Abstract
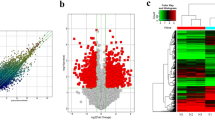
β-thalassemia is caused by β-globin gene mutations. However, heterogeneous phenotypes were found in individuals with same genotype, and still undescribed mechanism underlies such variation. We collected blood samples from 30 β-thalassemia major, 30 β-thalassemia minor patients, and 30 matched normal controls. Human lncRNA Array v2.0 (8 × 60 K, Arraystar) was used to detect changes in long non-coding RNAs (lncRNAs) and mRNAs in three samples each from β-thalassemia major, β-thalassemia minor, and control groups. Compared with normal controls, 1424 and 2045 lncRNAs were up- and downregulated, respectively, in β-thalassemia major patients, whereas 623 and 349 lncRNAs were up- and downregulated, respectively, in β-thalassemia minor patients. Compared with β-thalassemia minor group, 1367 and 2356 lncRNAs were up- and downregulated, respectively, in β-thalassemia major group. We selected five lncRNAs that displayed altered expressions (DQ583499, X-inactive specific transcript (Xist), lincRNA-TPM1, MRFS16P, and lincRNA-RUNX2-2) and confirmed their expression levels in all samples using real-time polymerase chain reaction. Based on coding-noncoding gene co-expression network and gene ontology biological process analyses, several signaling pathways were associated with three common organ systems exhibiting β-thalassemia phenotypes: hematologic, skeletal, and hepatic systems. This study implicates that abnormal expression levels of lncRNAs and mRNA in β-thalassemia cases may be correlated with its various clinical phenotypes.
Similar content being viewed by others
References
Tuzmen S, Schechter AN. Genetic diseases of hemoglobin: diagnostic methods for elucidating β-thalassemia mutations. Blood Rev 2001; 15(1): 19–29
Cao A, Galanello R. β-thalassemia. Genet Med 2010; 12(2): 61–76
Galanello R, Ruggeri R, Paglietti E, Addis M, Melis MA, Cao A. A family with segregating triplicated α globin loci and β thalassemia. Blood 1983; 62(5): 1035–1040
Harteveld CL, Refaldi C, Cassinerio E, Cappellini MD, Giordano PC. Segmental duplications involving the α-globin gene cluster are causing β-thalassemia intermedia phenotypes in β-thalassemia heterozygous patients. Blood Cells Mol Dis 2008; 40(3): 312–316
Sollaino MC, Paglietti ME, Perseu L, Giagu N, Loi D, Galanello R. Association of α globin gene quadruplication and heterozygous β thalassemia in patients with thalassemia intermedia. Haematologica 2009; 94(10): 1445–1448
Viprakasit V, Tanphaichitr VS, Chinchang W, Sangkla P, Weiss MJ, Higgs DR. Evaluation of α hemoglobin stabilizing protein (AHSP) as a genetic modifier in patients with β thalassemia. Blood 2004; 103(9): 3296–3299
Sankaran VGMT, Menne TF, Šćepanović D, Vergilio JA, Ji P, Kim J, Thiru P, Orkin SH, Lander ES, Lodish HF. MicroRNA-15a and- 16-1 act via MYB to elevate fetal hemoglobin expression in human trisomy 13. Proc Natl Acad Sci USA 2011; 108(4): 1519–1524
Lee YT, de Vasconcellos JF, Yuan J, Byrnes C, Noh SJ, Meier ER, Kim KS, Rabel A, Kaushal M, Miller JL. LIN28B-mediated expression of fetal hemoglobin and production of fetal-like erythrocytes from adult human erythroblasts ex vivo. Blood 2013; 122(6): 1034–1041
Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov 2011; 1(5): 391–407
Guttman M, Donaghey J, Carey BW, Garber M, Grenier JK, Munson G, Young G, Lucas AB, Ach R, Bruhn L, Yang X, Amit I, Meissner A, Regev A, Rinn JL, Root DE, Lander ES. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 2011; 477(7364): 295–300
Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell 2009; 136(4): 629–641
Guttman M, Rinn JL. Modular regulatory principles of large noncoding RNAs. Nature 2012; 482(7385): 339–346
Wapinski O, Chang HY. Long noncoding RNAs and human disease. Trends Cell Biol 2011; 21(6): 354–361
Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ, Shang JL, Gao CF, Zhang FR, Wang F, Sun SH. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology 2011; 54(5): 1679–1689
Yu G, Yao W, Wang J, Ma X, Xiao W, Li H, Xia D, Yang Y, Deng K, Xiao H, Wang B, Guo X, Guan W, Hu Z, Bai Y, Xu H, Liu J, Zhang X, Ye Z. lncRNAs expression signatures of renal clear cell carcinoma revealed by microarray. PLoS ONE 2012; 7(8): e42377
Akrami R, Jacobsen A, Hoell J, Schultz N, Sander C, Larsson E. Comprehensive analysis of long non-coding RNAs in ovarian cancer reveals global patterns and targeted DNA amplification. PLoS One 2013; 8(11): e80306
Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010; 329(5992): 689–693
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007; 129(7): 1311–1323
Alvarez-Dominguez JR, Hu W, Yuan B, Shi J, Park SS, Gromatzky AA, van Oudenaarden A, Lodish HF. Global discovery of erythroid long noncoding RNAs reveals novel regulators of red cell maturation. Blood 2014; 123(4): 570–581
Thein SL. The molecular basis of β-thalassemia. Cold Spring Harb Perspect Med 2013; 3(5): a011700
Colah R, Nadkarni A, Gorakshakar A, Phanasgaonkar S, Surve R, Subramaniam PG, Bondge N, Pujari K, Ghosh K, Mohanty D. Impact of β globin gene mutations on the clinical phenotype of beta thalassemia in India. Blood Cells Mol Dis 2004; 33(2): 153–157
Kim A, Kiefer CM, Dean A. Distinctive signatures of histone methylation in transcribed coding and noncoding human β-globin sequences. Mol Cell Biol 2007; 27(4): 1271–1279
Martin DI, Ward R, Suter CM. Germline epimutation: a basis for epigenetic disease in humans. Ann N Y Acad Sci 2005; 1054(1): 68–77
Sankaran VG, Menne TF, Xu J, Akie TE, Lettre G, Van Handel B, Mikkola HK, Hirschhorn JN, Cantor AB, Orkin SH. Human fetal hemoglobin expression is regulated by the developmental stagespecific repressor BCL11A. Science 2008; 322(5909): 1839–1842
Jawaid K, Wahlberg K, Thein SL, Best S. Binding patterns of BCL11A in the globin and GATA1 loci and characterization of the BCL11A fetal hemoglobin locus. Blood Cells Mol Dis 2010; 45(2): 140–146
Zhou D, Liu K, Sun CW, Pawlik KM, Townes TM. KLF1 regulates BCL11A expression and γ- to β-globin gene switching. Nat Genet 2010; 42(9): 742–744
Thein SL. Genetic association studies in β-hemoglobinopathies. Hematology Am Soc Hematol Educ Program 2013;354–361
Engreitz JM, Pandya-Jones A, McDonel P, Shishkin A, Sirokman K, Surka C, Kadri S, Xing J, Goren A, Lander ES, Plath K, Guttman M. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 2013; 341(6147): 1237973
Jin X, Kim JG, Oh MJ, Oh HY, Sohn YW, Pian X, Yin JL, Beck S, Lee N, Son J, Kim H, Yan C, Wang JH, Choi YJ, Whang KY. Opposite roles of MRF4 and MyoD in cell proliferation and myogenic differentiation. Biochem Biophys Res Commun 2007; 364(3): 476–482
Sweetman D, Goljanek K, Rathjen T, Oustanina S, Braun T, Dalmay T, Münsterberg A. Specific requirements of MRFs for the expression of muscle specific microRNAs, miR-1, miR-206 and miR-133. Dev Biol 2008; 321(2): 491–499
Perry SV. Vertebrate tropomyosin: distribution, properties and function. J Muscle Res Cell Motil 2001; 22(1): 5–49
van de Meerakker JB, Christiaans I, Barnett P, Lekanne Deprez RH, Ilgun A, Mook OR, Mannens MM, Lam J, Wilde AA, Moorman AF, Postma AV. A novel α-tropomyosin mutation associates with dilated and non-compaction cardiomyopathy and diminishes actin binding. Biochim Biophys Acta 2013; 1833(4): 833–839
Jones PA, Laird PW. Cancer epigenetics comes of age. Nat Genet 1999; 21(2): 163–167
Boyd J, Risinger JI, Wiseman RW, Merrick BA, Selkirk JK, Barrett JC. Regulation of microfilament organization and anchorageindependent growth by tropomyosin 1. Proc Natl Acad Sci USA 1995; 92(25): 11534–11538
Otto F, Kanegane H, Mundlos S. Mutations in the RUNX2 gene in patients with cleidocranial dysplasia. Hum Mutat 2002; 19(3): 209–216
Lanaya H, Natarajan A, Komposch K, Li L, Amberg N, Chen L, Wculek SK, Hammer M, Zenz R, Peck-Radosavljevic M, Sieghart W, Trauner M, Wang H, Sibilia M. EGFR has a tumour-promoting role in liver macrophages during hepatocellular carcinoma formation. Nat Cell Biol 2014; 16(10): 972–981, 1–7
Yannakoudakis BZ, Liu KJ. Common skeletal features in rare diseases: new links between ciliopathies and FGF-related syndromes. Rare Dis 2013; 1(1): e27109
Vasikova A, Belickova M, Budinska E, Cermak J. A distinct expression of various gene subsets in CD34+ cells from patients with early and advanced myelodysplastic syndrome. Leuk Res 2010; 34(12): 1566–1572
Lee K, Briehl MM, Mazar AP, Batinic-Haberle I, Reboucas JS, Glinsmann-Gibson B, Rimsza LM, Tome ME. The copper chelator ATN-224 induces peroxynitrite-dependent cell death in hematological malignancies. Free Radic Biol Med 2013; 60: 157–167
Acknowledgements
We thank all donors who participated in this program, all of our colleagues at Bao’an Maternal and Children Health Hospital, and all those who contributed to microarray service at Kang Chen Biotechnology Company in Shanghai. This work was supported by grants from the Science and Technology Plan of Shenzhen City, Guangdong Province, China (No. JCYJ20130402151000859), the Shanghai Science and Technology Commission Major Project (No. 11dz1950300), and the Key Project of the Shanghai Municipal Commission of Health and Family Planning (No. 2013ZYJB0015).
Author information
Authors and Affiliations
Corresponding authors
Additional information
These authors contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Ma, J., Liu, F., Du, X. et al. Changes in lncRNAs and related genes in β-thalassemia minor and β-thalassemia major. Front. Med. 11, 74–86 (2017). https://doi.org/10.1007/s11684-017-0503-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11684-017-0503-1