Abstract
Breast cancer ranks as the top reason for the oncologic mortality for female around the world. The occurrence rate of breast cancer is rapidly rising, especially in China. Although the therapeutic regimes for breast cancer are diverse, the treatment outcome in patients remains dismal. Long non-coding RNAs have been greatly reported as important participators in cancer progression during the past decades. FBXL19 antisense RNA 1 (FBXL19-AS1) has been identified as a novel oncogene in colorectal cancer recently, but its role in breast cancer remains unknown. Present study attempted to explore the functional role and mechanism of FBXL19-AS1 in breast cancer progression. Expression of FBXL19-AS1, lin-28 homolog A (LIN28A), and WD repeat domain 66 (WDR66) were detected by qPCR and Western blotting. Transwell assay was used to detect cell migration and invasion. RIP assay was used to examine interaction between LIN28A and FBXL19-AS1. First, FBXL19-AS1 was highly expressed in breast cancer cell lines. Loss-of-function assays indicated that FBXL19-AS1 promoted cell migration, invasion, and EMT in breast cancer. Mechanistically, FBXL19-AS1 interacted with and was stabilized by LIN28A, an RNA-binding protein which has been reported to be able to stabilize lncRNAs. Moreover, WDR66 expression was promoted by FBXL19-AS1 at mRNA and protein level. Finally, rescue assays suggested that FBXL19-AS1 promoted migration, invasion, and EMT through regulating WDR66 in breast cancer. Current study proved that LIN28A-stabilized FBXL19-AS1 promoted breast cancer metastasis by regulating WDR66, identifying FBXL19-AS1 as a new biological marker in breast cancer.





Similar content being viewed by others
References
Bai Y, Zhou X, Huang L, Wan Y, Li X, Wang Y (2018) Long noncoding RNA EZR-AS1 promotes tumor growth and metastasis by modulating Wnt/β-catenin pathway in breast cancer. Exp Ther Med 16:2235–2242
Becker S (2015) A historic and scientific review of breast cancer: the next global healthcare challenge. Int J Gynecol Obstet 131:S36–S39
Chen C, Zhou L, Wang H, Chen J, Li W, Liu W, Shen M, Liu H, Fu X (2018a) Long noncoding RNA CNALPTC1 promotes cell proliferation and migration of papillary thyroid cancer via sponging miR-30 family. Am J Cancer Res 8:192–206
Chen Y, Huang W, Sun W, Zheng B, Wang C, Luo Z, Wang J, Yan W (2018b) LncRNA MALAT1 promotes Cancer metastasis in osteosarcoma via activation of the PI3K-Akt signaling pathway. Cell Physiol Biochem 51:1313–1326
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, Ruan Y, Lassmann T, Carninci P, Brown JB, Lipovich L, Gonzalez JM, Thomas M, Davis CA, Shiekhattar R, Gingeras TR, Hubbard TJ, Notredame C, Harrow J, Guigó R (2012) The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 22:1775–1789
Fong HK, Hurley JB, Hopkins RS, Miake-Lye R, Johnson MS, Doolittle RF, Simon MI (1986) Repetitive segmental structure of the transducin beta subunit: homology with the CDC4 gene and identification of related mRNAs. Proc Natl Acad Sci 83:2162–2166
Gunsalus KTW, Wagoner MP, Meyer K, Potter WB, Schoenike B, Kim S, Alexander CM, Friedl A, Roopra A (2012) Induction of the RNA regulator LIN28A is required for the growth and pathogenesis of RESTless breast tumors. Cancer Res 72:3207–3216
Han D, Gao X, Wang M, Qiao Y, Xu Y, Yang J, Dong N, He J, Sun Q, Lv G, Xu C, Tao J, Ma N (2016) Long noncoding RNA H19 indicates a poor prognosis of colorectal cancer and promotes tumor growth by recruiting and binding to eIF4A3. Oncotarget 7:22159–22173
Hong W, Dong E (2014) The past, present and future of breast cancer research in China. Cancer Lett 351:1–5
Lei W, Wang ZL, Feng HJ, Lin XD, Li CZ, Fan D (2018) Long non-coding RNA SNHG12promotes the proliferation and migration of glioma cells by binding to HuR. Int J Oncol. https://doi.org/10.3892/ijo.2018.4478
Li Y, Jiang B, Wu X, Huang Q, Chen W, Zhu H, Qu X, Xie L, Ma X, Huang G (2018) Long non-coding RNA MIAT is estrogen-responsive and promotes estrogen-induced proliferation in ER-positive breast cancer cells. Biochem Biophys Res Commun 503:45–50
Lian Y, Yan C, Ding J, Xia R, Ma Z, Hui B, Ji H, Zhou J, Wang K (2017) A novel lncRNA, LL22NC03-N64E9.1, represses KLF2 transcription through binding with EZH2 in colorectal cancer. Oncotarget 8:59435–59445
Liu X, Zheng J, Xue Y, Qu C, Chen J, Wang Z, Li Z, Zhang L, Liu Y (2018) Inhibition of TDP43-mediated SNHG12-miR-195-SOX5 feedback loop impeded malignant biological behaviors of glioma cells. Mol Ther Nucleic Acids 10:142–158
Lu G, Li Y, Ma Y, Lu J, Chen Y, Jiang Q, Qin Q, Zhao L, Huang Q, Luo Z, Huang S, Wei Z (2018) Long noncoding RNA LINC00511 contributes to breast cancer tumourigenesis and stemness by inducing the miR-185-3p/E2F1/Nanog axis. J Exp Clin Cancer Res 37:289–289
Luo Y, Ouyang J, Zhou D, Zhong S, Wen M, Ou W, Yu H, Jia L, Huang Y (2018) Long noncoding RNA GAPLINC promotes cells migration and invasion in colorectal Cancer cell by regulating miR-34a/c-MET signal pathway. Dig Dis Sci 63:890–899
Piipponen M, Heino J, Kähäri V-M, Nissinen L (2018) Long non-coding RNA PICSAR decreases adhesion and promotes migration of squamous carcinoma cells by downregulating α2β1 and α5β1 integrin expression. Biology Open 7:bio037044
Qi D, Li J, Que B, Su J, Li M, Zhang C, Yang M, Zhou G, Ji W (2016) Long non-coding RNA DBCCR1-003 regulate the expression of DBCCR1 via DNMT1 in bladder cancer. Cancer Cell Int 16:81
Runowicz CD, Leach CR, Henry NL, Henry KS, Mackey HT, Cowens-Alvarado RL, Cannady RS, Pratt-Chapman ML, Edge SB, Jacobs LA, Hurria A, Marks LB, LaMonte SJ, Warner E, Lyman GH, Ganz PA (2016) American Cancer Society/American Society of Clinical Oncology breast Cancer survivorship care guideline. J Clin Oncol 34:611–635
Shen B, Yuan Y, Zhang Y, Yu S, Peng W, Xe H, Feng J (2017) Long non-coding RNA FBXL19-AS1 plays oncogenic role in colorectal cancer by sponging miR-203. Biochem Biophys Res Commun 488:67–73
Shen H, Zhao L, Feng X, Xu C, Li C, Niu Y (2016) Lin28A activates androgen receptor via regulation of c-myc and promotes malignancy of ER-/Her2+ breast cancer. Oncotarget 7:60407–60418
Shi D, Zhang Y, Lu R, Zhang Y (2018) The long non-coding RNA MALAT1 interacted with miR-218 modulates choriocarcinoma growth by targeting Fbxw8. Biomed Pharmacother 97:543–550
Shi Y, Li J, Liu Y, Ding J, Fan Y, Tian Y, Wang L, Lian Y, Wang K, Shu Y (2015) The long noncoding RNA SPRY4-IT1 increases the proliferation of human breast cancer cells by upregulating ZNF703 expression. Mol Cancer 14:51–51
Sun Y, Jin JG, Mi WY, Wu H, Zhang SR, Meng Q, Zhang ST (2017) Long non-coding RNA UCA1 targets miR-122 to promote proliferation, migration, and invasion of glioma cells. Oncol Res 26
Sun Y, Zeng C, Gan S, Li H, Cheng Y, Chen D, Li R, Zhu W (2018) LncRNA HOTTIP-mediated HOXA11 expression promotes cell growth, migration and inhibits cell apoptosis in breast Cancer. Int J Mol Sci 19:472
Wang C, Gu Y, Zhang E, Zhang K, Qin N, Dai J, Zhu M, Liu J, Xie K, Jiang Y, Guo X, Liu M, Jin G, Ma H, Jiang T, Yin R, Xia Y, Liu L, Wang S, Shen B, Huo R, Xu L, Sha J, Qu B, Shen H, Hu Z (2018a) A cancer-testis non-coding RNA LIN28B-AS1 activates driver gene LIN28B by interacting with IGF2BP1 in lung adenocarcinoma. Oncogene 38:1611–1624. https://doi.org/10.1038/s41388-018-0548-x
Wang Q, Ma C, Kemmner W (2013) Wdr66 is a novel marker for risk stratification and involved in epithelial-mesenchymal transition of esophageal squamous cell carcinoma. BMC Cancer 13:137
Wang Z, Pang J, Ji B, Zhang S, Cheng Y, Yu L, Pan W (2018b) RNA binding protein Lin28A promotes osteocarcinoma cells progression by associating with the long noncoding RNA MALAT1. Biotechnol Lett 40:493–500
Yang F, Wu Q, Zhang Y, Xiong H, Li X, Li B, Xie W, Zhang L, Xu M, Zhang K, He F (2018) LncRNA LOC653786 promotes growth of RCC cells via upregulating FOXM1. Oncotarget 9:12101–12111
Yong W, Yu D, Jun Z, Yachen D, Weiwei W, Midie X, Xingzhu J, Xiaohua W (2018) Long noncoding RNA NEAT1, regulated by LIN28B, promotes cell proliferation and migration through sponging miR-506 in high-grade serous ovarian cancer. Cell Death Dis 9:861–861
Zhang Y, Chen W, Pan T, Wang H, Zhang Y, Li C (2019) LBX2-AS1 is activated by ZEB1 and promotes the development of esophageal squamous cell carcinoma by interacting with HNRNPC to enhance the stability of ZEB1 and ZEB2 mRNAs. Biochem Biophys Res Commun 511:566–572. https://doi.org/10.1016/j.bbrc.2019.02.079
Zhang Y, Yu S, Zhang Z, Gang Z, Jia X (2018a) Long non-coding RNA AK096174 promotes cell proliferation and invasion in gastric cancer by regulating WDR66 expression. Biosci Rep 38:BSR20180277
Zhang Z, Sun L, Zhang Y, Lu G, Li Y, Wei Z (2018b) Long non-coding RNA FEZF1-AS1 promotes breast cancer stemness and tumorigenesis via targeting miR-30a/Nanog axis. J Cell Physiol 233:8630–8638
Zhong W, Yang J, Li M, Li L, Li A (2018) Long noncoding RNA NEAT1 promotes the growth of human retinoblastoma cells via regulation of miR-204/CXCR4 axis. J Cell Physiol:0
Acknowledgements
The authors sincerely appreciate all members contributed to this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Additional information
Editor: Tersuji Okamoto
Yayuan Zhang and Xiaojun Xiao are co-first author
Electronic supplementary material
Supplementary Figure 1
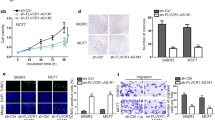
A Results of qPCR confirmed the overexpression of FBXL19-AS1 by pcDNA3.1/FBXL19-AS1 in BT474 and MDA-MB-231 cells. B–C Transwell assay showed that cell migration and invasion were enhanced by FBXL19-AS1 overexpression. D Western blot assay showed that E-cadherin level was decreased, whereas N-cadherin and Vimentin levels were increased by FBXL19-AS1 overexpression. E The 13 RNA-binding proteins predicted to interact with FBXL19-AS1 in Starbase. *P < 0.05, **P < 0.01, ***P < 0.001. (PNG 676 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Xiao, X., Zhou, W. et al. LIN28A-stabilized FBXL19-AS1 promotes breast cancer migration, invasion and EMT by regulating WDR66. In Vitro Cell.Dev.Biol.-Animal 55, 426–435 (2019). https://doi.org/10.1007/s11626-019-00361-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-019-00361-4