Abstract
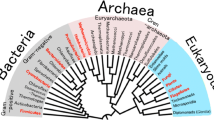
A highly efficient oil-degrading bacteria JZX-01 was isolated from the oil-contaminated soil of the seacoast near the Boxi Offshore Oil Field of China. Morphological, physiological, and 16S rDNA gene sequence analyses indicated that JZX-01 was assigned to the genus Rhodococcus sp. This strain decomposed 65.27 ± 5.63 % of the crude oil in 9 days. Gas chromatography–mass spectrometry analysis showed that even the long-chain hydrocarbons (C31–C38) and branched alkanes (pristine and phytane), which were regarded as the stubborn ones, could be degraded. Further study showed that the bacteria still has good oil degradation ability at low temperatures as well as under high salt conditions. Moreover, JZX-01 was found to have a biosurfactant-producing capacity, which significantly favors the surface tension reduction and crude oil degradation. The promising isolated strain Rhodococcus sp. JZX-01 could be further used for the bioremediation of oil-polluted soil or seawater in a wide range of temperatures and high salt conditions.





Similar content being viewed by others
References
Gojgic-Cvijovic, G. D., Milic, J. S., Solevic, T. M., Beskoski, V. P., Ilic, M. V., Djokic, L. S., et al. (2012). Biodegradation of petroleum sludge and petroleum polluted soil by a bacterial consortium: a laboratory study. Biodegradation, 23(1), 1–14.
Mnif, S., Chamkha, M., & Sayadi, S. (2009). Isolation and characterization of Halomonas sp. stain C2SS100, a hydrocarbon-degrading bacterium under hypersaline conditions. Journal of Applied Microbiology, 107(3), 785–794.
Rosenberg, A. (1983). Pseudomonas halodurans sp. nov., a halotolerant bacterium. Archives of Microbiology, 136(2), 117–123.
Gauthier, M. J., Lafay, B., Christen, R., Fernandez, L., Acquaviva, M., Bonin, P., et al. (1992). Marinobacter hydrocarbonoclasticus gen. nov., sp. nov., a new, extremely halotolerant, hydrocarbon-degrading marine bacterium. International Journal of Systematic Bacteriology, 42(4), 568–576.
Ryu, H. W., Joo, Y. H., An, Y., & Cho, K. (2006). Isolation and characterization of psychrotrophic and halotolerant Rhodococcus sp. YHLT-2. Journal of Microbiology and Biotechnology, 16(4), 605.
Margesin, R. (2000). Potential of cold-adapted microorganisms for bioremediation of oil-polluted Alpine soils. International Biodeterioration and Biodegradation, 46(1), 3–10.
Bharali, P., Das, S., Konwar, B. K., & Thakur, A. J. (2011). Crude biosurfactant from thermophilic Alcaligenes faecalis: feasibility in petro-spill bioremediation. International Biodeterioration and Biodegradation, 65(5), 682–690.
Cubitto, M. A., Moran, A. C., Commendatore, M., Chiarello, M. N., Baldini, M. D., & Sineriz, F. (2004). Effects of Bacillus subtilis O9 biosurfactant on the bioremediation of crude oil-polluted soils. Biodegradation, 15(5), 281–287.
Abalos, A., Vinas, M., Sabate, J., Manresa, M. A., & Solanas, A. M. (2004). Enhanced biodegradation of Casablanca crude oil by a microbial consortium in presence of a rhamnolipid produced by Pseudomonas aeruginosa AT10. Biodegradation, 15(4), 249–260.
Lai, C. C., Huang, Y. C., Wei, Y. H., & Chang, J. S. (2009). Biosurfactant-enhanced removal of total petroleum hydrocarbons from contaminated soil. Journal of Hazardous Materials, 167(1–3), 609–614.
Weisburg, W. G., Barns, S. M., Pelletier, D. A., & Lane, D. J. (1991). 16S ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology, 173(2), 697–703.
Saitou, N., & Nei, M. (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4(4), 406–425.
Mishra, S., Jyot, J., Kuhad, R. C., & Lal, B. (2001). In situ bioremediation potential of an oily sludge-degrading bacterial consortium. Current Microbiology, 43(5), 328–335.
Auffret, M., Labbe, D., Thouand, G., Greer, C. W., & Fayolle-Guichard, F. (2009). Degradation of a mixture of hydrocarbons, gasoline, and diesel oil additives by Rhodococcus aetherivorans and Rhodococcus wratislaviensis. Applied and Environmental Microbiology, 75(24), 7774–7782.
Sharma, S. L., & Pant, A. (2000). Biodegradation and conversion of alkanes and crude oil by a marine Rhodococcus sp. Biodegradation, 11(5), 289–294.
van Beilen, J. B., Wubbolts, M. G., & Witholt, B. (1994). Genetics of alkane oxidation by Pseudomonas oleovorans. Biodegradation, 5(3–4), 161–174.
Maier, T., Förster, H. H., Asperger, O., & Hahn, U. (2001). Molecular Characterization of the 56-kDa CYP153 from Acinetobacter sp. EB104. Biochemical and Biophysical Research Communications, 286(3), 652–658.
van Beilen, J. B., Funhoff, E. G., van Loon, A., Just, A., Kaysser, L., Bouza, M., et al. (2006). Cytochrome P450 alkane hydroxylases of the CYP153 family are common in alkane-degrading eubacteria lacking integral membrane alkane hydroxylases. Applied and Environmental Microbiology, 72(1), 59–65.
Throne-Holst, M., Wentzel, A., Ellingsen, T. E., Kotlar, H. K., & Zotchev, S. B. (2007). Identification of novel genes involved in long-chain n-alkane degradation by Acinetobacter sp. strain DSM 17874. Applied and Environmental Microbiology, 73(10), 3327–3332.
Wang, W., & Shao, Z. (2012). Genes involved in alkane degradation in the Alcanivorax hongdengensis strain A-11-3. Applied Microbiology and Biotechnology, 94(2), 437–448.
Simoni, S., Klinke, S., Zipper, C., Angst, W., & Kohler, H. E. (1996). Enantioselective metabolism of chiral 3-phenylbutyric acid, an intermediate of linear alkylbenzene degradation, by Rhodococcus rhodochrous PB1. Applied and Environmental Microbiology, 62(3), 749–755.
Brakstad, O. G., & Bonaunet, K. (2006). Biodegradation of petroleum hydrocarbons in seawater at low temperatures (0–5 degrees C) and bacterial communities associated with degradation. Biodegradation, 17(1), 71–82.
Kunihiro, N., Haruki, M., Takano, K., Morikawa, M., & Kanaya, S. (2005). Isolation and characterization of Rhodococcus sp. strains TMP2 and T12 that degrade 2,6,10,14-tetramethylpentadecane (pristane) at moderately low temperatures. Journal of Biotechnology, 115(2), 129–136.
Nhi-Cong, L. T., Mikolasch, A., Klenk, H.-P., & Schauer, F. (2009). Degradation of the multiple branched alkane 2,6,10,14-tetramethyl-pentadecane (pristane) in Rhodococcus ruber and Mycobacterium neoaurum. International Biodeterioration and Biodegradation, 63(2), 201–207.
Huy, N. Q., Jin, S., Amada, K., Haruki, M., Huu, N. B., Hang, D. T., et al. (1999). Characterization of petroleum-degrading bacteria from oil-contaminated sites in Vietnam. Journal of Bioscience and Bioengineering, 88(1), 100–102.
Obuekwe, C. O., Al-Jadi, Z. K., & Al-Saleh, E. S. (2009). Hydrocarbon degradation in relation to cell-surface hydrophobicity among bacterial hydrocarbon degraders from petroleum-contaminated Kuwait desert environment. International Biodeterioration and Biodegradation, 63(3), 273–279.
Kuyukina, M. S., & Ivshina, I. B. (2010). Multifunctional biosurfactant from non-pathogenic Rhodococcus ruber for diverse industrial applications. Journal of Biotechnology, 150, S83–S84.
Pacheco, G. J., Prioli Ciapina, E. M., Gomes, E. B., & Pereira Junior, N. (2010). Biosurfactant production by Rhodococcus erythropolis and its application to oil removal. Brazilian Journal of Microbiology, 41(3), 685–693.
Shavandi, M., Mohebali, G., Haddadi, A., Shakarami, H., & Nuhi, A. (2011). Emulsification potential of a newly isolated biosurfactant-producing bacterium, Rhodococcus sp. strain TA6. Colloids and Surfaces B-Biointerfaces, 82(2), 477–482.
Acknowledgments
The authors wish to acknowledge the financial support provided by the National Key Basic Research Program of China (No. 2014CB745100), the National Natural Science Foundation of China (No. 20906070), the Seed Foundation of Tianjin University and the Program of Introducing Talents of Discipline to Universities (No. B06006).
Author information
Authors and Affiliations
Corresponding author
Additional information
Chen Li and Zheng-Xi Zhou contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, C., Zhou, ZX., Jia, XQ. et al. Biodegradation of Crude Oil by a Newly Isolated Strain Rhodococcus sp. JZX-01. Appl Biochem Biotechnol 171, 1715–1725 (2013). https://doi.org/10.1007/s12010-013-0451-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-013-0451-4