Abstract
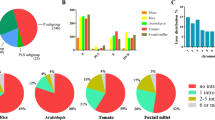
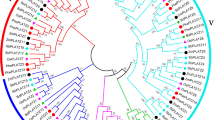
Pentatricopeptide repeat (PPR) gene family plays an essential role in the regulation of plant growth and organelle gene expression. Some PPR genes are related to fertility restoration in plant, but there is no detailed information in Gossypium. In the present study, we identified 482 and 433 PPR homologues in Gossypium raimondii (\(\hbox {D}_{5}\)) and G. arboreum (\(\hbox {A}_{2}\)) genomes, respectively. Most PPR homologues showed an even distribution on the whole chromosomes. Given an evolutionary analysis to PPR genes from G. raimondii (\(\hbox {D}_{5}\)), G. arboreum (\(\hbox {A}_{2}\)) and G. hirsutum genomes, eight PPR genes were clustered together with restoring genes of other species. Most cotton PPR genes were qualified with no intron, high proportion of \(\upalpha \)-helix and classical tertiary structure of PPR protein. Based on bioinformatics analyses, eight PPR genes were targeted in mitochondrion, encoding typical P subfamily protein with protein binding activity and organelle RNA metabolism in function. Further verified by RNA-seq and quantitative real-time PCR (qRT-PCR) analyses, two PPR candidate genes, Gorai.005G0470 (\(\hbox {D}_{5}\)) and Cotton_A_08373 (\(\hbox {A}_{2}\)), were upregulated in fertile line than sterile line. These results reveal new insights into PPR gene evolution in Gossypium.






Similar content being viewed by others
References
Akagi H., Nakamura A., Yokozeki-Misono Y., Inagaki A., Takahashi H., Mori K. et al. 2004 Positional cloning of the rice Rf-1 gene, a restorer of BT-type cytoplasmic male sterility that encodes a mitochondria-targeting PPR protein. Theor. Appl. Genet. 108, 1449–1457.
Altschul S. F., Gish W., Miller W., Myers E. W. and Lipman D. J. 1990 Basic local alignment search tool. J. Mol. Biol. 215, 403–410.
Aubourg S., Boudet N., Kreis M. and Lecharny A. 2000 In Arabidopsis thaliana, 1% of the genome codes for a novel protein family unique to plants. Plant Mol. Biol. 42, 603–613.
Barkan A. and Small I. 2014 Pentatricopeptide repeat proteins in plants. Annu. Rev. Plant Biol. 65, 415–442.
Bentolila S., Alfonso A. A. and Hanson M. R. 2002 A pentatricopeptide repeat-containing gene restores fertility to cytoplasmic male-sterile plants. Proc. Natl. Acad. Sci. USA 99, 10887–10892.
Brown G. G., Formanova N., Jin H., Wargachuk R., Dendy C., Patil P. et al. 2003 The radish Rfo restorer gene of Ogura cytoplasmic male sterility encodes a protein with multiple pentatricopeptide repeats. Plant J. 35, 262–272.
Carlsson J., Leino M., Sohlberg J., Sundstrom J. F. and Glimelius K. 2008 Mitochondrial regulation of flower development. Mitochondrion 8, 74–86.
Chen Z., Feng K., Grover C. E, Li P., Liu F., Wang Y. et al. 2016 Chloroplast DNA structural variation, phylogeny, and age of divergence among diploid cotton species. PLoS One 11, e0157183.
Chen Z., Grover C. E., Li P., Wang Y., Nie H., Zhao Y. et al. 2017a Molecular evolution of the plastid genome during diversification of the cotton genus Mol. Phylogenet. Evol. 112, 268–278.
Chen Z., Nie H., Grover C. E., Wang Y., Li P., Wang M. et al. 2017b Entire nucleotide sequences of Gossypium raimondii and G. arboreum mitochondrial genomes revealed A-genome species as cytoplasmic donor of the allotetraploid species. Plant Biol. 19, 484–493.
Chen Z., Zhao N., Li S., Grover C. E., Nie H., Wendel J. F. et al. 2017c Plant mitochondrial genome evolution and cytoplasmic male sterility. Crit. Rev. Plant Sci. 36, 55–69.
Cui X., Wise R. P. and Schnable P. S. 1996 The rf2 nuclear restorer gene of male-sterile T-cytoplasm maize. Science 272, 1334–1336.
Cushing D. A., Forsthoefel N. R., Gestaut D. R. and Vernon D. M. 2005 Arabidopsis emb175 and other ppr knockout mutants reveal essential roles for pentatricopeptide repeat (PPR) proteins in plant embryogenesis. Planta 221, 424–436.
Desloire S., Gherbi H., Laloui W., Marhadour S., Clouet V., Cattolico L. et al. 2003 Identification of the fertility restoration locus, Rfo, in radish, as a member of the pentatricopeptide-repeat protein family. EMBO Rep. 4, 588–594.
Dewey R. E., Timothy D. H. and Levings C. S. 1987 A mitochondrial protein associated with cytoplasmic male sterility in the T cytoplasm of maize. Proc. Natl. Acad. Sci. USA 84, 5374–5378.
Feng C. D., Stewart J. M. and Zhang J. F. 2005 STS markers linked to the Rf1 fertility restorer gene of cotton. Theor. Appl. Genet. 110, 237–243.
Fujii S. and Toriyama K. 2009 Suppressed expression of retrograde-regulated male sterility restores pollen fertility in cytoplasmic male sterile rice plants. Proc. Natl. Acad. Sci. USA 106, 9513–9518.
Fujii S., Bond C. S. and Small I. D. 2011 Selection patterns on restorer-like genes reveal a conflict between nuclear and mitochondrial genomes throughout angiosperm evolution. Proc. Natl. Acad. Sci. USA 108, 1723–1728.
Gallagher J. P., Grover C. E., Rex K., Moran M. and Wendel J. F. 2017 A new species of cotton from Wake Atoll, Gossypium stephensii (Malvaceae). Syst. Bot. 42, 115–123.
Galtier N. 2011 The intriguing evolutionary dynamics of plant mitochondrial DNA. BMC Biol. 9, 61.
Geddy R. and Brown G. G. 2007 Genes encoding pentatricopeptide repeat (PPR) proteins are not conserved in location in plant genomes and may be subject to diversifying selection. BMC Genomics 8, 130.
Germain A., Hotto A. M., Barkan A. and Stern D. B. 2013 RNA processing and decay in plastids. Wiley Interdiscip. Rev. RNA 4, 295–316.
Giancola S., Marhadour S., Desloire S., Clouet V., Falentin-Guyomarc’h H., Laloui W. et al. 2003 Characterization of a radish introgression carrying the Ogura fertility restorer gene Rfo in rapeseed, using the Arabidopsis genome sequence and radish genetic mapping. Theor. Appl. Genet. 107, 1442–1451.
Gillman J. D., Bentolila S. and Hanson M. R. 2007 The petunia restorer of fertility protein is part of a large mitochondrial complex that interacts with transcripts of the CMS-associated locus. Plant J. 49, 217–227.
Hashimoto M., Endo T., Peltier G., Tasaka M. and Shikanai T. 2003 A nucleus-encoded factor, CRR2, is essential for the expression of chloroplast ndhB in Arabidopsis. Plant J. 36, 541–549.
Howell M. D., Fahlgren N., Chapman E. J., Cumbie J. S., Sullivan C. M., Givan S. A. et al. 2007 Genome-wide analysis of the RNA-denpendent RNA polymerase6/DICER-like4 pathway in Arabidopsis reveals dependency on miRNA- and tasiRNA-directed targeting. Plant Cell 19, 926–942.
Hu J., Wang K., Huang W., Liu G., Gao Y., Wang J. et al. 2012 The rice pentatricopeptide repeat protein RF5 restores fertility in Hong-Lian cytoplasmic male-sterile lines via a complex with the glycine-rich protein GRP162. Plant Cell 24, 109–122.
Itabashi E., Iwata N., Fujii S., Kazama T. and Toriyama K. 2011 The fertility restorer gene, Rf2, for lead rice-type cytoplasmic male sterility of rice encodes a mitochondrial glycine-rich protein. Plant J. 65, 359–367.
Janska H., Sarria R., Woloszynska M., Arrieta-Montiel M. and Mackenzie S. A. 1998 Stoichiometric shifts in the common bean mitochondrial genome leading to male sterility and spontaneous reversion to fertility. Plant Cell 10, 1163–1180.
Jore M. M., Lundgren M., van Duijn E., Bultema J. B., Westra E. R., Waghmare S. P. et al. 2011 Structural basis for CRISPR RNA-guided DNA recognition by Cascade. Nat. Struct. Mol. Biol. 18, 529–536.
Katoh K. and Standley D. M. 2013 MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780.
Kazama T. and Toriyama K. 2003 A pentatricopeptide repeat-containing gene that promotes the processing of aberrant atp6 RNA of cytoplasmic male-sterile rice. FEBS Lett. 544, 99–102.
Klein R. R., Klein P. E., Mullet J. E., Minx P., Rooney W. L. and Schertz K. F. 2005 Fertility restorer locus Rf1 of sorghum (Sorghum bicolor L.) encodes a pentatricopeptide repeat protein not present in the colinear region of rice chromosome 12. Theor. Appl. Genet. 111, 994–1012.
Koizuka N., Imai R., Fujimoto H., Hayakawa T., Kimura Y., Kohno-Murase J. et al. 2003 Genetic characterization of a pentatricopeptide repeat protein gene, orf687, that restores fertility in the cytoplasmic male-sterile Kosena radish. Plant J. 34, 407–415.
Komori T., Ohta S., Murai N., Takakura Y., Kuraya Y., Suzuki S. et al. 2004 Map-based cloning of a fertility restorer gene,Rf-1, in rice (Oryza sativa L.). Plant J. 37, 315–325.
Lei B., Li S., Liu G., Chen Z., Su A., Li P. et al. 2013 Evolution of mitochondrial gene content: loss of genes, tRNAs and introns between Gossypium harknessii and other plants. Plant Syst. Evol. 299, 1889–1897.
Li F., Fan G., Wang K., Sun F., Yuan Y., Song G. et al. 2014 Genome sequence of the cultivated cotton Gossypium arboreum. Nat. Genet. 46, 567–572.
Li P., Cao M., Yang L., Xu A. and Liu H. 2007 Mapping of fertility restorer gene for cotton cytoplasmic male sterile line Jin A. Acta Bot. Bor-Occid. Sin. 27 1937–1942.
Li S., Liu G., Chen Z., Wang Y., Li P., and Hua J. 2013. Construction and initial analysis of five Fosmid libraries of mitochondrial genomes of cotton (Gossypium). Chinese Sci. Bull. 58, 4608–4615.
Liu F., Cui X., Horner H. T., Weiner H. and Schnable P. S. 2001 Mitochondrial aldehyde dehydrogenase activity is required for male fertility in maize. Plant Cell 13, 1063–1078.
Liu L., Guo W., Zhu X. and Zhang T. 2003 Inheritance and fine mapping of fertility restoration for cytoplasmic male sterility in Gossypium hirsutum L. Theor. Appl. Genet. 106, 461–469.
Luo D., Xu H., Liu Z., Guo J., Li H., Chen L. et al. 2013 A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice. Nat. Genet. 45, 573–577.
Lurin C., Andres C., Aubourg S., Bellaoui M., Bitton F., Bruyere C. et al. 2004 Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 16, 2089–2103.
Mackenzie S. A. and Chase C. D. 1990 Fertility restoration is associated with loss of a portion of the mitochondrial genome in cytoplasmic male-sterile common bean. Plant Cell 2, 905–912.
Matsuhira H., Kagami H., Kurata M., Kitazaki K., Matsunaga M., Hamaguchi Y. et al. 2012 Unusual and typical features of a novel restorer-of-fertility gene of sugar beet (Beta vulgaris L.). Genetics 192, 1347–1358.
Meierhoff K., Felder S., Nakamura T., Bechtold N. and Schuster G. 2003 HCF152, an Arabidopsis RNA binding pentatricopeptide repeat protein involved in the processing of chloroplast psbB-psbT-psbH-petB-petD RNAs. Plant Cell 15, 1480–1495.
Melonek J., Stone J. D. and Small I. 2016 Evolutionary plasticity of restorer-of-fertility-like proteins in rice. Sci. Rep. UK 6, 35152.
Meyer V. G. 1975 Male sterility from Gossypium harknessii. J. Hered. 66, 23–27.
Mistry J., Finn R. D., Eddy S. R. Bateman A. and Punta M. 2013 Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res. 41, e121.
Nakamura T., Meierhoff K., Westhoff P. and Schuster G. 2003 RNA-binding properties of HCF152, an Arabidopsis PPR protein involved in the processing of chloroplast RNA. Eur. J. Biochem. 270, 4070–4081.
O’Toole N., Hattori M., Andres C., Iida K., Lurin C., Schmitz-Linneweber C. et al. 2008 On the expansion of the pentatricopeptide repeat gene family in plants. Mol. Biol. Evol. 25, 1120–1128.
Paterson A. H.,Wendel J. F., Gundlach H., Guo H., Jenkins J., Jin D. et al. 2012 Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492, 423–427.
Schnable P. S. and Wise R. P. 1998 The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci. 3, 175–180.
Small I. D. and Peeters N. 2000 The PPR motif - a TPR-related motif prevalent in plant organellar proteins. Trends Biochem. Sci. 25, 46–47.
Suzuki H., Yu J., Ness S. A., O’Connell M. A. and Zhang J. 2013 RNA editing events in mitochondrial genes by ultra-deep sequencing methods: a comparison of cytoplasmic male sterile, fertile and restored genotypes in cotton. Mol. Genet. Genomics 288, 445–457.
Sykes T., Yates S., Nagy I., Asp T., Small I. and Studer B. 2017 In silico identification of candidate genes for fertility restoration in cytoplasmic male sterile perennial ryegrass (Lolium perenne L.). Genome Biol. Evol. 9, 351–362.
Tamura K., Peterson D., Peterson N., Stecher G., Nei M. and Kumar S. 2011 MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739.
Wang F., Yue B., Hu J. G., Stewart J. M. and Zhang J. F. 2009 A target region amplified polymorphism marker for fertility restorer gene Rf1 and chromosomal localization of Rf1 and Rf2 in cotton. Crop Sci. 49, 1602–1608.
Wang Z., Zou Y., Li X., Zhang Q., Chen L., Wu H. et al. 2006 Cytoplasmic male sterility of rice with boro II cytoplasm is caused by a cytotoxic peptide and is restored by two related PPR motif genes via distinct modes of mRNA silencing. Plant Cell 18, 676–687.
Wang Z. W., De Wang C., Gao L., Mei S. Y., Zhou Y., Xiang C. P. et al. 2013 Heterozygous alleles restore male fertility to cytoplasmic male-sterile radish (Raphanus sativus L.): a case of overdominance. J. Exp. Bot. 64, 2041–2048.
Wendel J. F. 1989 New World tetraploid cottons contain Old World cytoplasm. Proc. Natl. Acad. Sci. USA 86, 4132–4136.
Wendel J. F. and Grover C. E. 2015 Taxonomy and evolution of the cotton genus. In Cotton (ed. D. Fang and R. Percy), pp. 25–44. American Society of Agronomy, Crop Science Society of America, and Soil Science Society of America, Madison.
Wu J. Y., Cao X. X., Guo L. P., Qi T. X., Wang H. L., Tang H. N. et al. 2014 Development of a candidate gene marker for Rf1 based on a PPR gene in cytoplasmic male sterile CMS-\(\text{ D }_{2}\) upland cotton. Mol. Breed. 34, 231–240.
Xia X. and Xie Z. 2001 DAMBE: Software package for data analysis in molecular biology and evolution. J. Hered. 92, 371–373.
Yang Z. and Nielsen R. 2000 Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17, 32–43.
Yin J., Guo W., Yang L., Liu L. and Zhang T. 2006 Physical mapping of the Rf1 fertility-restoring gene to a 100 kb region in cotton. Theor. Appl. Genet. 112, 1318–1325.
Zabala G., Gabay-Laughnan S. and Laughnan J. R. 1997 The nuclear gene Rf3 affects the expression of the mitochondrial chimeric sequence R implicated in S-type male sterility in maize. Genetics 147, 847–860.
Zhang J. F. and Stewart J. M. 2004 Identification of molecular markers linked to the fertility restorer genes for \(\text{ CMS }\text{-D }_{8}\) in cotton. Crop Sci. 44, 1209–1217.
Zhang T., Hu Y., Jiang W., Fang L., Guan X. and Chen J. 2015 Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 33, 531–537.
Zhang X., Wang L., Xu X., Cai C. and Guo W. 2014 Genome-wide identification of mitogen-activated protein kinase gene family in Gossypium raimondii and the function of their corresponding orthologs in tetraploid cultivated cotton. BMC Plant Biol. 14, 345.
Zhao L., Yuanda L., Caiping C., Xiangchao T., Xiangdong C., Wei Z. et al. 2012 Toward allotetraploid cotton genome assembly: integration of a high-density molecular genetic linkage map with DNA sequence information. BMC Genomics 13, 539.
Acknowledgements
We are indebted to Dr. Anming Ding (Tobacco Research Institute, Chinese Academy of Agricultural Sciences, Qingdao, China) for supplying HMMER matrix of PPR gene family in Arabidopsis (defined by Prof. Small Ian). We thank Dr. Zhen Su (State Key Laboratory of Plant Physiology and Biochemistry, College of Biological Sciences, China Agricultural University, Beijing, China) for helpful advises and discussion. This work was supported by the National Natural Science Foundation of China (31671741) and National Key R & D Program for Crop Breeding (2016YFD0100203).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Manoj Prasad
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, N., Wang, Y. & Hua, J. Genomewide identification of PPR gene family and prediction analysis on restorer gene in Gossypium. J Genet 97, 1083–1095 (2018). https://doi.org/10.1007/s12041-018-0993-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-018-0993-x