I (1999) Metapopulation ecology. Oxford University Press
Hanski I, Ovaskainen O (2000) The metapopulation capacity of a fragmented landscape. Nature 404 (6779):755–758
Hill M, Caswell H (1999) Habitat fragmentation and extinction thresholds on fractal landscapes. Ecol Lett 2(2):121–127
Lande R (1987) Extinction thresholds in demographic models of territorial populations. Am Nat 130(4):624–635. https://doi.org/10.1086/284734
Leung B, Drake JM, Lodge DM (2004) Predicting invasions: propagule pressure and the gravity of allee effects. Ecology 85(6):1651–1660. http://www.esajournals.org/doi/abs/10.1890/02-0571
Marrotte R, Gonzalez A, Millien V (2014) Landscape resistance and habitat combine to provide an optimal model of genetic structure and connectivity at the range margin of a small mammal. Molecul Ecol http://onlinelibrary.wiley.com/doi/10.1111/mec.12847/abstract
Marrotte RR, Bowman J (2017) The relationship between least-cost and resistance distance. PloS one 12(3):e0174,212
Mateo-Sánchez MC, Balkenhol N, Cushman S, Pérez T, Domnguez A, Saura S (2015) Estimating effective landscape distances and movement corridors: comparison of habitat and genetic data. Ecosphere 6(4):1–16. https://doi.org/10.1890/ES14-00387.1. art59
Matthysen E (2012) Multicausality of dispersal: a review. In: Clobert J, Baguette M, Benton TG, Bullock JM (eds). Dispersal Ecology and Evolution. Oxford University Press. pp 3–18
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 89(10):2712–2724. http://www.esajournals.org/doi/abs/10.1890/07-1861.1
Morales JM, Moorcroft PR, Matthiopoulos J, Frair JL, Kie JG, Powell RA, Merrill EH, Haydon DT (2010) Building the bridge between animal movement and population dynamics. Philos Trans R Soc B: Biol Sci 365(1550):2289–2301
North AR, Godfray HCJ (2017) The dynamics of disease in a metapopulation: the role of dispersal range. J Theor Biol 418:57–65
Parvinen K (2002) Evolutionary branching of dispersal strategies in structured metapopulations. J Math Biol 45(2):106–124
Peacock SJ, Bateman AW, Krkoṡek M, Lewis MA (2016) The dynamics of coupled populations subject to control. Theor Ecol 9(3):365–380
Peterman WE, Connette GM, Semlitsch RD, Eggert LS (2014) Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Molec Ecol 23(10):2402–2413. http://onlinelibrary.wiley.com/doi/10.1111/mec.12747/full
Potapov A, Muirhead JR, Lele SR, Lewis MA (2011) Stochastic gravity models for modeling lake invasions. Ecol Modell 222(4):964–972. http://www.sciencedirect.com/science/article/pii/S0304380010003820
Sæther BE, Engen S, Lande R (1999) Finite metapopulation models with density–dependent migration and stochastic local dynamics. Proc R Soc London B: Biol Sci 266(1415):113–118
Silva JA, De Castro ML, Justo DA (2001) Stability in a metapopulation model with density-dependent dispersal. Bull Math Biol 63(3):485–505
Soares-Filho BS, Cerqueira GC, Pennachin CL (2002) Dinamicaa stochastic cellular automata model designed to simulate the landscape dynamics in an amazonian colonization frontier. Ecol Modell 154(3):217–235
Söndgerath D, Schröder B (2002) Population dynamics and habitat connectivity affecting the spatial spread of populations–a simulation study. Landsc Ecol 17(1):57–70
Spear SF, Balkenhol N, Fortin MJ, McRae BH, Scribner K (2010) Use of resistance surfaces for landscape genetic studies: considerations for parameterization and analysis. Molec Ecol 19(17):3576–3591. http://onlinelibrary.wiley.com/doi/10.1111/j.1365-294X.2010.04657.x/full
Strevens CM, Bonsall MB (2011) Density-dependent population dynamics and dispersal in heterogeneous metapopulations. J Anim Ecol 80(1):282–293
Taylor CM, Laughlin AJ, Hall RJ (2016) The response of migratory populations to phenological change: a migratory flow network modelling approach. J Anim Ecol 85(3):648–659
Truscott J, Ferguson NM (2012) Evaluating the adequacy of gravity models as a description of human mobility for epidemic modelling. PLoS Comput Biol 8(10):e1002,699
Xia Y, Bjørnstad ON, Grenfell BT (2004) Measles metapopulation dynamics: a gravity model for epidemiological coupling and dynamics. Am Nat 164(2):267–281
Funding
Funding for this work was made available from the ByWater Institute at Tulane University and the National Science Foundation (BCS-1313703).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rael, R., Taylor, C. A flow network model for animal movement on a landscape with application to invasion. Theor Ecol 11, 271–280 (2018). https://doi.org/10.1007/s12080-018-0373-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12080-018-0373-4