Abstract
Synaptic protein shanks (SH3 and multiple ankyrin repeat domains protein, Shank) have emerged as an important mediator of synaptic remodeling. Synaptic remodeling is a common pathogenic process in various neurological disorders including epilepsy. However, the expression and function of shanks gene in epileptogenesis has not been investigated to date. Herein, we investigated the expression of shanks (shank1/2/3) mRNA expression in both epileptic rats and epilepsy patients. Furthermore, methyl target sequencing was applied to explore the relationship between shank mRNA expression and DNA methylation in both rats and human patients. In general rat model, shank1/2/3 mRNA was downregulated at acute stage, upregulated at latent stage, and returned to the basal level at chronic stage. Ten CpG sites of shank1 was found differentially methylated, out of which 6 were hypermethylated. Seventeen CpG sites of shank3 were differentially methylated, out of which 8 were hypermethylated. In human epilepsy patients, decreased shank2 mRNA was detected from the brain tissue, with DNA hypermethylation dominant from both brain (18 out of 30) and blood tissue (58 out of 80), indicating the regulation role of DNA methylation on shank2 expression. In conclusion, our finding suggests the participation of the shanks gene in the pathophysiology of seizure, out of which 2 shank3 CpG sites (Chr7: 130473419, and Chr7: 130473405) may play an important role in shank3 expression at both the acute and latent stages in the SE rat model.
Similar content being viewed by others
Introduction
There is convincing evidence that rapid and well-coordinated changes in the quantitative and qualitative content of proteins at the synapse ensure its dynamic plasticity in response to external stimuli and underlie physical function. Disruption of local translation control at synapses has been thought of as a common pathophysiological mechanism underlying the aggravation of a wide range of neurological conditions including epilepsy [1, 2], autism [1, 2], and Parkinson’s [3] and Alzheimer’s diseases [4]. Prolonged seizure causes a shift in the balance of synaptic plasticity and leads to changes in behavior, cognitive abilities, and memory. In turn, the disruption of the synaptic structure and function such as the imbalance between excitatory and inhibitory neurotransmitters, the loss of presynaptic protein such as Caspr4p [5], the disturbance of neurotransmitter release [6], and the impaired signal transmission on the postsynaptic membrane [7] all contribute the to the pathophysiology of epileptic seizure.
Among the synaptic proteins, the family of SHANK proteins acts as scaffold proteins required for the development and function of neuronal synapses. SHANK proteins were encoded by shanks genes, including shank1, shank2, and shank3. Shank proteins are multidomain scaffold proteins located at the postsynaptic density (PSD) area, playing an important role in regulating the postsynaptic signaling pathway. Shank proteins connect neurotransmitter receptors such as glutamate receptors (GKAP, PSD95, and Homer) and ion channels to the G protein-coupled cAMP/PKA and IP3/PKC signaling pathways [8], interact with N-methyl-d-aspartic acid receptor (NMDAR) [8, 9], and regulate synaptic plasticity such as long-term potentiation or long-term depression [9], playing a crucial role in regulating synaptic function. Deletion of shanks gene was demonstrated to be associated with autism spectrum disorder [10], Phelan-McDermid syndrome [11], and schizophrenia [12]. However, the expression and regulation of shanks gene in the pathophysiology of epilepsy has not been investigated to date.
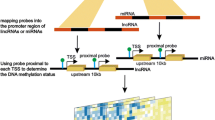
Epigenetic signaling such as DNA methylation exerts high fidelity regulation on gene expression. DNA methylation occurs on the cytosine bases in DNA and on the lysine and arginine residues on histone tails. Methyl group was transferred from S-adenosylmethionine to cytosine bases at CpG (5′-cytosine-phosphate-guanine-3′) dinucleotide sequences forming 5-methylcytosine. Symmetrical methylation of DNA induces a conformational change to its structure that prevents transcription factors from recognizing the DNA site, thus leading to a silencing effect on gene expression. Currently, there is strong evidence that DNA methylation is playing a crucial role in the pathophysiology of epilepsy [13, 14]. At the onset of epilepsy, an increase in DNA methylation takes place, which leads to increased seizure severity and contributes to the development of epilepsy [15]. Hypermethylation of certain extracellular matrix proteins such as reelin is directly implicated in the pathophysiology of temporal lobe epilepsy [14]. Inhibiting DNA methylation via DNA methyltransferase inhibitor resulted in Gria2 upregulation and blocked the seizure-inducing effect of kainic acid in the hippocampus [13]. Recently, in 2019, it was also reported that DNA methylation regulates neural function by modifying synaptic structure and function, thus contributing to the epileptogenesis [16]. DNA methylation of shank3 was reported to downregulate the protein expression in rat hippocampal neurons [17]. Whether shanks gene expression was regulated by DNA methylation in epilepsy model dynamic time was unclear.
As a crucial regulator of the synaptic structure and function, the role of shanks gene in the pathophysiology of epilepsy has not been fully studied to date. The present study was the first to explore the role of shanks gene in epilepsy in both rats and human patients. We examined the dynamic change of mRNA expression and DNA methylation of shanks gene in rat hippocampus following pilocarpine-induced seizure. Furthermore, brain tissue from epilepsy patients was collected for mRNA and DNA methylation sequencing. We proposed that the shanks gene participates in epileptogenesis, with the mRNA and methylation expression distinct between rats and human; and the gene expression of shank3 might be regulated by DNA methylation as a negative control in the epileptic rat.
Materials and Methods
Chemicals and Drugs
Pilocarpine and lithium chloride were purchased from Sigma-Aldrich (St. Louis, MO). TRIZOL Reagent, PureLink RNA Mini Kit, and SuperScript II Reverse Transcriptase were from Invitrogen (Carlsbad, CA). SYBR® Green SuperMix was from Quanta BioSciences (MD, USA). PrimeScript RT reagent kit was from Takara Bio (Tonomachi, Japan). DNA extraction kit was from Generay Biotech Company (Shanghai, China). Illumina HiSeq reagent kit was from Illumina Inc. (San Diego, CA, USA). EZ DNA Methylation-Gold Kit was from Zymo Research Corp (Irvine, CA). Chloroform, isopropanol, and ethanol were from Shanghai Chemical Corp (Shanghai, China). DNeasy Blood & Tissue Kit was from Qiagen (Crawley, UK). The mouse monoclonal anti-5-methyl cytosine antibody was from Diagenode (NJ, USA). BioMag magnetic beads were from Bangs Laboratories (IN, USA). The CpG promoter microarray was from NimbleGen Systems (WI, USA). Infinium HumanMethylation450 BeadChip Kit was from Illumina (San Diego, CA, USA).
Experimental Animals
Animal use and care was performed in accordance with the Central South University Ethics Review of Experimental Animal Welfare, Administration Committee of Affairs Concerning Experimental Animals in Hunan Province, China. All experimental protocols were confirmed by the Animal Ethics Committee of Central South University, Changsha, China (number 201603296). Sprague-Dawley (SD) male rats (6–8 weeks old, 220–250 g) were obtained from the Animal Center of Central South University, Changsha, China. And then they were randomly divided into six groups (6 rats per group): a sham control group, and five groups with rats sacrificed at different time points after SE, particularly 1 day, 7 days, 14 days, 30 days, and 60 days after SE, respectively. Animals were raised in a temperature-controlled room with 12-h light/dark cycle and free access to water and food and acclimatized for at least 7 days before use.
Seizure Induction
Seizure was inducted as we previously published [18]. SD rats (6–8 weeks old, male) were used in the post-PILO treatment experiment: 8 rats for control and 40 for seizure induction. Rats were injected with lithium chloride (125 mg/kg, i.p.) 18–20 min before PILO hydrochloride (25 mg/kg, i.p.). Saline injection was used for the 8 control mice. About 40 min after PILO injection, thirty-two out of 40 rats developed status epilepticus (SE), the onset of which was defined as the occurrence of three stage 4 seizures (Supplement Table 1). To reduce the variation that might be introduced by repeated doses of pilocarpine, no additional pilocarpine was administered to those 8 rats that did not enter SE initially; they thus were removed from the experiment. Forty-five minutes after SE began, rats were treated with 10% chloral hydrate to interrupt seizures. During recovery, animals were fed moistened rodent chow, monitored daily, and injected with 5% dextrose in lactated Ringer’s solution (Baxter) (0.5 ml, i.p.) when necessary. At the appropriate time (1 day after SE, 7 days after SE, 14 days after SE, 30 days after SE, 60 days after SE), the animals were euthanized under deep anesthesia with 10% chloral hydrate (3 ml/kg, i.p.). The brains were collected and dissected for their hippocampus for biochemical study. Based on the epilepsy development phases, the rats were grouped into 3 stages: acute stage (1 day), latent stage (7 days, 14 days, 30 days), and chronic stage (60 days).
Quantitative Real-Time PCR (qPCR)
The total RNA from rat hippocampus was isolated using TRIzol (Invitrogen) with the PureLink RNA Mini Kit (Invitrogen). RNA purity and concentration were measured by A260/A280 ratio and A260 value, respectively. The first-strand complementary DNA (cDNA) synthesis was performed with 1 μg RNA, 0.25 μg random primers, and 200 units of SuperScript II Reverse Transcriptase (Invitrogen) in a reaction volume of 20 μl at 42 °C for 50 min. The reaction was ended by heating at 70 °C for 15 min. The qPCR was performed using 8 μl of 50× diluted cDNA, 0.4 μM of primers, and 2× B-R SYBR® Green SuperMix (Quanta BioSciences) with a final volume of 20 μl in the iQ5 Multicolor Real-Time PCR Detection System (Bio-Rad Laboratories) [19]. Cycling conditions were as follows: 95 °C for 2 min followed by 40 cycles of 95 °C for 15 s and then 60 °C for 1 min. Melting curve analysis was used to verify single-species PCR product. Fluorescent data were acquired at the 60 °C step. Three repetitions for each sample were done, with six animals at each time point. GAPDH was set as the reference gene, and the cycle threshold for GAPDH was subtracted from the cycle threshold measured for each gene of interest to yield ∆CT. And the expression relative to GAPDH was calculated using 2∆∆Ct. Samples without cDNA template served as negative controls. Primers used for qPCR are listed in Supplement Table 2.
Methyl Target Sequencing in Rats
Analysis of CpG Island
By the bioinformatics method, CpG island of shanks gene was identified between 2000 bp upstream the transcription start site and 1000 bp downstream the first exon (all phases included). The standard for a CpG island is defined as CpG frequency > 0.6, CG content > 50%, and length > 200 bp. Methylation-specific primers were designed with Primer3 software (Supplement Table 3).
DNA Extraction
Rats were sacrificed 24 h after SE, and the hippocampus was dissected carefully. DNA was extracted with a commercial kit (Generay Biotech Company, Shanghai, China) according to the manufacturer’s instruction. DNA purity and concentration were determined with a NanoDrop spectrophotometer (BioTek Company, USA), by measuring the optic density ratio of maximal absorbent wavelengths at 260 to 280 nm (OD260/280) with salmon sperm DNA as an internal standard. The extracted DNA sample was sodium bisulfite converted with EZ DNA Methylation-Gold Kit.
Reduced Representation Bisulfite Sequencing (RRBS)
Illumina adapter sequences and unique barcodes were introduced through an additional PCR step. A total of 1 U HotStarTaq polymerase (TAKARA) and 2 ml template DNA were added to 1× reaction buffer (TAKARA), 2 mM Mg2+, 0.2 mM dNTP, and 0.1 μM of each primer to a total volume of 20 ml. Samples were amplified as follows: 95 °C for 2 min; 11 cycles of 94 °C for 20 s, 63–0.5 °C per cycle for 40 s, 72 °C for 1 min; 24 cycles of 94 °C for 20 s, 65 °C for 30 s, 72 °C for 1 min; 72 °C for 2 min. Samples were amplified. Specific sequence was added to system for PCR reaction. A 20 μl mixture was prepared for each reaction and included 1× reaction buffer (NEB Q5TM), 0.3 mM dNTP, 0.25 μM of F primer, 0.25 μM of index primer, 1 U Q5TM DNA polymerase (NEB), and 1 μl diluted template. The cycling program was 98 °C for 30 s; 11 cycles of 98 °C for 10 s, 65 °C for 30 s, 72 °C for 30 s; 72 °C for 5 min. Libraries were then sequenced on an Illumina HiSeq before alignment and analysis.
Methyl Target Sequencing in Brain Tissue from Epilepsy Patients
Subjects
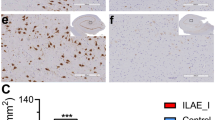
As is previously published [20], brain tissue was obtained from 9 epilepsy patients (Supplement Table 4A) with drug-refractory seizure who had undergone surgical removal of epileptogenic zone in their temporal neocortex. The epileptogenic zones were determined with a combination of ictal semiology, brain imaging (magnetic resonance imaging, MRI), video-EEG, sphenoidal electrode monitoring, and intraoperative electrocorticography (EcoG) considered (Fig. 5a). For comparison, histologically normal anterior temporal neocortex samples were obtained from 7 patients (Supplement Table 4B) who were treated for post-trauma intracranial hypertension. These subjects had no history of epilepsy. The dissected brain tissue was immediately placed in cryovials containing buffered diethylpyrocarbonate and stored in liquid nitrogen until use. All experimental procedures were confirmed by the Ethics Committee of Central South University, Changsha, China (number 201603297).
Methylated-Cytosine DNA Immunoprecipitation Chip
Genomic DNA was extracted from brain tissues using the DNeasy Blood & Tissue Kit according to the manufacturer’s protocol (Qiagen, Crawley, UK). Immunoprecipitation of methylated DNA was performed using a mouse monoclonal anti-5-methyl cytosine antibody (Diagenode) and BioMag magnetic beads (Bangs Laboratories, USA). The MeDIP quality was assessed by qPCR of H19 (positive control) and GAPDH (negative control). For each sample, 1 μg of input and IP DNA was labeled with Cy3- and Cy5-labeled random 9-mers, respectively, and co-hybridized to the NimbleGen HG 18 CpG promoter microarray (NimbleGen, WI, USA). Scanning was performed with an Axon GenePix 4000B microarray scanner. Raw data were extracted as pair files by NimbleScan software (NimbleGen, USA). Several procedures were then sequentially performed to obtain peak data with specified parameters (sliding window width 750 bp; mini probes per peak 2; P value minimum cutoff 2; maximum spacing between nearby probes per peak 500 bp).
Bisulfite Sequencing PCR
The methylation status of the CpG islands in candidate gene promoters was determined by bisulfite sequencing as described previously. The primers were designed using MethPrimer software. (Primer information is shown in Supplemental Table 3). DNA was bisulfite-treated and amplified by PCR. To determine the methylation status of the candidate gene promoters, an average of five positive clones were sequenced using the M13 reverse primer and an automated ABI prism 3730xl Genetic Analyzer (Applied Biosystems, USA). The sequencing results were analyzed using QUMA.
RT-PCR
Total RNA was isolated from brain tissue using TRIzol reagent according to the manufacturer’s protocol (Invitrogen, Paisley, UK). cDNA was synthesized from 1 μg total RNA using the PrimeScript RT reagent Kit (Takara). For PCR reactions, 2 ng cDNA was amplified in a 25 μl reaction volume containing 0.5 μM of each forward and reverse primer and 12.5 μl 2× PCR TaqMix (TianGen Beijing, China) under the following conditions: 5 min of denaturation at 95 °C; followed by 35 cycles of 30 s at 95 °C, 30 s at 60 °C, and 60 s at 72 °C; and a final extension at 72 °C for 10 min. The amplified products were separated on 2% agarose gels. Densitometry analysis was performed with Quantity One software (Bio-Rad Laboratories, USA).
mRNA Sequencing in Brain Tissue from Epilepsy Patients
Samples
Brain tissue was obtained from 6 epilepsy patients with drug-refractory seizure who had undergone surgical removal of epileptogenic zone in their temporal neocortex. The epileptogenic zones were determined with a combination of ictal semiology, brain imaging (magnetic resonance imaging, MRI), video-EEG, sphenoidal electrode monitoring, and intraoperative electrocorticography (EcoG) considered. For comparison, histologically normal anterior temporal neocortex samples were obtained from 6 patients who were treated for post-trauma intracranial hypertension. These subjects had no history of epilepsy. All experimental procedures were confirmed by the Animal Ethics Committee of Central South University, Changsha, China (number 201603297).
RNA Extraction
Total RNA was isolated using TRIzol Reagent (Invitrogen Life Technologies). The total RNA was tested by Agilent 2100 BioAnalyzer and RIN ≥ 7, the ratio of 28S to 18S ≥ 1.5, and the recommended starting dose was 0.1–1.0 μg. The total RNA was quantitatively evaluated with Qubit RNA Assay Kit.
Sequencing Libraries
Sequencing libraries were generated using the NEBNext Ultra RNA Library Prep Kit for Illumina (Illumina, San Diego, CA, USA). Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out with 5 μl rRNA depleted total RNA, 4 μl NEBNext First Strand Synthesis Reaction Buffer (5×), and 1 μl random primers under 94 °C for 15 min. First-strand cDNA was synthesized with fragmented and primed mRNA, murine RNase inhibitor, and ProtoScript II Reverse Transcriptase under 25 °C for 10 min, 42 °C for 50 min, and 70 °C for 15 min. Second-strand cDNA synthesis was subsequently performed with first-strand cDNA, second-strand synthesis reaction buffer (10×), and second-strand synthesis enzyme mix under 16 °C for 1 h. The product was purified with 144 μl AMPure XP Beads, and then resuspended by 60 μl nuclear-free water. In total, 55.5 μl cs cDNA was taken out to perform end repair, together with 6.5 μl NEBNext End Repair Reaction Buffer (10×) and 3 μl NEBNext End Prep Mix under 20 °C for 30 min, and 65 °C for 30 min. Adaptor ligation was performed with 65 μl dA-Tailed cDNA, 15 μl blunt/TA ligase Master Mix, and 1 μl diluted NEBNext Adaptor under 20 °C for 15 min, followed by 3 μl USER enzyme added into the system under 37 °C for 15 min. Product was purified with AMPure XP Beads and resuspended by resuspension buffer. DNA fragments with ligated adaptor molecules were selectively enriched with NEBNext Q5 Hot Start HiFi PCR Master Mix, Universal PCR Primer, and Index Primer under using Illumina PCR Primer Cocktail in a 15-cycle PCR reaction. Products were purified and quantified using the Qubit DNA HS Assay Kit. The sequencing library was then sequenced on a HiSeq platform (Illumina).
DNA Methylation Sequencing in Blood Sample from Epilepsy Patients
As is published in our previous paper [21], briefly, a total of 30 (18 males and 12 females) and 30 sex- and age-matched healthy controls were recruited through the epilepsy clinic of Xiangya Hospital from October 2010 to October 2014. Patients were treated with common anti-epileptics to control symptoms. DNA was extracted from blood samples from patients and control subjects using a commercial kit (Beijing Adly biological company, Beijing, China) according to manufacturer’s instructions. DNA concentration and purity were determined in a NanoDrop spectrophotometer. Samples with DNA concentration greater than 50 ng/μl and OD260/280 ratio from 1.60 to 2.10 were used for further experimentations. The pattern and extent of DNA methylation were assayed using the Infinium HumanMethylation450 BeadChip Kit (Illumina, Inc., San Diego, CA, USA). DNA samples were processed via a series of steps, including DNA denaturation, whole-genome amplification, fragmentation, precipitation, resuspension, and hybridization, according to manufacturer’s protocols. The signal of the final hybridization products was captured with the iScan system (Illumina, Inc.).
Statistical Analysis
Means of designated comparing groups were statistically analyzed using SPSS 18.0 Software Student t test (e.g., age comparison) or nonparametric test with Bonferroni correction (SPSS21.0). P < 0.05 was considered statistically significant. All data are presented as mean ± SEM. Correlation analyses and curve fitting were performed using the Prism 4 program (Prism GraphPad, San Diego, CA). Figure panels were assembled with Photoshop. The data analysis of shanks CpG distribution and DNA methylation was obtained from the Generay Biotech Company. The data analysis of differential DNA methylation from blood was seen in our previously published paper [21].
Results
Dynamic Change of Shanks mRNA Expression After SE
Status epilepticus can trigger a series of reactions and processes within the brain that can precede the development of epilepsy, i.e., epitogenesis [19]. The mRNA expression of shanks1/2/3 within rat hippocampus after pilocarpine-induced SE at different time points was measured by qRT-PCR (Fig. 1 and Table 1). Compared with the control, the mRNA levels of shank1/2/3 were dramatically all downregulated 24 h after SE. Shank1 mRNA was decreased to 0.288 ± 0.163 fold change in 24 h, and returned to the basal level 7 days after SE; shank2 mRNA was decreased to 0.439 ± 0.134 fold change in 24 h, and upregulated 7 days (1.76 ± 0.618) and 14 days (1.632 ± 0.621) after SE, and returned to the basal level after 60 days; shank3 mRNA was decreased to 0.392 ± 0.206 fold change in 24 h, and upregulated 7 days (1.008 ± 0.257) and 14 days (1.366 ± 0.326) after SE, and returned to the basal level 60 days after SE. The qPCR results suggest that pilocarpine-induced SE triggered dynamic change of shanks mRNA, which might participate in epileptogenesis. Complex factors might be implicated in regulating shanks gene expression. Herein, whether shanks gene expression was regulated by DNA methylation epigenetically was investigated.
DNA Methylation Analysis of Shanks Gene After SE
Distribution of CpG Islands
Aberrant DNA methylation contributes to the pathogenesis of epilepsy, and CpG island is the common site for DNA methylation. Therefore, we examined the methylation profile of shank 1/2/3 in rat hippocampus tissue. First, CpG island of shanks gene was analyzed by CpG Island Finder Software (Table 2). CpG island was detected in the shanks gene within 10 kb upstream the promoter, and CpG island was identified within the red box. Three CpG islands of shank1 were identified containing 72 CpG sites, out of which 18 CpG sites were located in the upstream region of the transcription start site (TSS), and 54 CpG sites downstream the TSS. One CpG island of shank2 was identified containing 13 CpG sites, all of which are located in the upstream region of the TSS. Two CpG islands of shank3 were identified containing 83 CpG sites, out of which 24 CpG sites are located in the upstream of the TSS, while 59 sites are located in the downstream region of the TSS. The CpG distribution of shanks gene in human brain tissue was also analyzed, which exhibited pretty similar distribution (Supplemental Figs. 1, 2, 3).
Identification of Differentially Methylated CpG Sites at Various Time Points
Next, the CpG methylation of shank1/2/3 after SE at different time points was measured with methyl target technique and bioinformatics analysis (Figs. 2, 3; Tables 3, 4). Dynamic change of CpG methylation was identified in the shanks gene. Those CpG sites that were differentially methylated at more than one time point were considered to be of more importance. Eleven CpG sites of shank1 exhibited methylation change (Table 3, Fig. 2), out of which one CpG site (Chr1: 100297972) displayed differential methylation at two time points with hypermethylation 7 days after SE and hypomethylation 30 days after. None of the CpG sites of shank2 exhibited significant methylation change at all time points. Seventeen CpG sites of shank3 exhibited distinct methylation (Table 4, Fig. 3), out of which two CpG sites (Chr7: 130473419, and Chr7: 130473405) displayed differential methylation at more than one time point. CpG site Chr7: 130473419 showed hypomethylation 1 day after SE, with hypermethylation 14 and 60 days after, while CpG site Chr7: 130473405 exhibited hypomethylation 1 day after SE and hypermethylation 30 days after. CpG site (Chr1: 100297972) of shank1 is located downstream the transcription start site (TSS), while both the two CpG sites of shank3 (Chr7: 130473419, and Chr7: 130473405) are located upstream the TSS. Based on the location of the CpG sites, the two CpG sites of shank3 are located upstream the TSS might influence the shank3 expression via CpG methylation. Next, we analyzed the relationship between DNA methylation and shanks gene expression.
The distribution of differentially methylated CpG sites of shank1. TSS, transcription start site. Distance2TSS, the relative distance of the CpG site compared with TSS. The red dot, the number of CpG sites that displayed hypermethylation compared with the control group. The blue dot, the number of CpG sites that displayed hypomethylation compared with the control group (P < 0.05, t test)
The distribution of differentially methylated CpG sites of shank3. TSS, transcription start site. Distance2TSS, the relative distance of the CpG site compared with TSS. The red dot, the number of CpG sites that displayed hypermethylation compared with the control group. The blue dot, the number of CpG sites that displayed hypomethylation compared with the control group (P < 0.05, t test)
The Dynamic Balance Between mRNA Expression and CpG Methylation of Shanks Gene
Shanks mRNA displays dynamic change after SE. Generally, shanks mRNA exhibited significant decrease at acute stage, which progressively increased to a level higher than the control in the latent stage, and finally returned to the basal level in the chronic stage. The dynamic change of shanks mRNA was accurately regulated by complex factors including drug, vigorous seizure attack, transcription factors, and epigenetic control such as DNA methylation, environment, etc. In the acute stage 1 day after SE, the downregulation of shanks gene might possibly be induced by pilocarpine-induced SE, meanwhile DNA methylation might exert little influence on shanks expression. In the following stage, whether shanks mRNA expression was adjusted by DNA methylation was analyzed (Fig. 4a). We focus on those CpG sites that display methylation change at more than time point, including 1 CpG site of shank 1 (Chr1: 100297972) and 2 CpG sites of shank 3 (Chr7: 130473419 and Chr7: 130473405). DNA methylation is known to induce a silencing effect on gene expression, and we analyzed the relationship between DNA methylation and gene expression.
The DNA methylation of CpG site Chr1: 100297972 in shank1 was increased (2.29) 7 days after SE and decreased (0.59) 30 days after SE (Fig. 4a), with shank1 mRNA returned to the normal level 7 days and 30 days after SE; no specific relationship was exhibited between CpG methylation and shank1 mRNA. Since this CpG site is located downstream the TSS, the CpG methylation might exert little influence on the shank1 expression.
At CpG Chr7: 130473419 of shank3 (Fig. 4b), hypomethylation (0.78, P < 0.05) was exhibited 1 day after SE with mRNA downregulation (0.245, P < 0.01), followed by hypermethylation (1.74, P < 0.05) 14 days after SE with mRNA upregulation (1.366, P < 0.01), and hypermethylation (1.64, P < 0.05) 60 days later with increased mRNA (1.162, P > 0.05) (Fig. 4b). Generally, synchronized change was found between shank3 mRNA and CpG methylation, and the methylation change at Chr7: 130473419 lags behind the mRNA change. We speculate that the expression of shank3 mRNA was first downregulated by pilocarpine-induced SE, and the subsequent demethylation was triggered by the decreased shank3 for self-balance as a self-negative control to upregulate shank3. When the shank3 mRNA expression reaches a level higher than the control group, the body then tries to bring down the shank3 by hypermethylation for internal homeostasis, which explains why the DNA methylation lags behind the mRNA change.
At CpG Chr7: 130473405 (Fig. 4c) of shank3, hypomethylation (0.64, P < 0.05) was exhibited 1 day after SE with downregulated mRNA (0.245, P < 0.01), and hypermethylation (1.50, P < 0.05) 30 days later with upregulated mRNA (1.282, P > 0.05), no methylation change 60 days after SE, and mRNA returned to the basal level. This also indicates the synchronized change between mRNA and CpG methylation, and the methylation change at Chr7: 130473405 lags behind the mRNA change. The dynamic change between the mRNA and CpG methylation at this CpG site could also have resulted from a self-negative control as that was exhibited at CpG Chr7: 130473419 of shank3. Since both of the two CpG sites are located upstream the TSS, we speculate that both CpG Chr7: 130473405 and CpG Chr7: 130473419 contribute to the shank3 mRNA expression in pilocarpine-induced SE in rats.
mRNA Expression and DNA Methylation Profiling of Brain and Blood Tissue from Epileptic Patients
Brain tissue from 6 epileptic patients and 6 controls was collected for mRNA sequencing, which demonstrate that the mRNA expression of shank2 (2.1 times lower) was downregulated, while shank1 (5.7 times higher) and shank3 (2.9 times higher) were upregulated (Fig. 5). However, no statistical significance was found, which might result from the relatively small number of patients.
The mRNA expression of shanks gene from epilepsy patients. The epileptogenic zones were determined with a combination of ictal semiology, brain imaging (magnetic resonance imaging, MRI, or positron emission tomography–computed tomography also known as PET-CT), video-EEG, sphenoidal electrode monitoring, and intraoperative electrocorticography (EcoG) considered (a). The shanks gene mRNA expression from the brain tissue was measured by mRNA sequencing. Brain tissue from 6 epileptic patients and 6 controls was collected, which reveals that the mRNA expression of shank2 (2.1 times lower) (c and d) was downregulated, while shank1 (5.7 times higher) (b) and shank3 (2.9 times higher) (e) were upregulated. Data are shown as mean + SEM (P > 0.05, t test)
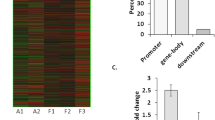
Brain tissue from 9 patients and 7 controls was analyzed for genome-wide DNA methylation profiling. Generally, 35 CpG sites were differentially methylated, out of which 20 CpG sites exhibited hypermethylation (Table 5). All three CpG sites of shank1 were hypomethylated compared with the control, which is consistent with the increased shank1 mRNA. Thirty CpG sites of shank2 were found differentially methylated, out of which 18 were hypermethylated and 12 were demethylated. Generally, shank2 DNA is hypermethylated, which is consistent with the decreased shank2 mRNA. Both the two CpG sites of shank3 were hypermethylated. And this is not in accord with the increased shank1 mRNA.
Blood samples from epilepsy patients were collected from 48 patients and 48 controls for DNA methylation sequencing (Fig. 6). Generally, 101 differentially methylated CpG sites were defined between epilepsy patients and the control, out of which 78 CpG were hypermethylated and 23 sites were demethylated. Most of the CpG sites were from shank2 (80 CpG sites), out of which 58 sites were hypermethylated and 22 were demethylated, while almost all CpG sites from shank1 (11 CpG sites) and shank3 (10 CpG sites) were hypermethylated (Table 6).
A comparison of the differentially methylated CpG sites between brain tissue and blood sample was conducted, which reveals that 3 CpG sites were found to be overlapped, including cg00001136, cg00369590, and cg00670575, all of which belong to shank2 and were hypermethylated.
In general, shank1/2/3 DNA were hypermethylated in epilepsy patients both in the brain and blood samples. This finding is consistent with our data that hypermethylation of shank1/2/3 was detected in rat hippocampus 1 day after SE at acute stage. These data suggest that the DNA hypermethylation might participate in epileptogenesis in human patients.
Discussion
As a crucial regulator of synaptic structure and function, the shanks gene has been implicated in various neurological disorders including autism spectrum disorder [10], Phelan-McDermid syndrome [11], and schizophrenia [12]. However, as far as the literature we searched, the expression and function of shanks gene in epilepsy has not been reported yet. Our study was designated to evaluate whether shanks gene was involved in the pathophysiology of epileptic seizure. For the first time to date, we investigated the dynamic change of shank1/2/3 mRNA expression and DNA methylation in both epileptic rats and epilepsy patients.
SHANK proteins are encoded by shank1, shank2, and shank3. The qRT-PCR data (Fig. 1) revealed the dynamic change of shank1/2/3 mRNA expression after SE, all of which in general decreased at acute stage, gradually upregulated in latent stage, and returned to the basal level at chronic stage. The dynamic changes of shanks gene were regulated by complex factors including drug, transcription factors, and epigenetic control such as DNA methylation, miRNA [22], environment, etc. Our study focuses on the role of DNA methylation in shanks mRNA expression. With the methyl target technique and bioinformatics analysis, dynamic methylation change was detected for shank1 and shank3, while no CpG methylation change was of statistical significance for shank2. In general, hypomethylation of shank1/3 was exhibited at acute stage with small variations, and hypermethylation gradually began to occur and became dominant at latent stage, while at chronic stage, DNA methylation exhibited relatively big variation. Classically, DNA hypermethylation leads to the silencing of the certain gene, which contradicts with our data in the animal model. We speculate that at acute stage, the inhibition of shank1/3 mRNA might be induced by pilocarpine-induced status epilepticus. The significant decrease of shanks gene might trigger the reaction of the body for self-balance, which upregulates shanks gene expression through a series of reactions including transcription factors, posttranscriptional modification, non-coding RNA regulation, etc. Among those regulation factors, DNA methylation of shank1/3 plays a crucial role and restored the shank1/3 mRNA expression via demethylation; at latent stage, upregulated shank1/3 again leads to CpG hypermethylation, which brings shank1/3 expression back to the basal level and maintains the internal homeostasis. The dynamic change of the mRNA expression of shank1/2/3 after SE implicates the possible role of shanks gene in epileptic seizure, participating in the acute stage of SE via downregulation of the mRNA and returns to the basal level by DNA methylation change as self-feedback control.
DNA hypermethylation often leads to the silencing of a certain gene. Shanks gene methylation was found to be of dynamic change; only three CpG sites were found to be differentially methylated at least at two time points, including one CpG site of shank1 (Chr1: 100297972) and two CpG sites of shank3 (Chr7: 130473419 and Chr7: 130473405). Based on our data, the expression of shank3 might possibly be regulated by DNA methylation at both Chr7: 130473419 and CpG Chr7: 130473405 in a pilocarpine-induced epilepsy model of rat. A similar result was found in the hippocampal neuron culture model, which demonstrates that the expression of shank3, but not shank1 and shank2, was regulated by epigenetic mechanisms such as DNA methylation [17].
We also extended the discovery from the animal model to epilepsy patients. Brain tissue and blood samples were collected from epilepsy patients for mRNA and genome-wide DNA methylation profiling. Based on the RNA-seq in the brain tissue, shank2 was significantly inhibited (2.3 times lower) compared with the control, while both shank1 (5.7 timer higher) and shank3 (2.9 times higher) were increased. Through a comparison, in the rat SE model, shank1/2/3 were all decreased at acute stage, and returned to the basal level at chronic stage. The difference of shank1/2/3 mRNA expression between rat and human might be explained by the different seizure types, the time when tissue was collected, frequency of seizure attack, etc. In the rat model, hippocampus tissue was dissected soon after one episode of epilepticus status, while in human patients, most of the patients had experienced years of epilepsy with various types of seizure attack other than SE. Therefore, the mRNA expression in the human brain provided more valuable information on shank gene profiling in human patients.
Furthermore, whether shank gene expression was regulated by DNA methylation in epilepsy patients was investigated. The genome-wide DNA methylation profiling reveals 35 CpG sites from the brain tissue were differentially methylated, out of which 30 CpG sites were from shank2, 3 from shank1, and 2 from shank3. And 101 CpG sites from the blood were differentially methylated, out of which 80 CpG sites were from shank2, 11 from shank1, and 10 from shank3. For shank2 in general, 60% (18 out of 30) CpG sites from the brain tissue and 72.5% (58 out of 80) CpG sites from the blood sample exhibited hypermethylation, which is consistent with the decreased shank2 mRNA from the brain tissue. Meanwhile, no specific relationship between the mRNA expression and DNA methylation was detected in shank1 or shank3. Since the number of differentially methylated CpG sites of shank1 and shank3 was relatively small, the data may not be statistically significant. Taken together, in epilepsy patients, shank2 expression might possibly be regulated by DNA methylation, while the expression of shank1 and shank3 is to be examined.
SHANKS protein has been investigated in neurological disorders such as autism spectrum disorders, Alzheimer’s disease, schizophrenia, Phelan-McDermid syndrome, and bipolar disorder. However, the function of SHANKS protein in epilepsy has not been investigated yet. Studies demonstrate that the deletion of shanks gene in mice causes synaptic plasticity defects [9, 10, 23]. Since it is convincing that synaptic plasticity was playing an important role in epilepsy, together with the dynamic change of shank mRNA in SE rat in our data, we propose that shanks gene was involved in the pathophysiology of epilepsy. The SHANK family of scaffold proteins contains multiple domains, including a PSD-95-discs large-zonula occludens-1 (PDZ) domain, a proline-rich region, a sterile alpha motif (SAM) domain, ankyrin repeats (ANK), and the SRC Homology (SH3) domain [10]. Through these domains, SHANK proteins interact with multiple synaptic proteins, such as glutamatergic receptors, cytoskeletal proteins, and other scaffold molecules [10, 12]. The deletion of shank genes in mice alters the PSD levels of several proteins including Homer, SAPAP proteins, NMDARs, and AMPARs, leading to impaired social behaviors, self-grooming, and impaired memory: all of which could be found in epilepsy patients [9, 10, 23]. The PSD protein 95-discs large homolog 1-zonula occludens-1 (PDZ) domain of SHANK 1/2/3 interacts with SAP90/PSD95-associated proteins (SAPAPs) which bind to PSD95 that interacts with NMDA receptors (NMDARs), the mutation of which has been reported to cause epileptic seizure [24, 25] that was treatable with NMDAR antagonism [26]. The class I PDZ domain of shank3 interacts with glutamate receptor 1 (GluR1) subunit of AMPA receptors (AMPARs) [23], the inhibition of which has been demonstrated to induce a strong inhibitory effect on the excitatory neurotransmission in rat hippocampus and provide seizure control [27, 28]. The deletion of shank3 alters the PSD levels of several proteins including AMPARs and NMDARs [29, 30], leading to impaired AMPA transmission, and finally causes synaptic plasticity defects [30, 31]. Taken together with the dynamic change of shanks mRNA and methylation in SE rats, we speculate that in the rat epilepsy model the shanks mRNA was regulated by NDA methylation epigenetically, and the dynamic balance between shank3 mRNA and DNA might possibly contribute to the pathophysiology of epilepsy.
Our work examined the mRNA expression and DNA methylation of shanks gene in both rats and epilepsy patients, and propose the possible role of shanks gene in the pathophysiology of epilepsy; however, the function of shanks gene is yet to be investigated. Moreover, further work is to be done to elucidate the difference of shanks mRNA expression and DNA methylation between rats and human, and to validate whether shanks gene expression is regulated by DNA methylation.
As far as the literature we searched, our work is the first to examine the dynamic change of the shank1/2/3 gene at both mRNA and DNA methylation levels in both rat and human epilepsy models. Our data presents that the downregulation of shanks gene might possibly participate in the acute stage of status epilepticus, and the dynamic balance between the mRNA expression and DNA methylation contributes to the pathophysiology of epilepsy.
References
Keller R, Basta R, Salerno L, Elia M (2017) Autism, epilepsy, and synaptopathies: a not rare association. Neurol Sci 38:1353–1361
Krishnan V, Stoppel DC, Nong Y, Johnson MA, Nadler MJS, Ozkaynak E, Teng BL, Nagakura I et al (2017) Autism gene Ube3a and seizures impair sociability by repressing VTA Cbln1. Nature 543:507–512
Abeliovich A, Gitler AD (2016) Defects in trafficking bridge Parkinson’s disease pathology and genetics. Nature 539:207–216
Lourenco MV, Frozza RL, de Freitas GB, Zhang H, Kincheski GC, Ribeiro FC, Gonçalves RA, Clarke JR et al (2019) Exercise-linked FNDC5/irisin rescues synaptic plasticity and memory defects in Alzheimer’s models. Nat Med 25:165–175
Karayannis T, Au E, Patel JC, Kruglikov I, Markx S, Delorme R, Héron D, Salomon D et al (2014) Cntnap4 differentially contributes to GABAergic and dopaminergic synaptic transmission. Nature 511:236–240
Soltesz I, Alger BE, Kano M, Lee SH, Lovinger DM, Ohno-Shosaku T, Watanabe M (2015) Weeding out bad waves: towards selective cannabinoid circuit control in epilepsy. Nat Rev Neurosci 16:264–277
Khlebodarova TM, Kogai VV, Trifonova EA, Likhoshvai VA (2018) Dynamic landscape of the local translation at activated synapses. Mol Psychiatry 23:107–114
Shin SM, Zhang N, Hansen J, Gerges NZ, Pak DTS, Sheng M, Lee SH (2012) GKAP orchestrates activity-dependent postsynaptic protein remodeling and homeostatic scaling. Nat Neurosci 15:1655–1666
Yuan T, Mameli M, O'Connor EC, Dey PN, Verpelli C, Sala C, Perez-Otano I, Lüscher C et al (2013) Expression of cocaine-evoked synaptic plasticity by GluN3A-containing NMDA receptors. Neuron 80:1025–1038
Monteiro P, Feng G (2017) SHANK proteins: roles at the synapse and in autism spectrum disorder. Nat Rev Neurosci 18:147–157
Berg EL, Copping NA, Rivera JK, Pride MC, Careaga M, Bauman MD, Berman RF, Lein PJ et al (2018) Developmental social communication deficits in the Shank3 rat model of Phelan-McDermid syndrome and autism spectrum disorder. Autism Res 11:587–601
de Sena Cortabitarte A, Degenhardt F, Strohmaier J, Lang M, Weiss B, Roeth R, Giegling I, Heilmann-Heimbach S et al (2017) Investigation of SHANK3 in schizophrenia. Am J Med Genet Part B Neuropsychiatr Genet 174:390–398
Machnes ZM, Huang TCT, Chang PKY, Gill R, Reist N, Dezsi G, Ozturk E, Charron F et al (2013) DNA methylation mediates persistent epileptiform activity in vitro and in vivo. PLoS One 8:e76299
Kobow K, Jeske I, Hildebrandt M, Hauke J, Hahnen E, Buslei R, Buchfelder M, Weigel D et al (2009) Increased reelin promoter methylation is associated with granule cell dispersion in human temporal lobe epilepsy. J Neuropathol Exp Neurol 68:356–364
Kobow K, Blümcke I (2011) The methylation hypothesis: do epigenetic chromatin modifications play a role in epileptogenesis? Epilepsia 52(Suppl 4):15–19
Campbell RR, Wood MA (2019) How the epigenome integrates information and reshapes the synapse. Nat Rev Neurosci 20:133–147
Beri S, Tonna N, Menozzi G, Bonaglia MC, Sala C, Giorda R (2007) DNA methylation regulates tissue-specific expression of Shank3. J Neurochem 101:1380–1391
Luo ZH, Walid A A, Xie Y, Long H, Xiao W, Xu L, Fu Y, Feng L et al (2019) Construction and analysis of a dysregulated lncRNA-associated ceRNA network in a rat model of temporal lobe epilepsy. Seizure 69:105–114
Fu Y, Yang MS, Jiang J, Ganesh T, Joe E, Dingledine R (2015) EP2 receptor signaling regulates microglia death. Mol Pharmacol 88:161–170
Xiao W, Cao Y, Long H, Luo Z, Li S, Deng N, Wang J, Lu X et al (2018) Genome-wide DNA methylation patterns analysis of noncoding RNAs in temporal lobe epilepsy patients. Mol Neurobiol 55:793–803
Long H-Y, Feng L, Kang J, Luo ZH, Xiao WB, Long LL, Yan XX, Zhou L et al (2017) Blood DNA methylation pattern is altered in mesial temporal lobe epilepsy. Sci Rep 7:43810
Xiao W, Wu Y, Wang J, Luo Z, Long L, Deng N, Ning S, Zeng Y et al (2019) Network and pathway-based analysis of single-nucleotide polymorphism of miRNA in temporal lobe epilepsy. Mol Neurobiol 56:7022–7031
Uchino S, Wada H, Honda S, Nakamura Y, Ondo Y, Uchiyama T, Tsutsumi M, Suzuki E et al (2006) Direct interaction of post-synaptic density-95/Dlg/ZO-1 domain-containing synaptic molecule Shank3 with GluR1 alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid receptor. J Neurochem 97:1203–1214
Marwick K, Skehel P, Hardingham G, Wyllie D (2015) Effect of a GRIN2A de novo mutation associated with epilepsy and intellectual disability on NMDA receptor currents and Mg(2+) block in cultured primary cortical neurons. Lancet Lond Engl 385(Suppl 1):S65
Li D, Yuan H, Ortiz-Gonzalez XR, Marsh ED, Tian L, McCormick EM, Kosobucki GJ, Chen W et al (2016) GRIN2D recurrent de novo dominant mutation causes a severe epileptic encephalopathy treatable with NMDA receptor channel blockers. Am J Hum Genet 99:802–816
Rothan HA, Amini E, Faraj FL, Golpich M, Teoh TC, Gholami K, Yusof R (2017) NMDA receptor antagonism with novel indolyl, 2-(1,1-dimethyl-1,3-dihydro-benzo[e]indol-2-ylidene)-malonaldehyde, reduces seizures duration in a rat model of epilepsy. Sci Rep 7:45540
Chang P, Augustin K, Boddum K, Williams S, Sun M, Terschak JA, Hardege JD, Chen PE et al (2016) Seizure control by decanoic acid through direct AMPA receptor inhibition. Brain J Neurol 139:431–443
Yelshanskaya MV, Singh AK, Sampson JM, Narangoda C, Kurnikova M, Sobolevsky AI (2016) Structural bases of noncompetitive inhibition of AMPA-subtype Ionotropic glutamate receptors by antiepileptic drugs. Neuron 91:1305–1315
Peça J, Feliciano C, Ting JT, Wang W, Wells MF, Venkatraman TN, Lascola CD, Fu Z et al (2011) Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472:437–442
Mei Y, Monteiro P, Zhou Y, Kim JA, Gao X, Fu Z, Feng G (2016) Adult restoration of Shank3 expression rescues selective autistic-like phenotypes. Nature 530:481–484
Zhou Y, Kaiser T, Monteiro P, Zhang X, van der Goes MS, Wang D, Barak B, Zeng M et al (2016) Mice with Shank3 mutations associated with ASD and schizophrenia display both shared and distinct defects. Neuron 89:147–162
Funding
This work was supported through funding from the National Natural Science Foundation of China (grant numbers 81671299 to B.X., and 81701182 to Y.F.) and the Natural Science Foundation of Hunan Province (grant number: 2018JJ2648) to Z.L.
Author information
Authors and Affiliations
Contributions
Du Liu and Zhaohui Luo conducted the majority of the experiments. Yujiao Fu analyzed and interpreted the data and drafted the manuscript. Jialing Guo and Wei Xiao prepared the epilepsy animal experiment. Du Liu and Zhaohui Luo prepared the CpG detective experiments. Hongyu Long and Wenbiao Xiao analyzed and interpreted the data. Li Feng prepared for language editing and revising. Bo Xiao and Zhaohui Luo contributed to the study design. Bo Xiao provided financial support. All authors consent to the publication of the manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethics Approval
All procedures were followed in accordance with the ethical standards of the responsible committee on animal and human experimentation (the Ethics Review Committee of Xiangya Hospital, numbers 201603296 and 201603297) and with the Helsinki Declaration of 1964 and later versions.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Fu, Y., Liu, D., Guo, J. et al. Dynamic Change of Shanks Gene mRNA Expression and DNA Methylation in Epileptic Rat Model and Human Patients. Mol Neurobiol 57, 3712–3726 (2020). https://doi.org/10.1007/s12035-020-01968-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-020-01968-5