Abstract
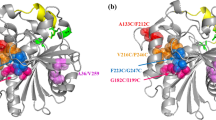
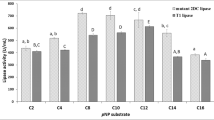
Cysteine mutants of a cold-active lipase (PcLipI) from Penicillium cyclopium were designed by the software Disulfide by Design Ver. 1.20 in an effort to improve enzyme thermostability by addition of a disulfide bridge. Those mutants predicted by molecular dynamics simulation to have better thermostability than the wild type were first expressed in Escherichia coli BL21(DE3) and then, for further investigation, in Pichia pastoris GS115. By replacing Val248 and Thr251 with cysteines to create a disulfide bridge, the recombinant lipases reE-PcLipV248C-T251C (expressed in E. coli) and reP-PcLipV248C-T251C (expressed in P. pastoris) were obtained. Both had enhanced thermostability with half-lives at 35 °C about 4.5- and 12.8-fold longer than that of the parent PcLipI expressed in E. coli and P. pastoris, respectively. The temperature optima of reE-PcLipV248C-T251C and reP-PcLipV248C-T251C were 35 and 30 °C, which were each 5 °C higher than those of the parent PcLipI expressed in E. coli and P. pastoris. The K ms of reE-PcLipV248C-T251C and reP-PcLipV248C-T251C toward tributyrin were 53.2 and 39.5 mM, while their V maxs were 1,460 and 3,800 U/mg, respectively. PcLipV248C-T251C had better thermostability and catalytic efficiency than the other mutants and the parent PcLipI.






Similar content being viewed by others
References
Jaeger, K. E., & Eggert, T. (2002). Current Opinion in Biotechnology, 13, 390–397.
Feller, G., Narinx, E., Arpigny, J. L., Aittaleb, M., Baise, E., Genicot, S., et al. (1996). FEMS Microbiology Reviews, 18, 189–202.
Hasan, F., Shah, A. A., & Hameed, A. (2006). Enzyme and Microbial Technology, 39, 235–251.
Alquati, C., De Gioia, L., Santarossa, G., Alberghina, L., Fantucci, P., & Lotti, M. (2002). European Journal of Biochemistry, 269, 3321–3328.
Liebeton, K., Zacharias, A., & Jaeger, K. E. (2001). Journal of Bacteriology, 183, 597–603.
Han, Z. L., Han, S. Y., Zheng, S. P., & Lin, Y. (2009). Applied Microbiology and Biotechnology, 85, 117–126.
Yu, X. W., Tan, N. J., Xiao, R., & Xu, Y. (2012). PLoS One, 7, e46388.
Le, Q. A. T., Joo, J. C., Yoo, Y. J., & Kim, Y. H. (2012). Biotechnology and Bioengineering, 109, 867–876.
Sha, C., Yu, X. W., Lin, N. X., Zhang, M., & Xu, Y. (2013). Enzyme and Microbial Technology, 53, 438–443.
Tan, Z. B., Li, J. F., Wu, M. C., Tang, C. D., Zhang, H. M., & Wang, J. Q. (2011). World Journal of Microbiology and Biotechnology, 27, 2767–2774.
Sambrook, J., & Russell, D. W. (2001). Molecular cloning: A laboratory manual (3rd ed.). New York: Cold Spring Harbor Laboratory.
Grochulski, P., Bouthillier, F., Kazlauskas, R. J., Serreqi, A. N., Schrag, J. D., Ziomek, E., et al. (1994). Biochemistry, 33, 3494–3500.
Uppenberg, J., Ohrner, N., Norin, M., Hult, K., Kleywegt, G. J., Patkar, S., et al. (1995). Biochemistry, 34, 16838–16851.
Lang, D. A., Mannesse, M. L., de Haas, G. H., Verheij, H. M., & Dijkstra, B. W. (1998). European Journal of Biochemistry, 254, 333–340.
Nardini, M., Lang, D. A., Liebeton, K., Jaeger, K. E., & Dijkstra, B. W. (2000). Journal of Biological Chemistry, 275, 31219–31225.
Dombkowski, A. A. (2003). Bioinformatics, 19, 1852–1853.
Badieyan, S., Bevan, D. R., & Zhang, C. (2012). Biotechnology and Bioengineering, 109, 31–44.
Pronk, S., Páll, S., Schulz, R., Larsson, P., Bjelkmar, P., Apostolov, R., et al. (2013). Bioinformatics, 29, 845–854.
Xie, Z. H., & Shi, X. J. (2009). Progress in Biochemistry and Biophysics, 36, 1490–1494.
Zhou, C., Bai, H., Deng, S., Wang, J., Zhu, J., Wu, M., et al. (2008). Bioresource Technology, 99, 831–838.
Zhang, H., Wu, M., Li, J., Gao, S., & Yang, Y. (2012). Applied Biochemistry and Biotechnology, 167, 2198–2211.
Wu, M. C., Qian, Z. K., Jiang, P. H., Min, T. S., Sun, C. R., & Huang, W. D. (2003). Lipids, 38, 191–199.
Laemmli, U. K. (1970). Nature, 227, 680–685.
Ollis, D. L., Cheah, E., Cygler, M., Dijkstra, B., Frolow, F., Franken, S. M., et al. (1992). Protein Engineering, 5, 197–211.
Liu, L., Gao, C., Lan, D., Yang, B., & Wang, Y. (2012). Biochemical and Biophysical Research Communications, 424, 285–289.
Radestock, S., & Gohlke, H. (2008). Engineering in Life Sciences, 8, 507–522.
Austin, B. P., & Waugh, D. S. (2012). Protein Expression and Purification, 82, 116–214.
Warsame, A., Vad, R., Kristensen, T., & Oyen, T. B. (2001). Biochemical and Biophysical Research Communications, 281, 1176–1182.
Cereghino, J. L., & Cregg, J. M. (2000). FEMS Microbiology Reviews, 24, 45–66.
Horchani, H., Ouertani, S., Gargouri, Y., & Sayari, A. (2009). Journal of Molecular Catalysis B: Enzymatic, 61, 194–201.
Acknowledgments
This work was financially supported by the National Natural Science Foundation of China (no. 20776061) and the Postgraduate Innovation Training Project of Jiangsu (no. CXZZ12-0758). We are grateful to Prof. Xianzhang Wu (School of Biotechnology, Jiangnan University) for providing technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Zhongbiao Tan and Jianfang Li, the two first authors, contributed equally to this work
Rights and permissions
About this article
Cite this article
Tan, Z., Li, J., Wu, M. et al. Enhancing the Thermostability of a Cold-Active Lipase from Penicillium cyclopium by In Silico Design of a Disulfide Bridge. Appl Biochem Biotechnol 173, 1752–1764 (2014). https://doi.org/10.1007/s12010-014-0962-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-014-0962-7