Abstract
Xylanase encoding gene (1,224 bp) from Geobacillus thermodenitrificans was cloned in pET28a (+) vector and successfully expressed in Escherichia coli BL21 (DE3). The deduced amino acid sequence analysis revealed homology with that of glycosyl hydrolase (GH) 10 family with a high molecular mass (50 kDa). The purified recombinant xylanase is optimally active at pH 9.0 and 70 °C with T 1/2 of 10 min at 80 °C, and retains greater than 85 % activity after exposure to 70 °C for 180 min. The enzyme liberates xylose as well as xylooligosaccharides from birchwood xylan and agro-residues, and therefore, this is an endoxylanase. The xylan hydrolytic products (xylooligosaccharides, xylose, and xylobiose) find application as prebiotics and in the production of bioethanol. The xylanase being thermostable and alkalistable, it has released chromophores and phenolics from the residual lignin of pulps, suggesting its utility in mitigating chlorine requirement in pulp bleaching.




Similar content being viewed by others
References
Collins, T., Gerday, C., & Feller, G. (2005). FEMS Microbiology Reviews, 29, 3–23.
Bazzicalupo, M., & Fani, R. (1995). In: Methods in molecular biology, species diagnostic protocols: PCR and other nucleic acids methods. (Eds. Clapp, J.P.), pp. 155–77. Totowa: Humana Press.
Verma, D., & Satyanarayana, T. (2012). Bioresource Technology, 107, 333–338.
Archana, A., & Satyanarayana, T. (1997). Enzyme and Microbial Technology, 21, 12–17.
Miller, G. L. (1959). Anaytical Chemistry, 31, 426–428.
Patel, R. N., Grabski, A. C., & Jeffries, T. W. (1993). Applied Microbiology and Biotechnology, 39, 405–412.
Lambert, C., Leonard, N., De Bolle, X., & Depiereux, E. (2002). Bioinformatics, 18, 1250–1256.
Gerasimova, J., & Kuisiene, N. (2012). Microbiology, 81, 418–424.
Saksono, B., & Sukmarini, L. (2010). HAYATI Journal of Biosciences, 17, 189–197.
Canakci, S., Cevher, Z., Inan, K., Tokgoz, M., Bahar, F., Kacagan, M., et al. (2012). World Journal of Microbiology and Biotechnology, 28, 1981–1988.
Mamo, G., Hatti-Kaul, R., & Mattiasson, B. (2007). Extremophiles, 11, 169–177.
Guo, B., Chen, X. L., Sun, C. Y., Zhou, B. C., & Zhang, Y. Z. (2009). Applied Microbiology and Biotechnology, 84, 1107–1115.
Sinnot, M. L. (1990). Chemical Reviews, 90, 1171–1202.
McCarter, J. D., & Withers, S. G. (1994). Current Opinion in Structure Biology, 4, 885–892.
Shi, P., Tian, J., Yuan, T., Liu, X., Huang, H., et al. (2011). Applied and Environmental Microbiology, 76, 3620–3624.
Ko, E. P., Akatsuka, H., Moriyama, H., Shinmyo, A., Hata, Y., Kastube, Y., et al. (1992). Journal of Biochemistry, 288, 117–121.
Coughlan, M., & Visser, J. (1992). In J. Visser, G. Beldman, M. A. K. Someren, & A. G. J. Voragen (Eds.), Xylans and xylanases (pp. 111–140). Amsterdam: Elsevier.
Paes, G., Berrin, J. G., & Beaugrand, J. (2012). Advances in Biotechnology, 30, 564–592.
Jalal, A., Rashid, N., Rasool, N., & Akhtar, M. (2009). Journal of Bioscience and Bioengineering, 107, 360–365.
Hyeong, H. L., Lim, P. O., & Lee, Y. E. (2007). Journal of Microbiology and Biotechnology, 17, 29–36.
Son-Ng, I., Li, C. W., Yeh, Y., Chen, P. T., Chir, J. L., Ma, C. H., et al. (2009). Extremophiles, 13, 425–435.
Chang, W. S., Tsai, C. L., & Tseng, M. J. (2004). Biochemical and Biophysical Research Communications, 319, 1017–1025.
Cheng, Y. F., Yang, C. H., & Liu, W. H. (2005). Enzyme and Microbial Technology, 37, 541–546.
Gupta, N., Vohra, R. M., & Hoondal, G. S. (1992). Biotechnology Letters, 14, 1045–1046.
Bajaj, B. K., Razdan, K., & Sharma, A. (2010). Indian Journal of Chemical Technology, 17, 375–380.
Zhang, G., Mao, L., Zhao, Y., Xue, Y., & Ma, Y. (2010). Biotechnology Letters, 32, 1915–1920.
Khasin, A., Alchanati, I., & Shoham, Y. (1993). Applied and Environmental Microbiology, 59, 1725–1730.
Shrinivas, D., Savitha, G., & Naik, G. R. (2010). Applied Biochemistry and Biotechnology, 162, 2049–2057.
Viikari, L., Ranua, M., Kantelinen, A., Sunduist, J., & Linko, M. (1986). In: Proc 3rd Int Conf Biotechnology in the Pulp and Paper Industry, STFI, Stockholm, Sweden, pp.67–9.
Kulkarni, N., & Rao, N. (1996). Journal of Biotechnology, 51, 167–173.
Zhang, G. M., Huang, J., Huang, G. R., Ma, L. X., & Zhang, X. E. (2007). Applied Microbiology and Biotechnology, 74, 339–346.
Liu, W., Shi, P., Chen, Q., Yang, P., Wang, G., Wang, Y., et al. (2010). Applied Biochemistry and Biotechnology, 162, 1–12.
Yin, L., Lin, H., Chiang, Y., & Jiang, S. (2010). Journal of Agriculture & Food Chemistry, 58, 557–562.
Gupta, S., Bhushan, B., & Hoondal, G. S. (2000). Journal of Applied Microbiology, 88, 325–334.
Kamble, R. D., & Jadhav, A. R. (2012). ISRN Biotechnology, 2013, 1–5.
Salama, M. A., Ismail, K. M. I., Amany, H. A., Abo, E.-L., & Genweely, N. S. I. (2008). International Journal of Botany, 4, 41–48.
Lu, F., Lu, M., Lu, Z., Bie, X., Zhao, H., & Wang, Y. (2008). Bioresource Technology, 99, 5938–5941.
Cardoso, O. A. V., & Filho, E. X. F. (2003). FEMS Microbiology Letters, 223, 309–314.
Wu, S., Liu, B., & Zhang, X. (2006). Applied Microbiology and Biotechnology, 72, 1210–1216.
Do, T. T., & Quyen, D. T. (2010). Middle East Journal of Scientific Research, 6, 392–397.
Torronen, A., Mach, R. L., Messner, R., Gonzalez, R., Kakkinen, N., Harkki, A., et al. (1992). Biotechnology (NY), 10, 1461–1465.
Khandeparkar, R. D. S., & Bhosle, N. B. (2006). Enzyme and Microbial Technology, 4, 732–742.
Gruppen, H., Hamer, R. J., & Voragen, A. G. J. (1992). Journal of Cereal Science, 16, 53–67.
Kormelink, F. J. M., & Voragen, A. G. J. (1993). Applied Microbiology and Biotechnology, 38, 688–695.
Jiang, Z. Q., Deng, W., Zhu, Y. P., Li, L. T., Sheng, Y. J., & Hayashi, K. (2004). Journal of Molecular Catalysis B Enzymatic, 27, 207–213.
Ratanakhanokchai, K., Kyu, K. L., & Tanticharoen, M. (1999). Applied and Environmental Microbiology, 65, 694–697.
Damiano, V. B., Bocchini, D. A., Gomes, E., & Da Silva, R. (2003). World Journal of Microbiology and Biotechnology, 19, 139–144.
Choudhury, B., Sunita, C., Singh, S. N., & Ghosh, P. (2006). World Journal of Microbiology and Biotechnology, 22, 283–288.
Kaur, A., Mahajan, R., Singh, A., Garg, G., & Sharma, J. (2010). Bioresource Technology, 101, 9150–9155.
Acknowledgments
The authors are grateful to the Ministry of Environment and Forests, Govt. of India, New Delhi for partial financial assistance while carrying out this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Nucleotide and deduced amino acid sequences of the G. thermodenitrificans xylanase gene. The underlined regions I (I V A E N V M K), II (R F H T L V W H), III (D V V N E), IV (L Y I N D Y N), V (I G H Q S H I), VI (I T E L D V), VII (T F W G I A D N H T W), and VIII (D Y I K V A F Q T A) denote highly conserved GH10 xylanases. Glu187, Asp230, and Glu293 are crucial catalytic residues. (DOC 40 kb)
Supplementary Fig. 2
Secondary structure of rXyl-gtd was proposed using ESPript 2.2 software. The structure is based on the available template of xylanase from 1HIZ chain A of G. stearothermophilus. Symbols α, β, and TT denote the helix, sheets, and turns, respectively. (DOC 237 kb)
Supplementary Fig.3
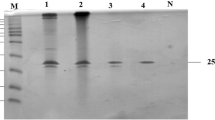
Electrophoretic analysis of the rXyl-gtd using 15 % SDS-PAGE. a Purified rXyl-gtd of 50 kDa; M, standard protein markers; lanes 1 and 2 show eluted proteins with 200 and 250 mM imidazole; b zymogram analysis of purified rXyl-gtd using Congo red. (DOC 644 kb)
Rights and permissions
About this article
Cite this article
Verma, D., Anand, A. & Satyanarayana, T. Thermostable and Alkalistable Endoxylanase of the Extremely Thermophilic Bacterium Geobacillus thermodenitrificans TSAA1: Cloning, Expression, Characteristics and Its Applicability in Generating Xylooligosaccharides and Fermentable Sugars. Appl Biochem Biotechnol 170, 119–130 (2013). https://doi.org/10.1007/s12010-013-0174-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-013-0174-6