Abstract
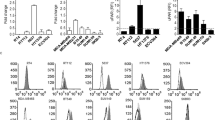
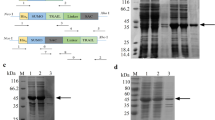
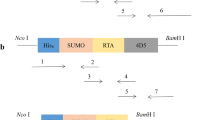
Prostate apoptosis response-4 (Par-4), an anticancer protein that interacts with cell surface receptor GRP78, can selectively suppress proliferation and induce apoptosis of cancer cells. The core domain of Par-4 (aa 137–195), designated as SAC, is sufficient to inhibit tumor growth and metastasis without harming normal tissues and organs. Nevertheless, the anticancer effects of SAC have not been determined in ovarian cancer cells. Here, we developed a novel method for producing native SAC in Escherichia coli using a small ubiquitin-related modifier (SUMO) fusion system. This fusion system not only greatly improved the solubility of target protein but also enhanced the expression level of SUMO-SAC. After purified by Ni-NTA affinity chromatography, SUMO tag was cleaved from SUMO-SAC fusion protein using SUMO protease to obtain recombinant SAC. Furthermore, we simplified the purification process by combining the SUMO-SAC purification and SUMO tag cleavage into one step. Finally, the purity of recombinant SAC reached as high as 95% and the yield was 25 mg/L. Our results demonstrated that recombinant SAC strongly inhibited proliferation and induced apoptosis in ovarian cancer cells SKOV-3. Immunofluorescence analysis and competitive binding reaction showed that recombinant SAC could specifically induce apoptosis of SKOV-3 cells through combination with cell surface receptor, GRP78. Therefore, we have developed an effective strategy for expressing bioactive SAC in prokaryotic cells, which supports the application of SAC in ovarian cancer therapy.






Similar content being viewed by others
References
Hebbar, N., Wang, C., & Rangnekar, V. M. (2012). Mechanisms of apoptosis by the tumor suppressor Par-4. Journal of Cellular Physiology, 227, 3715–3721.
El-Guendy, N., & Rangnekar, V. M. (2003). Apoptosis by Par-4 in cancer and neurodegenerative diseases. Experimental Cell Research, 283, 51–66.
El-Guendy, N., Zhao, Y., Gurumurthy, S., Burikhanov, R., & Rangnekar, V. M. (2003). Identification of a unique core domain of par-4 sufficient for selective apoptosis induction in cancer cells. Molecular and Cellular Biology, 23, 5516–5525.
Zhao, Y., & Rangnekar, V. M. (2008). Apoptosis and tumor resistance conferred by Par-4. Cancer Biology & Therapy, 7, 1867–1874.
Burikhanov, R., Zhao, Y., Goswami, A., Qiu, S., Schwarze, S. R., & Rangnekar, V. M. (2009). The tumor suppressor Par-4 activates an extrinsic pathway for apoptosis. Cell, 138, 377–388.
Zhao, Y., Burikhanov, R., Brandon, J., Qiu, S., Shelton, B. J., Spear, B., Bondada, S., Bryson, S., & Rangnekar, V. M. (2011). Systemic Par-4 inhibits non-autochthonous tumor growth. Cancer Biology & Therapy, 12, 152–157.
Shrestha-Bhattarai, T., & Rangnekar, V. M. (2010). Cancer-selective apoptotic effects of extracellular and intracellular Par-4. Oncogene, 29, 3873–3880.
Sayers, T. J. (2011). Targeting the extrinsic apoptosis signaling pathway for cancer therapy. Cancer Immunology, Immunotherapy, 60, 1173–1180.
Chaudhry, P., Singh, M., Parent, S., & Asselin, E. (2012). Prostate apoptosis response 4 (Par-4), a novel substrate of caspase-3 during apoptosis activation. Molecular and Cellular Biology, 32, 826–839.
Braun, P., Hu, Y., Shen, B., Halleck, A., Koundinya, M., Harlow, E., & LaBaer, J. (2002). Proteome-scale purification of human proteins from bacteria. Proceedings of the National Academy of Sciences of the United States of America, 99, 2654–2659.
Zafar, A., Aftab, M. N., ud Din, Z., Aftab, S., Iqbal, I., & ul Haq, I. (2016). Cloning, purification and characterization of a highly thermostable amylase gene of Thermotoga petrophila into Escherichia coli. Applied Biochemistry and Biotechnology, 178, 831–848.
Baneyx, F. (1999). Recombinant protein expression in Escherichia coli. Current Opinion in Biotechnology, 10, 411–421.
Mahamad, P., Boonchird, C., & Panbangred, W. (2016). High level accumulation of soluble diphtheria toxin mutant (CRM197) with co-expression of chaperones in recombinant Escherichia coli. Applied Microbiology and Biotechnology, 100, 6319–6330.
Tian, H., Zhao, Y., Chen, N., Wu, M., Gong, W., Zheng, J., Fernig, D. G., Jungbauer, A., Wang, D., Li, X., & Jiang, C. (2016). High production in E. coli of biologically active recombinant human fibroblast growth factor 20 and its neuroprotective effects. Applied Microbiology and Biotechnology, 100, 3023–3034.
Rueda, F., Cano-Garrido, O., Mamat, U., Wilke, K., Seras-Franzoso, J., García-Fruitós, E., & Villaverde, A. (2014). Production of functional inclusion bodies in endotoxin-free Escherichia coli. Applied Microbiology and Biotechnology, 98, 9229–9238.
Gao, J., & Wang, H. (2015). Prokaryotic expression, refolding and purification of high-purity mouse Midkine in Escherichia coli. Applied Biochemistry and Biotechnology, 176, 454–466.
Kong, B., & Guo, G. L. (2011). Enhanced in vitro refolding of fibroblast growth factor 15 with the assistance of SUMO fusion partner. PloS One, 6, e20307.
Fan, J., Huang, L., Sun, J., Qiu, Y., Zhou, J., & Shen, Y. (2015). Strategy for linker selection to enhance refolding and bioactivity of VAS-TRAIL fusion protein based on inclusion body conformation and activity. Journal of Biotechnology, 209, 16–22.
Kapust, R. B., & Waugh, D. S. (1999). Escherichia coli maltose-binding protein is uncommonly effective at promoting the solubility of polypeptides to which it is fused. Protein Science, 8, 1668–1674.
Li, Y. (2013). Production of human antimicrobial peptide LL-37 in Escherichia coli using a thioredoxin-SUMO dual fusion system. Protein Expression and Purification, 87, 72–78.
Zhang, M., Qiu, Z., Li, Y., Yang, Y., Zhang, Q., Xiang, Q., Su, Z., & Huang, Y. (2013). Construction and characterization of a recombinant human beta defensin 2 fusion protein targeting the epidermal growth factor receptor: in vitro study. Applied Microbiology and Biotechnology, 97, 3913–3923.
Dian, C., Eshaghi, S., Urbig, T., McSweeney, S., Heijbel, A., Salbert, G., & Birse, D. (2002). Strategies for the purification and on-column cleavage of glutathione-S-transferase fusion target proteins. Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences, 769, 133–144.
Chaubey, N., & Ghosh, S. S. (2013). Molecular cloning, purification and functional implications of recombinant GST tagged hGMCSF cytokine. Applied Biochemistry and Biotechnology, 169, 1713–1726.
Ichikawa, Y., Kagawa, W., Saito, K., Chikashige, Y., Haraguchi, T., Hiraoka, Y., & Kurumizaka, H. (2013). Purification and characterization of the fission yeast telomere clustering factors, Bqt1 and Bqt2. Protein Expression and Purification, 88, 207–213.
Ye, T., Lin, Z., & Lei, H. (2008). High-level expression and characterization of an anti-VEGF165 single-chain variable fragment (scFv) by small ubiquitin-related modifier fusion in Escherichia coli. Applied Microbiology and Biotechnology, 81, 311–317.
Li, Y. (2013). Recombinant production of crab antimicrobial protein scygonadin expressed as thioredoxin and SUMO fusions in Escherichia coli. Applied Biochemistry and Biotechnology, 169, 1847–1857.
Upadhyay, S. K., Saurabh, S., Rai, P., Singh, R., Chandrashekar, K., Verma, P. C., Singh, P. K., & Tuli, R. (2010). SUMO fusion facilitates expression and purification of garlic leaf lectin but modifies some of its properties. Journal of Biotechnology, 146, 1–8.
Xu, R., Dong, Y., Wang, L., Tao, X., Sun, A., & Wei, D. (2014). TAT-RhoGDI2, a novel tumor metastasis suppressor fusion protein: expression, purification and functional evaluation. Applied Microbiology and Biotechnology, 98, 9633–9641.
Lv, X., Zhang, J., Xu, R., Dong, Y., Sun, A., Shen, Y., & Wei, D. (2016). Gigantoxin-4-4D5 scFv is a novel recombinant immunotoxin with specific toxicity against HER2/neu-positive ovarian carcinoma cells. Applied Microbiology and Biotechnology, 100, 6403–6413.
Lee, T. J., Jang, J. H., Noh, H. J., Park, E. J., Choi, K. S., & Kwon, T. K. (2010). Overexpression of Par-4 sensitizes TRAIL-induced apoptosis via inactivation of NF-κB and Akt signaling pathways in renal cancer cells. Journal of Cellular Biochemistry, 109, 885–895.
Lee, T. J., Lee, J. T., Kim, S. H., Choi, Y. H., Song, K. S., Park, J. W., & Kwon, T. K. (2008). Overexpression of Par-4 enhances thapsigargin-induced apoptosis via downregulation of XIAP and inactivation of Akt in human renal cancer cells. Journal of Cellular Biochemistry, 103, 358–368.
Jagtap, J. C., Dawood, P., Shah, R. D., Chandrika, G., Natesh, K., Shiras, A., Hegde, A. S., Ranade, D., & Shastry, P. (2014). Expression and regulation of prostate apoptosis response-4 (Par-4) in human glioma stem cells in drug-induced apoptosis. PloS One, 9, e88505.
Meynier, S., Kramer, M., Ribaux, P., Tille, J. C., Delie, F., Petignat, P., & Cohen, M. (2015). Role of PAR-4 in ovarian cancer. Oncotarget, 6, 22641–22652.
Kline, C. L., & Irby, R. B. (2011). The pro-apoptotic protein prostate apoptosis response protein-4 (Par-4) can be activated in colon cancer cells by treatment with Src inhibitor and 5-FU. Apoptosis, 16, 1285–1294.
Zhang, X. X., Li, H. D., Zhao, L., Song, H. J., Wang, G., Guo, Q. J., Luan, Z. D., & Su, R. J. (2013). The cell surface GRP78 facilitates the invasion of hepatocellular carcinoma cells. BioMed Research International, 2013, 917296.
Li, Z., Zhang, L., Zhao, Y., Li, H., Xiao, H., Fu, R., Zhao, C., Wu, H., & Li, Z. (2013). Cell-surface GRP78 facilitates colorectal cancer cell migration and invasion. The International Journal of Biochemistry & Cell Biology, 45, 987–994.
Irby, R. B., & Kline, C. L. (2013). Par-4 as a potential target for cancer therapy. Expert Opinion on Therapeutic Targets, 17, 77–87.
Funding
This work was funded by the National Natural science Foundation of China (No. 21646005/B060806) and China Postdoctoral Science Foundation funded project (No. 2016M601529).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhang, J., Sun, A., Dong, Y. et al. Recombinant Production and Characterization of SAC, the Core Domain of Par-4, by SUMO Fusion System. Appl Biochem Biotechnol 184, 1155–1167 (2018). https://doi.org/10.1007/s12010-017-2599-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-017-2599-9