Abstract
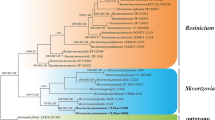
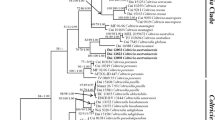
Two new species of Conidiobolus were isolated from Anhui Province, China. A polyphasic taxonomic approach comprising morphological characteristics and molecular data (the nuclear large subunit of ribosomal DNA and the translation elongation factor 1 alpha gene sequences) was applied to determine their novel taxonomic status in the genus Conidiobolus. The new species C. mirabilis is a sister group to C. thromoides, but morphologically differs by smaller primary conidia and a unique formation of zygospores among two to four adjacent hyphal segments. The other new species C. pachyzygosporus characterized with thick-walled zygospores is phylogenetically closely related to C. antarcticus, C. couchii, and C. osmodes.



Similar content being viewed by others
References
Bałazy S, Wiśniewski J, Kaczmarek S (1987) Some note worthy fungi occurring on mites. Bull Pol Ac: Biol 35:199–224
Bałazy S (1993) Entomophthorales: Flora of Poland (Flora Polska), Fungi (Mycota), vol 24. Polish Academy of Science, W. Szafer Inst. Botany, Kraeeków
Ben-Ze’ev IS, Kenneth RG (1982) Features-criteria of taxonomic value in the Entomophthorales: I. A revision of the Batkoan classification. Mycotaxon 14:393–455
Brefeld O (1884) Conidiobolus utriculosus und minor. Untersuchungen aus der Gesammtgebietee der Mykologie 6(2):35–78
Drechsler C (1952) Widespread distribution of Delacroixia coronata and other saprophytic Entomophthoraceae in plant detritus. Science 115:575–576. https://doi.org/10.1126/science.115.2995.575
Drechsler C (1953) Three new species of Conidiobolus isolated from leaf mold. J Wash Acad Sci 43(2):29–34
Drechsler C (1955) Three new species of Conidiobolus isolated from decaying plant detritus. Am J Bot 42(5):437–443. https://doi.org/10.1002/j.1537-2197.1955.tb11144.x
Drechsler C (1957) Two small species of Conidiobolus forming lateral zygospores. Bull Torrey Bot Club 84(4):268–280. https://doi.org/10.2307/2482673
Drechsler C (1960) Two new species of Conidiobolus found in plant detritus. Am J Bot 47:368–377. https://doi.org/10.1002/j.1537-2197.1960.tb07138.x
Felsenstein J (1985) Confidence limits on the bootstrap: an approach using the bootstrap. Evolution 38:783–791
Gryganskyi AP, Humber RA, Smith ME, Hodge K, Huang B, Voigt K, Vilgalys R (2013) Phylogenetic lineages in Entomophthoromycota. Persoonia 30:94–105. https://doi.org/10.3767/003158513X666330
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Hall TA (1999) Bioedit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Han ML, Chen YY, Shen LL, Song J, Vlasák J, Dai YC, Cui BK (2016) Taxonomy and phylogeny of the brown-rot fungi: Fomitopsis and its related genera. Fungal Divers 80:343–373. https://doi.org/10.1007/s13225-016-0364-y
Huang B, Humber RA, Hodge KT (2007) A new species of Conidiobolus from Great Smoky Mountains National Park. Mycotaxon 100:227–233
Humber RA (1981) An alternative view of certain taxonomic criteria used in the Entomophthorales (Zygomycetes). Mycotaxon 13:191–240
Humber RA (1989) Synopsis of a revised classification for the Entomophthorales (Zygomycotina). Mycotaxon 34:441–460
Humber RA (1997) Fungi: identification. In: Lacey LA (ed) Manual of techniques in insect pathology. Academic Press, London
Humber RA (2012) Entomophthoromycota: a new phylum and reclassification for entomophthoroid fungi. Mycotaxon 120:477–492. https://doi.org/10.5248/120.477
Jensen AB, Gargas A, Eilenberg J, Rosendahl S (1998) Relationships of the insect-pathogenic order Entomophthorales (Zygomycota, Fungi) based on phylogenetic analyses of nucleus small subunit ribosomal DNA sequences (SSU rDNA). Fungal Genet Biol 24(3):325–334. https://doi.org/10.1006/fgbi.1998.1063
Keller S (2008) The arthropod-pathogenic Entomophthorales from Switzerland – is central Europe the centre of their global species-richness? Mitt Schweiz Entomol Ges 81:39–51
King DS (1976a) Systematics of Conidiobolus (Entomophthorales) using numerical taxonomy I. Biology and cluster analysis. Can J Bot 54:45–65. https://doi.org/10.1139/b76-008
King DS (1976b) Systematics of Conidiobolus (Entomophthorales) using numerical taxonomy II. Taxonomic considerations. Can J Bot 54:1285–1296. https://doi.org/10.1139/b76-141
King DS (1977) Systematics of Conidiobolus (Entomophthorales) using numerical taxonomy III. Descriptions of recognized species. Can J Bot 55:718–729. https://doi.org/10.1139/b77-086
Kwon-Chung KJ (2012) Taxonomy of fungi causing mucormycosis and entomophthoramycosis (zygomycosis) and nomenclature of the disease: molecular mycologic perspectives. Clin Infect Dis 54:S8–S15. https://doi.org/10.1093/cid/cir864
Liu M, Rombach MC, Humber RA, Hodge KT (2005) What’s in a name? Aschersoniainsperata: a new pleoanamorphic fungus with characteristics of Aschersonia and Hirsutella. Mycologia 97:249–256. https://doi.org/10.3852/mycologia.97.1.246
Mendoza L, Vilela R, Voelz K, Ibrahim AS, Voigt K, Lee SC (2014) Human fungal pathogens of Mucorales and Entomophthorales. Cold Spring Harb Perspect Med 5(4):1–33. https://doi.org/10.1101/cshperspect.a019562
Nie Y, Yu CZ, Liu XY, Huang B (2012) A new species of Conidiobolus (Ancylistaceae) from Anhui, China. Mycotaxon 120:427–435. https://doi.org/10.5248/120.427
Nie Y, Tang XX, Liu XY, Huang B (2016) Conidiobolus stilbeus, a new species with mycelial strand and two types of primary conidiophores. Mycosphere 7(6):801–809. https://doi.org/10.5943/mycosphere/7/6/11
Nie Y, Tang XX, Liu XY, Huang B (2017) A new species of Conidiobolus with chlamdosporus from Dabie Mountains, eastern China. Mycosphere 8(7):809–816. https://doi.org/10.5943/mycosphere/8/7/1
Page RD (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818. https://doi.org/10.1093/bioinformatics/14.9.817
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Spatafora JW, Chang Y, Benny GL, Lazarus K, Smith ME, Berbee ML, Bonito G, Corradi N, Grigoriev I, Gryganskyi A, James TY, O’Donnell K, Roberson RW, Taylor TN, Uehling J, Vilgalys R, White MM, Stajich JE (2016) A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108(5):1028–1046. https://doi.org/10.3852/16-042
Srinivasan MC, Thirumalachar MJ (1961) Studies on species of Conidiobolus from India-I. Sydowia, Annales Mycologici 15:237–241
Srinivasan MC, Thirumalachar MJ (1962a) Studies on species of Conidiobolus from India-II. Sydowia, Annales Mycologici 16:60–66
Srinivasan MC, Thirumalachar MJ (1962b) Studies on species of Conidiobolus from India-III. Mycologia 54(6):685–693. https://doi.org/10.2307/3756504
Srinivasan MC, Thirumalachar MJ (1965) Studies on species of Conidiobolus from India-IV. Sydowia 19:86–91
Srinivasan MC, Thirumalachar MJ (1968a) Studies on species of Conidiobolus from India-V. Mycopathol Mycol Appl 36:341–346. https://doi.org/10.1007/BF02050380
Srinivasan MC, Thirumalachar MJ (1968b) Two new species of Conidiobolus from India. Journal of the Mitchell Society 84:211–212
Swofford DL (2002) PAUP*: Phylogenetic analysis using parsimony (*and other methods), Version 4.0b10. Sinauer Associates, Sunderland
Thompson JD, Gibson TJ, Plewniak F (1997) The Clustal-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 63:215–228
Vilela R, Silva SMS, Correa FR, Dominguez E, Mendoza L (2010) Morphologic and phylogenetic characterization of Conidiobolus lamprauges recovered from infected sheep. J Clin Microbiol 48:427–432. https://doi.org/10.1128/JCM.01589-09
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/JCM.01589-09
Waingankar VM, Singh SK, Srinivasan MC (2008) A new thermophilic species of Conidiobolus from India. Mycopathologia 165:173–177. https://doi.org/10.1007/s11046-007-9088-6
Wang CF, Li KP, Huang B (2010a) A new record to China — Conidiobolus iuxtagenitus. J Fungal Res 8:12–14
Wang CF, Li KP, Liu YJ, Li ZZ, Huang B (2010b) Three new Chinese records of Conidiobolus. Mycosystema 29:595–599
Watanabe M, Lee K, Goto K, Kumagai S, Sugita-Konishi Y, Hara-Kudo Y (2010) Rapid and effective DNA extraction method with bead grinding for a large amount of fungal DNA. J Food Prot 73(6):1077–1084. https://doi.org/10.4315/0362-028X-73.6.1077
Waters SD, Callaghan AA (1989) Conidiobolus iuxtagenitus, a new species with discharge delongate repetitional conidia and conjugation tubes. Mycol Res 93:223–226. https://doi.org/10.1016/S0953-7562(89)80121-2
Acknowledgements
We thank Dr. Rabern Simmons (University of Michigan) for improving the manuscript.
Funding
This project was supported by the National Natural Science Foundation of China (Nos. 30770008 and 31670019) and the Key Science Research Project of Anhui Province (No. TD200708).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Section Editor: Marco Thines
Rights and permissions
About this article
Cite this article
Nie, Y., Qin, L., Yu, DS. et al. Two new species of Conidiobolus occurring in Anhui, China. Mycol Progress 17, 1203–1211 (2018). https://doi.org/10.1007/s11557-018-1436-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-018-1436-z