Abstract
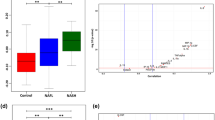
Liver fibrosis (LF) is the final stage of liver dysfunction, characterized by diffuse fibrosis which is the main response to the liver injury. Haptoglobin (HP) protein, produced as an acute phase reactant during LF, preventing liver damage, may be potential molecular targets for early LF diagnostics and therapeutic applications. However, protein networks associated with the HP are largely unknown. To address this issue, we used a pathological mouse model of LF that was induced by treatment with carbon tetrachloride for 8 days. HP protein was separated and identified by two-dimensional gel electrophoresis and matrix-assisted laser desorption/ionization time-of-flight/time-of-flight mass spectrometry. HP protein was subjected to functional pathway analysis using STRING and Cytoscape software for better understanding of the protein–protein interaction (PPI) networks in biological context. Bioinformatics analyses revealed that HP expression associated with fibrosis was upregulated, and suggested that HP responsible for fibrosis may precede the onset and progression of LF. Using the web-based database, functional pathway analysis suggested the modulation of multiple vital physiological pathways, including antioxidation immunity, signal transduction, metabolic process, energy production, cell apoptosis, oxidation reduction, DNA repair process, cell communication, and regulation of cellular process. The generation of protein interaction networks clearly enhances the interpretation and understanding of the molecular mechanisms of HP. HP protein represents targets for further experimental investigation that will provide biological insight and potentially could be exploited for novel therapeutic approaches to combat LF.




Similar content being viewed by others
Reference
Behrends, C., Sowa, M. E., Gygi, S. P., & Harper, J. W. (2010). Network organization of the human autophagy system. Nature, 466(7302), 68–76.
Stuart, L. M., Boulais, J., Charriere, G. M., Hennessy, E. J., Brunet, S., Jutras, I., Goyette, G., Rondeau, C., Letarte, S., Huang, H., Ye, P., Morales, F., Kocks, C., Bader, J. S., Desjardins, M., & Ezekowitz, R. A. (2007). A systems biology analysis of the Drosophila phagosome. Nature, 445(7123), 95–101.
Jørgensen, C., Sherman, A., Chen, G. I., Pasculescu, A., Poliakov, A., Hsiung, M., Larsen, B., Wilkinson, D. G., Linding, R., & Pawson, T. (2009). Cell-specific information processing in segregating populations of Eph receptor ephrin-expressing cells. Science, 326(5959), 1502–1509.
Zhang, K., Yuan, K., Wu, H., Li, Q., Wang, Y., Chen, S., Zhang, L., Gu, H., & Fu, R. (2012). Identification of potential markers related to neoadjuvant chemotherapy sensitivity of breast cancer by SELDI-TOF MS. Applied Biochemistry and Biotechnology, 166(3), 753–763.
Tang, C., Iwahara, J., & Clore, G. M. (2006). Visualization of transient encounter complexes in protein–protein association. Nature, 444(7117), 383–386.
Jielile, J., Jialili, A., Sabirhazi, G., Shawutali, N., Redati, D., Chen, J., Tang, B., Bai, J., & Aldyarhan, K. (2011). Proteomic analysis of differential protein expression of Achilles tendon in a rabbit model by two-dimensional polyacrylamide gel electrophoresis at 21 days postoperation. Applied Biochemistry and Biotechnology, 165(3-4), 1092–1106.
Yu, H., Braun, P., Yildirim, M. A., Lemmens, I., Venkatesan, K., Sahalie, J., Hirozane-Kishikawa, T., Gebreab, F., Li, N., Simonis, N., Hao, T., Rual, J. F., Dricot, A., Vazquez, A., Murray, R. R., Simon, C., Tardivo, L., Tam, S., Svrzikapa, N., Fan, C., de Smet, A. S., Motyl, A., Hudson, M. E., Park, J., Xin, X., Cusick, M. E., Moore, T., Boone, C., Snyder, M., Roth, F. P., Barabási, A. L., Tavernier, J., Hill, D. E., & Vidal, M. (2008). High-quality binary protein interaction map of the yeast interactome network. Science, 322(5898), 104–110.
Hew, C. S., & Gam, L. H. (2011). Proteome analysis of abundant proteins extracted from the leaf of Gynura procumbens (Lour.) Merr. Applied Biochemistry and Biotechnology, 165(7-8), 1577–1586.
Wang, X., Zhang, A., Sun, H., Wu, G., Sun, W., & Yan, G. (2012). Network generation enhances interpretation of proteomics data sets by a combination of two-dimensional polyacrylamide gel electrophoresis and matrix-assisted laser desorption/ionization-time of flight mass spectrometry. Analyst, 137(20), 4703–4711.
Li, F., Glinskii, O. V., Zhou, J., Wilson, L. S., Barnes, S., Anthony, D. C., & Glinsky, V. V. (2011). Identification and analysis of signaling networks potentially involved in breast carcinoma metastasis to the brain. PLoS One, 6(7), e21977.
Bonetta, L. (2010). Protein–protein interactions: interactome under construction. Nature, 468(7325), 851–854.
Franzosa, E. A., & Xia, Y. (2011). Structural principles within the human-virus protein–protein interaction network. Proceedings of the National Academy of Sciences of the United States of America, 108(26), 10538–41053.
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., Amin, N., Schwikowski, B., & Ideker, T. (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Research, 13, 2498–2504.
Deighton, R. F., Kerr, L. E., Short, D. M., Allerhand, M., Whittle, I. R., & McCulloch, J. (2010). Network generation enhances interpretation of proteomic data from induced apoptosis. Proteomics, 10(6), 1307–1315.
Koh, G. C., Porras, P., Aranda, B., Hermjakob, H., & Orchard, S. E. (2012). Analyzing protein–protein interaction networks. Journal of Proteome Research, 11(4), 2014–2031.
White, I. R., Patel, K., Symonds, W. T., Dev, A., Griffin, P., Tsokanas, N., Skehel, M., Liu, C., Zekry, A., Cutler, P., Gattu, M., Rockey, D. C., Berrey, M. M., & McHutchison, J. G. (2007). Serum proteomic analysis focused on fibrosis in patients with hepatitis C virus infection. Journal of Translational Medicine, 5, 33.
Ngai, H. H., Sit, W. H., Jiang, P. P., Thongboonkerd, V., & Wan, J. M. (2007). Markedly increased urinary preprohaptoglobin and haptoglobin in passive Heymann nephritis: a differential proteomics approach. Journal of Proteome Research, 6(8), 3313–3320.
Ang, I. L., Poon, T. C., Lai, P. B., Chan, A. T., Ngai, S. M., Hui, A. Y., Johnson, P. J., & Sung, J. J. (2006). Study of serum haptoglobin and its glycoforms in the diagnosis of hepatocellular carcinoma: a glycoproteomic approach. Journal of Proteome Research, 5(10), 2691–2700.
Ahmed, N., Barker, G., Oliva, K. T., Hoffmann, P., Riley, C., Reeve, S., Smith, A. I., Kemp, B. E., Quinn, M. A., & Rice, G. E. (2004). Proteomic-based identification of haptoglobin-1 precursor as a novel circulating biomarker of ovarian cancer. British Journal of Cancer, 91(1), 129–140.
Zhang, A., Sun, H., Yuan, Y., Sun, W., Jiao, G., & Wang, X. (2011). An in vivo analysis of the therapeutic and synergistic properties of Chinese medicinal formula Yin-Chen-Hao-Tang based on its active constituents. Fitoterapia, 82(8), 1160–1168.
Heo, M., Maslov, S., & Shakhnovich, E. (2011). Topology of protein interaction network shapes protein abundances and strengths of their functional and nonspecific interactions. Proceedings of the National Academy of Sciences of the United States of America, 108(10), 4258–4263.
Lim, S. R., Gooi, B. H., Singh, M., & Gam, L. H. (2011). Analysis of differentially expressed proteins in colorectal cancer using hydroxyapatite column and SDS-PAGE. Applied Biochemistry and Biotechnology, 165(5-6), 1211–1224.
Jensen, L. J., Kuhn, M., Stark, M., Chaffron, S., Creevey, C., Muller, J., Doerks, T., Julien, P., Roth, A., Simonovic, M., Bork, P., & von Mering, C. (2009). STRING 8—a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Research, 37, D412–D416.
Bakal, C., Linding, R., Llense, F., Heffern, E., Martin-Blanco, E., Pawson, T., & Perrimon, N. (2008). Phosphorylation networks regulating JNK activity in diverse genetic backgrounds. Science, 322(5900), 453–456.
Gutiérrez-Sánchez, G., Atwood, J., Kolli, V. S., Roussos, S., & Augur, C. (2012). Initial proteome analysis of caffeine-induced proteins in Aspergillus tamarii using two-dimensional fluorescence difference gel electrophoresis. Applied Biochemistry and Biotechnology, 166(8), 2064–2077.
Karthik, D., Ilavenil, S., Kaleeswaran, B., Sunil, S., & Ravikumar, S. (2012). Proteomic analysis of plasma proteins in diabetic rats by 2D electrophoresis and MALDI-TOF-MS. Applied Biochemistry and Biotechnology, 166(6), 1507–1519.
Wang, X., Zhang, A., Han, Y., Wang, P., Sun, H., Song, G., Dong, T., Yuan, Y., Yuan, X., Zhang, M., Xie, N., Zhang, H., Dong, H., & Dong, W. (2012). Urine metabolomics analysis for biomarker discovery and detection of jaundice syndrome in patients with liver disease. Molecular & Cellular Proteomics, 11(8), 370–380.
McDermott, J. E., Diamond, D. L., Corley, C., Rasmussen, A. L., Katze, M. G., & Waters, K. M. (2012). Topological analysis of protein co-abundance networks identifies novel host targets important for HCV infection and pathogenesis. BMC Systems Biology, 6, 28.
Chakraborty, C., Pal, S., Doss, C. G., Wen, Z. H., & Lin, C. S. (2012). In silico analyses of COMT, an important signaling cascade of dopaminergic neurotransmission pathway, for drug development of Parkinson’s disease. Applied Biochemistry and Biotechnology, 167(4), 845–860.
Acknowledgments
This work was supported by grants from the Key Program of Natural Science Foundation of State (grant no. 90709019), the National Specific Program on the Subject of Public Welfare (grant no. 200807014), National Key Subject of Drug Innovation (grant no. 2009ZX09502-005), and National Program on Key Basic Research Project of China (grant no. 2005CB523406).
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 66 kb)
Rights and permissions
About this article
Cite this article
Zhang, A., Sun, H., Sun, W. et al. Proteomic Identification Network Analysis of Haptoglobin as a Key Regulator Associated with Liver Fibrosis. Appl Biochem Biotechnol 169, 832–846 (2013). https://doi.org/10.1007/s12010-012-0001-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-012-0001-5