Abstract
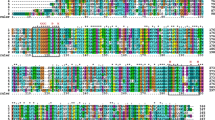
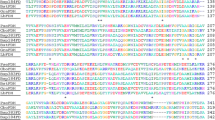
The budC gene encoding a meso-2,3-butanediol dehydrogenase (BlBDH) from Bacillus licheniformis was cloned and overexpressed in Escherichia coli BL21(DE3). Sequence analysis reveals that this BlBDH belongs to short-chain dehydrogenase/reductase (SDR) superfamily. In the presence of NADH, BlBDH catalyzes the reduction of diacetyl to (3S)-acetoin (97.3 % ee), and further to (2S,3S)-2,3-butanediol (97.3 % ee and 96.5 % de). Similar to other meso-2,3-BDHs, it shows oxidative activity to racemic 2,3-butanediol whereas no activity toward racemic acetoin in the presence of NAD+. For diacetyl reduction and 2,3-butanediol oxidation, the pH optimum of BlBDH is 5.0 and 10.0, respectively. Unusually, it shows relatively high activity over a wide pH range from 5.0 to 8.0 for racemic acetoin reduction. BlBDH shows lower K m and higher catalytic efficiency toward racemic acetoin (K m = 0.47 mM, k cat /K m = 432 s–1·mM–1) when compared with 2,3-butanediol (K m = 7.25 mM, k cat /K m = 81.5 s–1·mM–1), indicating its physiological role in favor of reducing racemic acetoin into 2,3-butanediol. The enzymatic characterization of BlBDH provides evidence for the directed engineering of B. licheniformis for producing enantiopure 2,3-butanediol.






Similar content being viewed by others
References
Li, L. X., Zhang, L. J., Li, K., Wang, Y., Gao, C., Han, B. B., Ma, C. Q., & Xu, P. (2013). A newly isolated Bacillus licheniformis strain thermophilically produces 2,3-butanediol, a platform and fuel biochemical. Biotechnology for Biofuels, 6, 123.
Ji, X. J., Huang, H., & Ouyang, P. K. (2011). Microbial 2,3-butanediol production: a state-of-the-art review. Biotechnology Advances, 29, 351–364.
Li, L. X., Wang, Y., Zhang, L. J., Ma, C. Q., Wang, A. L., Tao, F., & Xu, P. (2012). Biocatalytic production of (2S,3S)-2,3-butanediol from diacetyl using whole cells of engineered Escherichia coli. Bioresource Technology, 115, 111–116.
Qi, G. F., Kang, Y. F., Li, L., Xiao, A. F., Zhang, S. M., Wen, Z. Y., Xu, D. H., & Chen, S. W. (2014). Deletion of meso-2,3-butanediol dehydrogenase gene budC for enhanced D-2,3-butanediol production in Bacillus licheniformis. Biotechnology for Biofuels, 7, 16.
Xu, G. C., & Ni, Y. (2015). Bioreductive preparation of ACE inhibitors precursor (R)-2-hydroxy-4-phenylbutanoate esters: recent advances and future perspectives. Bioresources and Bioprocessing, 2, 15.
Ma, C. Q., Wang, A. L., Qin, J. Y., Li, L. X., Ai, X. L., Jiang, T. Y., Tang, H. Z., & Xu, P. (2009). Enhanced 2,3-butanediol production by Klebsiella pneumoniae SDM. Applied Microbiology and Biotechnology, 82, 49–57.
Zhang, L. Y., Yang, Y. L., Sun, J. A., Shen, Y. L., Wei, D. Z., Zhu, J. W., & Chu, J. (2010). Microbial production of 2,3-butanediol by a mutagenized strain of Serratia marcescens H30. Bioresource Technology, 101, 1961–1967.
Ji, X. J., Huang, H., Du, J., Zhu, J. G., Ren, L. J., Hu, N., & Li, S. (2009). Enhanced 2,3-butanediol production by Klebsiella oxytoca using a two-stage agitation speed control strategy. Bioresource Technology, 100, 3410–3414.
Wang, A., Xu, Y. Q., Ma, C., Gao, C., Li, L. X., Wang, Y., Tao, F., & Xu, P. (2012). Efficient 2,3-butanediol production from cassava powder by a crop-biomass-utilizer, Enterobacter cloacae subsp. dissolvens SDM. PLoS One, 7, e40442.
Jurchescu, I. M., Hamann, J., Zhou, X., Ortmann, T., Kuenz, A., Prüβe, U., & Lang, S. (2013). Enhanced 2,3-butanediol production in fed-batch cultures of free and immobilized Bacillus licheniformis DSM 8785. Applied Microbiology and Biotechnology, 97, 6715–6723.
Hassler, T., Schieder, D., Pfaller, R., Faulstich, M., & Sieber, V. (2012). Enhanced fed-batch fermentation of 2,3-butanediol by Paenibacillus polymyxa DSM 365. Bioresource Technology, 124, 237–244.
Biswas, R., Yamaoka, M., Nakayama, H., Kondo, T., Yoshida, K., Bisaria, V. S., & Kondo, A. (2012). Enhanced production of 2,3-butanediol by engineered Bacillus subtilis. Applied Microbiology and Biotechnology, 94, 651–658.
Yan, Y. J., Lee, C. C., & Liao, J. C. (2009). Enantioselective synthesis of pure (R, R)-2,3-butanediol in Escherichia coli with stereospecific secondary alcohol dehydrogen. Organic and Biomolecular Chemistry, 7, 3914–3917.
González, E., Fernández, M. R., Larroy, C., Solà, L., Pericàs, M. A., Parés, X., & Biosca, J. A. (2000). Characterization of a (2R,3R)-2,3-butanediol dehydrogenase as the Saccharomyces cerevisiae YAL060W gene product. Disruption and induction of the gene. Journal of Biological Chemistry, 275, 35876–35885.
Yu, B., Sun, J., Bommareddy, R. R., Song, L., & Zeng, A. P. (2011). Novel (2R,3R)-2,3-butanediol dehydrogenase from potential industrial strain Paenibacillus polymyxa ATCC 12321. Applied and Environmental Microbiology, 77, 4230–4233.
Nicholson, W. L. (2008). The Bacillus subtilis ydjL (bdhA) gene encodes acetoin reductase/2,3-butanediol dehydrogenase. Applied and Environmental Microbiology, 74, 6832–6838.
Zhang, L. Y., Xu, Q. M., Zhan, S. R., Li, Y. Y., Lin, H., Sun, S. J., Sha, L., Hu, K. H., Guan, X., & Shen, Y. L. (2014). A new NAD(H)-dependent meso-2,3-butanediol dehydrogenase from an industrially potential strain Serratia marcescens H30. Applied Microbiology and Biotechnology, 98, 1175–1184.
Giovannini, P. P., Medici, A., Bergamini, C. M., & Rippa, M. (1996). Properties of diacetyl (acetoin) reductase from Bacillus stearothermophilus. Bioorganic and Medicinal Chemistry, 4, 1197–1201.
Wang, Z., Song, Q. Q., Yu, M. L., Wang, Y. F., Xiong, B., Zhang, Y. J., Zheng, J. Y., & Ying, X. X. (2014). Characterization of a stereospecific acetoin (diacetyl) reductase from Rhodococcus erythropolis WZ010 and its application for the synthesis of (2S,3S)-2,3-butanediol. Applied Microbiology and Biotechnology, 98, 641–650.
Zhang, L. Y., Xu, Q. M., Peng, X. Q., Xu, B. H., Wu, Y. H., Yang, Y. L., Sun, S. J., Hu, K. H., & Shen, Y. L. (2014). Cloning, expression and characterization of glycerol dehydrogenase involved in 2,3-butanediol formation in Serratia marcesce. Journal of Industrial Microbiology and Biotechnology, 41, 1319–1327.
Wang, Y., Tao, F., & Xu, P. (2014). Glycerol dehydrogenase plays a dual role in glycerol metabolism and 2,3-butanediol formation in Klebsiella pneumoniae. Journal of Biological Chemistry, 289, 6080–6090.
Thanh, T. N., Jürgen, B., Bauch, M., Liebeke, M., Lalk, M., Ehrenreich, A., Evers, S., Maurer, K. H., Antelmann, H., Ernst, F., Homuth, G., Hecker, M., & Schweder, T. (2010). Regulation of acetoin and 2,3-butanediol utilization in Bacillus licheniformis. Applied Microbiology and Biotechnology, 87, 2227–2235.
Raedts, J., Siemerink, M. A. J., Levisson, M., van der Oost, J., & Kengen, S. W. M. (2014). Molecular characterization of an NADPH-dependent acetoin reductase 2,3-butanediol dehydrogenase from Clostridium beijerinckii NCIMB8052. Applied and Environmental Microbiology, 80, 2011–2020.
Xu, G. C., Yu, H. L., Zhang, X. Y., & Xu, J. H. (2012). Access to optically active aryl halohyddrins using a substrate-tolerant carbonyl reductase discovered from Kluyveromyces thermotolerans. ACS Catalysis, 2, 2566–2571.
Filling, C., Berndt, K. D., Benach, J., Knapp, S., Prozorovski, T., Nordling, E., Ladenstein, R., Jörnvall, H., & Oppermann, U. (2002). Critical residues for structure and catalysis in short-chain dehydrogenases/reductases. Journal of Biological Chemistry, 277, 25677–25684.
Jörnvall, H., Hedlund, J., Bergman, T., Oppermann, U., & Persson, B. (2010). Superfamilies SDR and MDR: from early ancestry to present forms. Emergence of three lines, a Zn-metalloenzyme, and distinct variabilities. Biochemical and Biophysical Research Communications, 396, 125–130.
Acknowledgments
We are grateful to the National Natural Science Foundation of China (21276112 and 21506073), Natural Science Foundation of Jiangsu Province (BK20150003), the Fundamental Research Funds for the Central Universities (JUSRP51409B), the Program of Introducing Talents of Discipline to Universities (111-2-06), and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions for the financial support of this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
All the authors certify that this manuscript is original and has not been published and will not be submitted elsewhere for publication while being considered by Applied Biochemistry and Biotechnology. Moreover, the study is not split up into several parts to increase the quantity of submissions and submitted to various journals or to one journal over time. No data have been fabricated or manipulated (including images) to support your conclusions. No data, text, or theories by others are presented as if they were our own. The submission has been received explicitly from all coauthors. Furthermore, authors whose names appear on the submission have contributed sufficiently to the scientific work and therefore share collective responsibility and accountability for the results. The authors declare that they have no conflict of interest.
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Xu, GC., Bian, YQ., Han, RZ. et al. Cloning, Expression, and Characterization of budC Gene Encoding meso-2,3-Butanediol Dehydrogenase from Bacillus licheniformis . Appl Biochem Biotechnol 178, 604–617 (2016). https://doi.org/10.1007/s12010-015-1897-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1897-3