Abstract
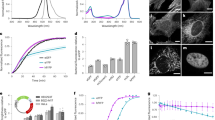
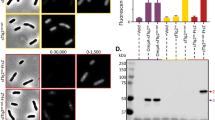
Directed evolution-based protein engineering usually generates large library contained insoluble mutants because of structural disturbance by mutation. To reduce the workload and costs, it is crucial to identify and eliminate those insoluble variants prior to dedicated analysis. Here, we demonstrate a method to visualize soluble protein mutants by using monomeric red fluorescent protein (mRFP) as a fusion tag. A plasmid was devised to express nicotinic acid mononucleotide adenylyltransferase (NadD) fused with a GGGS-linked mRFP tag at the C-terminus. The plasmid was subjected to site saturation mutagenesis within the nadD gene, used to transform Escherichia coli DH10B competent cells, leading to colonies with different red intensities. It was found that the fluorescence intensity of the cell culture correlated positively with the content of NadD-mRFP mutant in the supernatant. Mutation at position 132 led to a library of which most colonies lost the red phenotype, indicating that the position had a key role for proper protein folding. Similarly, mRFP enabled identification of soluble mutants of other enzymes including 1-deoxy-D-xylulose-5-phosphate reductoisomerase and phosphite dehydrogenase. These data suggested that mRFP can serve as a fusion reporter for visualizing soluble protein mutants to facilitate more efficient library screening in directed evolution.






Similar content being viewed by others
References
Dougherty, M. J., & Arnold, F. H. (2009). Directed evolution: new parts and optimized function. Current Opinion in Biotechnology, 20(4), 486–491.
Waldo, G. S. (2003). Genetic screens and directed evolution for protein solubility. Current Opinion in Chemical Biology, 7(1), 33–38.
Zhang, J., Campbell, R. E., Ting, A. Y., & Tsien, R. Y. (2002). Creating new fluorescent probes for cell biology. Nature Reviews. Molecular Cell Biology, 3(12), 906–918.
Waldo, G. S., Standish, B. M., Berendzen, J., & Terwilliger, T. C. (1999). Rapid protein-folding assay using green fluorescent protein. Nature Biotechnology, 17(7), 691–695.
Listwan, P., Terwilliger, T. C., & Waldo, G. S. (2009). Automated, high-throughput platform for protein solubility screening using a split-GFP system. Journal of Structural and Functional Genomics, 10(1), 47–55.
Pedelacq, J. D., Cabantous, S., Tran, T., Terwilliger, T. C., & Waldo, G. S. (2006). Engineering and characterization of a superfolder green fluorescent protein. Nature Biotechnology, 24(1), 79–88.
Heddle, C., & Mazaleyrat, S. L. (2007). Development of a screening platform for directed evolution using the reef coral fluorescent protein ZsGreen as a solubility reporter. Protein Engineering, Design & Selection, 20(7), 327–337.
Shaner, N. C., Steinbach, P. A., & Tsien, R. Y. (2005). A guide to choosing fluorescent proteins. Nature Methods, 2(12), 905–909.
Zhong, C., Wei, P., & Zhang, Y.-H. P. (2017). Enhancing functional expression of codon-optimized heterologous enzymes in Escherichia coli BL21(DE3) by selective introduction of synonymous rare codons. Biotechonology and Bioengineering, 114(5), 1054–1064.
Chen, P., Feng, X., Hu, R., Sun, J., Du, W., & Liu, B. F. (2010). Hydrodynamic gating valve for microfluidic fluorescence-activated cell sorting. Analytica Chimica Acta, 663, 1–6.
Cheong, D. E., Ko, K. C., Han, Y., Jeon, H. G., Sung, B. H., Kim, G.-J., Choi, J. H., & Song, J. J. (2015). Enhancing functional expression of heterologous proteins through random substitution of genetic codes in the 5′-coding region. Biotechonology and Bioengineering, 112(4), 822–826.
Lu, P., & Feng, M. G. (2008). Bifunctional enhancement of a beta-glucanase-xylanase fusion enzyme by optimization of peptide linkers. Applied Microbiology and Biotechnology, 79(4), 579–587.
Campbell, R. E., Tour, O., Palmer, A. E., Steinbach, P. A., Baird, G. S., Zacharias, D. A., & Tsien, R. Y. (2002). A monomeric red fluorescent protein. Proceedings of the National Academy of Sciences of the United States of America, 99(12), 7877–7882.
Mehl, R. A., Kinsland, C., & Begley, T. P. (2000). Identification of the Escherichia coli nicotinic acid mononucleotide adenylyltransferase gene. Journal of Bacteriology, 182(15), 4372–4374.
Wang, L., Ji, D., Liu, Y., Wang, Q., Wang, X., Zhou, Y. J., Zhang, Y., Liu, W., & Zhao, Z. K. (2017). Synthetic cofactor-linked metabolic circuits for selective energy transfer. ACS Catalysis, 7(3), 1977–1983.
Wang, J., Tan, H., & Zhao, Z. K. (2007). Over-expression, purification, and characterization of recombinant NAD-malic enzyme from Escherichia coli K12. Protein Expression and Purification, 53(1), 97–103.
van den Ent, F., & Lowe, J. (2006). RF cloning: a restriction-free method for inserting target genes into plasmids. Journal of Biochemical and Biophysical Methods, 67(1), 67–74.
Wang, J., Zhang, S., Tan, H., & Zhao, Z. (2007). PCR-based strategy for construction of multi-site-saturation mutagenic expression library. Journal of Microbiological Methods, 71(3), 225–230.
Wang, X., Zhou, Y. J., Wang, L., Liu, W., Liu, Y., Peng, C., & Zhao, Z. K. (2017). Engineering Escherichia coli nicotinic acid mononucleotide adenylyltransferase for fully active amidated NAD biosynthesis. Applied and Environmental Microbiology, 83(13), e00692–e00617.
Zhang, L., Patel, H. N., Lappe, J. W., & Wachter, R. M. (2006). Reaction progress of chromophore biogenesis in green fluorescent protein. Journal of the American Chemical Society, 128, 4766–4772.
Cha, H. J., Wu, C.-F., Valdes, J. J., Rao, G., & Bentley, W. E. (2000). Observations of green fluorescent protein as a fusion partner in genetically engineered Escherichia coli: monitoring protein expression and solubility. Biotechnology and Bioengineering, 67(5), 565–574.
Seitz, T., Thoma, R., Schoch, G. A., Stihle, M., Benz, J., D'Arcy, B., Wiget, A., Ruf, A., Hennig, M., & Sterner, R. (2010). Enhancing the stability and solubility of the glucocorticoid receptor ligand-binding domain by high-throughput library screening. Journal of Molecular Biology, 403, 562–577.
Gupta, R. D., & Tawfik, D. S. (2008). Directed enzyme evolution via small and effective neutral drift libraries. Nature Methods, 5(11), 939–942.
Jiang, S., Li, C., Zhang, W., Cai, Y., Yang, Y., Yang, S., & Jiang, W. (2007). Directed evolution and structural analysis of N-carbamoyl-D-amino acid amidohydrolase provide insights into recombinant protein solubility in Escherichia coli. The Biochemical Journal, 402, 429–437.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant numbers: 21325627, 21572227) and Dalian Institute of Chemical Physics, CAS (grant number: DICP ZZBS201605).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that there is no conflict of interest.
Electronic Supplementary Material
ESM 1
(DOC 2215 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Wang, L., Lin, X. et al. Visualizing Soluble Protein Mutants by Using Monomeric Red Fluorescent Protein as a Reporter for Directed Evolution. Appl Biochem Biotechnol 185, 81–90 (2018). https://doi.org/10.1007/s12010-017-2640-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-017-2640-z