Abstract
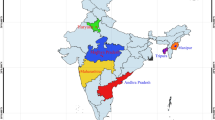
The phenotypic and microsatellite marker information of nine strains of catla (Catla catla) for growth trait was used to infer relationship within and between strains. This information helped in optimizing the proportion of individuals to be used from each strain while creating a base population for selective breeding. For this purpose, nine strains were collected from different sources and places of India namely West Bengal, Bihar, Odisha, Andhra Pradesh and Uttar Pradesh. Two riverine sources i.e. Ganga and Subarnarekha were also represented among the nine strains collected for base population. They were brought to Indian Council of Agricultural Research-Central Institute of Freshwater Aquaculture (ICAR-CIFA) at fry stage and reared separately till fingerlings. After passive integrated transponder tagging fingerlings were stocked in three communal ponds for one year culture. Live body weights were then measured and least square means were obtained after pond effect correction. A wide range of variation was observed among and between strains. Microsatellite markers were used to estimate genetic differences of different strains of catla using pair wise \(F_{\mathrm{ST}}\) estimates. Overall multi locus \(F_{\mathrm{ST}}\), including all loci was estimated to be 0.4137 (\(P<0.05\)), indicating genetic heterogeneity among them. Analysis of molecular variance revealed that, 58.63% of variation was due to within individual variation, 3.45% of variation was due to among individuals within strain and 37.92% was due to among strain variations. Both phenotypic as well as microsatellite data will be used to form a base population of catla with individuals from the stock having broad genetic variation for selective breeding programme.





Similar content being viewed by others
References
Abowei J. F. N. 2010 The condition factor, length – weight relationship and abundance of Ilisha africana (Block, 1795) from Nkoro River Niger Delta, Nigeria. Adv. J. Food Sci. Technol. 2, 6–11.
An H. S., Lee J. W., Kim H. Y., Kim J. B., Chang D. S., Park J. Y. et al 2013 Genetic differences between wild and hatchery populations of Korean spotted sea bass (Lateolabrax maculatus) inferred from microsatellite markers. Genes Genomics 35, 671–680.
Beverton R. J. H. and Holt S. J. 1957 On the dynamics of exploited fish populations. Bulletin Fishery Research Board, Canada. 19, 300
Brown B. and Epifanio J. 2003 Nuclear DNA. In Population genetics: principles and applications for fisheries scientists (ed. E. M. Hallermann), pp. 458. American Fisheries Society, Bethesda, MD.
Bunje P. M., Barluenga M. and Meyer A. 2007 Sampling genetic diversity in the sympatrically and allopatrically speciating Midas cichlid species complex over a 16 year time series. BMC Evol. Biol. 7, 25.
Caballero A. and Toro M. A. 2002 Analysis of genetic diversity for the management of conserved subdivided populations. Conserv. Genet. 3, 289–299.
Chauhan T. and Rajiv K. 2010 Molecular markers and their applications in fisheries and aquaculture. Adv. Biosci. Biotechnol. 1, 281–291.
DAHDF. 2016 Department of Animal Husbandry, Dairying and Fisheries, Ministry of Agriculture and Farmers Welfare, Government of India, Annual Report (2016–17) (http://dahd.nic.in/sites/default/files/Annual%20Report%202016-17.pdf).
Das Mahapatra K., Gjerde B., Reddy P. V. G. K., Sahoo M., Jana R. K., Saha J. N. et al 2001 Tagging: on the use of passive integrated transponder (PIT) tag for identification of fishes. Aquacult. Res. 32, 47–50.
Das P., Barat A., Meher P. K., Ray P. P. and Majumdar D. 2005 Isolation and characterization of polymorphic microsatellites in Labeo rohita and their cross-species amplification in related species. Mol. Ecol. Notes 5, 231–233.
Das R., Mohindra V., Singh R. K., Lal K. K., Punia P., Masih P. et al 2012 Intraspecific genetic diversity in wild Catla catla (Hamilton, 1822) populations assessed through mtDNA cytochrome b sequences. J. Appl. Ichthyol. 28, 280–283.
Das S. P., Swain S., Bej D., Jayasankar P., Jena J. K. and Das P. 2015 Length–weight relationships of four Cyprinid species in India. J. Appl. Ichthyol. 31, 583–584.
Earl D. A. 2012 STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361.
Eding H. and Meuwissen T. H. E. 2001 Marker based estimates of between and within population kinships for the conservation of genetic diversity. J. Anim. Breed. Genet. 118, 141–159.
Eding J. H., Crooijmans R. P. M. A., Groenen M. A. M. and Meuwissen T. H. E. 2002 Assessing the contribution of breeds to genetic diversity in conservation schemes. Genet. Sel. Evol. 34, 613–633.
Eknath A. E. and Doyle R. W. 1990 Effective population size and rate of inbreeding in aquaculture of Indian major carps. Aquaculture 85, 293–305.
Eknath A. E., Tayamen M. M., Palada-de-Vera M. S., Danting J. C., Reyes R. A., Dinosio E. E. et al 1993 Genetic improvement of farmed tilapia: the growth performance of eight strains of Oreochromis niloticus tested in different farm environments. Aquaculture 111, 171–188.
Evanno G., Regnaut S. and Goudet J. 2005 Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620.
Fernandez J., Toro M. A., Sonesson A. K. and Villanueva B. 2014 Optimizing the creation of base populations for aquaculture breeding programs using phenotypic and genomic data and its consequences on genetic progress. Front. Genet. 5, 414.
Froese R. 1998 Length–weight relationships for 18 less studied fish species. J. Appl. Ichthyol. 14, 117–118.
Froese R. 2006 Cube law, condition factor and weight–length relationships: history, meta-analysis and recommendations. J. Appl. Ichthyol. 22, 241–253.
Fulton T. W. 1904 The rate of growth of fishes. Twenty-second Annual Report, Part III. Fisheries Board of Scotland, Edinburgh, pp. 141–241.
Gjedrem T., Gjoen H. M. and Gjerde B. 1991 Genetic origin of Norwegian farmed Atlantic salmon. Aquaculture 98, 41–50.
Glaubitz J. C. 2004 CONVERT: a user-friendly program to reformat diploid genotypic data for commonly used population genetic software packages. Mol. Ecol. Notes 4, 309–310.
Hammer O., Harper D. A. T. and Ryan P. D. 2001 PAST: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 4, 9.
Holtsmark M., Sonesson A. K., Gjerde B. and Klemetsdal G. 2006 Number of contributing subpopulations and mating design in the base population when establishing a selective breeding program for fish. Aquaculture 258, 241–249.
Karlsson S. and Mork J. 2005 Deviation from Hardy–Weinberg equilibrium, and temporal instability in allele frequencies at microsatellite loci in a local population of Atlantic cod. ICES J. Mar. Sci. 62, 1588–1596.
Langen K., Schwarzer J., Kullmann H., Bakker T. C. and Thünken T. 2011 Microsatellite support for active inbreeding in a cichlid fish. PLoS One 6, e24689.
Le Cren E. D. 1951 The length–weight relationship and seasonal cycle in gonad weight and condition in Perch (Perca fluviatitus). J. Anim. Ecol. 20, 201–219.
Liu Z. J. and Cordes J. F. 2004 DNA marker technologies and their applications in aquaculture genetics. Aquaculture 238, 1–37.
Ma K. Y., Feng J. B. and Li J. L. 2012 Genetic variation based on microsatellite analysis of the oriental river prawn, Macrobrachium nipponense from Qiandao Lake in China. Genet. Mol. Res. 11, 4235–4244.
Mojekwu T. O. and Anumudu C. I. 2013 Microsatellite markers in Aquaculture: application in fish population genetics. IOSR-JESTFT 5, 43–48.
Oosterhout Van C., Hutchinson W. F., Wills D. P. M. and Shipley P. 2004 Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 4, 535–538.
Patel A., Das P., Swain S. K., Meher P. K., Jayasankar P. and Sarangi N. 2009 Development of 21 new microsatellite markers in Labeo rohita (rohu). Anim. Genet. 40, 253–254.
Pritchard J. K., Stephens M. and Donnelly P. 2000 Inference of population structure using multilocus genotype data. Genetics 155, 945–959.
Puurtinen M., Knott K. E., Suonpää S., Ooik T. V. and Kaitala V. 2004 Genetic variability and drift load in populations of an aquatic snail. Evolution 58, 749–756.
Rahman S. Z., Khan M. R., Islam S. and Alam S. 2009 Genetic variation of wild and hatchery populations of the catla Indian major carp (Catla catla Hamilton 1822: Cypriniformes, Cyprinidae) revealed by RAPD markers. Genet. Mol. Biol. 32, 197–201.
Rana R. S., Bhat K. V., Lakhanpal S. and Lakra W. S. 2004 Comparative genetic diversity in natural and hatchery populations of Indian major carps (C. catla and L. rohita). Asian-Australas J. Anim. Sci. 17, 1197–1203.
Reddy P. V. G. K., Gjerde B., Tripathy S. D., Jana R. K., Mahapatra K. D., Gupta S. D. et al 2002 Growth and survival of six stocks of rohu (Labeo rohita) in mono and polyculture system. Aquaculture 203, 239–250.
Saccheri I., Kuussaari M., Kankare M., Vikman P., Fortelius W. and Hanski I. 1998 Inbreeding and extinction in a butterfly metapopulation. Nature 392, 491–494.
Sambrook J., Fritsch E. F. and Maniatis T. 1989 Molecular cloning: a laboratory manual, 2nd edition, pp. 9.14–9.23. Cold Spring Harbor Laboratory Press, New York.
Sarang N., Shrivas A. K., Sao S. and Vardia H. K. 2015 Length–weight relationship and condition factor of Catla catla in Chhirpani Reservoir, Chhattisgarh, India. IOSR-JESTFT 1, 33–37.
Sekino M., Hara M. and Taniguchi N. 2002 Genetic diversity within and between hatchery strains of Japanese flounder Paralichthys olivaceus assessed by means of microsatellite and mitochondrial DNA sequencing analysis. Aquaculture 213, 101–122.
Thomas J., Venu S. and Kurup B. M. 2003 Length–weight relationship of some deep-sea fish inhabiting continental slope beyond 250 m depth along the West coast of India. NAGA, World Fish Center Q. 26, 17–21.
Acknowledgements
This work was carried out under an ICAR-CIFA Institute based project and DBT-COE project. The authors are thankful to the Directors of this Institute for providing facilities and funding for the project. We are also thankful to Dr Ramesh Rathod, scientist and Dr B. Mishra, technical officer for their help during collection of different strains of catla.
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Indrajit Nanda
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Das Mahapatra, K., Sahoo, L., Saha, J.N. et al. Establishment of base population for selective breeding of catla (Catla catla) depending on phenotypic and microsatellite marker information. J Genet 97, 1327–1337 (2018). https://doi.org/10.1007/s12041-018-1034-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-018-1034-5