Abstract
Purpose
Neoadjuvant chemotherapy (NAC) aims to minimize the tumor size before surgery. Predicting response to NAC could reduce toxicity and delays to effective intervention. Computational analysis of dynamic contrast-enhanced magnetic resonance images (DCE-MRI) through deep convolution neural network (CNN) has shown a significant performance to distinguish responders and no responder’s patients. This study intends to present a new deep learning (DL) model predicting the breast cancer response to NAC based on multiple MRI inputs.
Methods
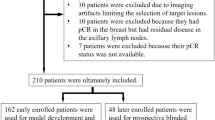
A cohort of 723 axial slices extracted from 42 breast cancer patients who underwent NAC therapy was used to train and validate the developed DL model. This dataset was provided by our collaborator institute of radiology in Brussels. Fourteen external cases were used to validate the best obtained model to predict pCR based on pre- and post-chemotherapy DCE-MRI. The model performance was assessed by area under the receiver operating characteristic curve (AUC), accuracy, sensitivity, specificity, and feature map visualization.
Results
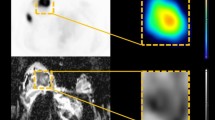
The developed multi-inputs deep learning architecture was able to predict the pCR to NAC treatment in the validation dataset with an AUC of 0.91 using combined pre- and post-NAC images. The visual results showed that the most important extracted features from non-pCR tumors are in the peripheral region. The proposed method was more productive than the previous ones.
Conclusion
Even with a limited training dataset size, the proposed and developed CNN model using DCE-MR images acquired before and after the first chemotherapy was able to classify pCR and non-pCR patients with substantial accuracy. This model could be used hereafter in clinical analysis after its evaluation based on more extra data.








Similar content being viewed by others
References
El Adoui M, Drisis S, Larhmam MA, Lemort M, Benjelloun M (2017) Breast cancer heterogeneity analysis as index of response to treatment using MRI images: a review. J Imaging Med 9(4):109–119
Drisis S, El Adoui M, Flamen P, Benjelloun M, Dewind R, Paesmans M, Ignatiadis M, Bali M, Lemort M (2019) Early prediction of neoadjuvant treatment outcome in locally advanced breast cancer using parametric response mapping and radial heterogeneity from breast MRI. J Magn Reson Imaging. https://doi.org/10.1002/jmri.26996
Meisamy S, Bolan PJ, Baker EH, Bliss RL, Gulbahce E, Everson LI, Nelson MT, Emory TH, Tuttle TM, Yee D, Garwood M (2004) Neoadjuvant chemotherapy of locally advanced breast cancer: predicting response with in vivo 1H MR spectroscopy—a pilot study at 4 T. Radiology 233(2):424–431
Liu SV, Melstrom L, Yao K, Russell CA, Sener SF (2010) Neoadjuvant therapy for breast cancer. J Surg Oncol 101(4):283–291
Kul S, Cansu A, Alhan E, Dinc H, Gunes G, Reis A (2011) Contribution of diffusion-weighted imaging to dynamic contrast-enhanced MRI in the characterization of breast tumors. Am J Roentgenol 196(1):210–217
Adoui ME, Drisis S, Benjelloun M (2017) Analyzing breast tumor heterogeneity to predict the response to chemotherapy using 3D MR images registration. In: Proceedings of the international conference on smart digital environment (ICSDE). ACM, pp 56–63
Theilmann RJ, Borders R, Trouard TP, Xia G, Outwater E, Ranger-Moore J, Gillies RJ, Stopeck A (2004) Changes in water mobility measured by diffusion MRI predict response of metastatic breast cancer to chemotherapy. Neoplasia 6(6):831
Rieber A, Brambs H-J, Gabelmann A, Heilmann V, Kreienberg R, Kühn T (2002) Breast MRI for monitoring response of primary breast cancer to neo-adjuvant chemotherapy. Eur Radiol 12(7):1711–1719
Sannachi L, Gangeh M, Tadayyon H, Sadeghi-Naini A, Gandhi S, Wright FC, Slodkowska E, Curpen B, Tran W, Czarnota GJ (2018) Response monitoring of breast cancer patients receiving neoadjuvant chemotherapy using quantitative ultrasound, texture, and molecular features. PLoS ONE 13(1):e0189634
Guenther LM, Rowe RG, Acharya PT, Swenson DW, Meyer SC, Clinton CM, Guo D, Sridharan M, London WB, Grier HE, Ecklund K (2018) Response Evaluation Criteria in Solid Tumors (RECIST) following neoadjuvant chemotherapy in osteosarcoma. Pediatr Blood Cancer 65(4):e26896
Cheung Y-C, Chen S-C, Su M-Y, See L-C, Hsueh S, Chang H-K, Lin Y-C, Tsai C-S (2003) Monitoring the size and response of locally advanced breast cancers to neoadjuvant chemotherapy (weekly paclitaxel and epirubicin) with serial enhanced MRI. Breast Cancer Res Treat 78(1):51–58
Cooper AB, Slack R, Fogelman D, Holmes HM, Petzel M, Parker N, Balachandran A, Garg N, Ngo-Huang A, Varadhachary G, Evans DB (2015) Characterization of anthropometric changes that occur during neoadjuvant therapy for potentially resectable pancreatic cancer. Ann Surg Oncol 22(7):2416–2423
El Adoui M, Drisis S, Benjelloun M (2018) A PRM approach for early prediction of breast cancer response to chemotherapy based on registered MR images. Int J Comput Assist Radiol Surg 13:1233–1243
El Adoui M, Larhmam MA, Drisis S, Benjelloun M (2019) Deep Learning approach predicting breast tumor response to neoadjuvant treatment using DCE-MRI volumes acquired before and after chemotherapy. In: SPIE medical imaging, 2019: computer-aided diagnosis, vol 10950, p 109502I. https://doi.org/10.1117/12.2505887
El Adoui M, Drisis S, Benjelloun M (2019) Predict breast tumor response to chemotherapy using a 3D deep learning architecture applied to DCE-MRI data. In: 7th international work-conference on bioinformatics and biomedical engineering, IWBBIO, 2019. Springer, Cham, pp 33–40. https://doi.org/10.1007/978-3-030-17935-9_4
Little RA, Barjat H, Hare JI, Jenner M, Watson Y, Cheung S, Holliday K, Zhang W, O’Connor JP, Barry ST, Puri S (2018) Evaluation of dynamic contrast-enhanced MRI biomarkers for stratified cancer medicine: how do permeability and perfusion vary between human tumours? Magn Reson Imaging 46:98–105
Drisis S, Metens T, Ignatiadis M, Stathopoulos K, Chao S-L, Lemort M (2016) Quantitative DCE-MRI for prediction of pathological complete response following neoadjuvant treatment for locally advanced breast cancer: the impact of breast cancer subtypes on the diagnostic accuracy. Eur Radiol 26(5):1474–1484
Drisis S, Flamen P, Ignatiadis M, Metens T, Chao S-L, Chintinne M, Lemort M (2018) Total choline quantification measured by 1H MR spectroscopy as early predictor of response after neoadjuvant treatment for locally advanced breast cancer: the impact of immunohistochemical status. J Magn Reson Imaging 48(4):982–993
Kerwin WS, Zhao X, Yuan C, Hatsukami TS, Maravilla KR, Underhill HR, Zhao X (2009) Contrast-enhanced MRI of carotid atherosclerosis: dependence on contrast agent. J Magn Resonan Imaging 30(1):35–40
Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA, Gee JC (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29(6):1310
El Adoui M, Mahmoudi SA, Larhmam MA, Benjelloun M (2019) MRI breast tumor segmentation using different encoder and decoder CNN architectures. Computers 8(3):52
Schlüter J, Grill T (2015) Exploring data augmentation for improved singing voice detection with neural networks. In: ISMIR, pp 121–126
Giusti A, Cireşan DC, Masci J, Gambardella LM, Schmidhuber J (2013) Fast image scanning with deep max-pooling convolutional neural networks. In: 2013 IEEE international conference on image processing. IEEE, pp 4034–4038
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. arXiv:1412.6980
Pham V, Bluche T, Kermorvant C, Louradour J (2014) Dropout improves recurrent neural networks for handwriting recognition. In: 2014 14th international conference on frontiers in handwriting recognition. IEEE, pp 285–290
Koutsoukas A, Monaghan KJ, Li X, Huan J (2017) Deep-learning: investigating deep neural networks hyper-parameters and comparison of performance to shallow methods for modeling bioactivity data. J Cheminform 9(1):42
Byrski T, Huzarski T, Dent R, Gronwald J, Zuziak D, Cybulski C, Kladny J, Gorski B, Lubinski J, Narod SA (2009) Response to neoadjuvant therapy with cisplatin in BRCA1-positive breast cancer patients. Breast Cancer Res Treat 115(2):359–363
Nachar N (2008) The Mann–Whitney U: a test for assessing whether two independent samples come from the same distribution. Tutor Quant Methods Psychol 4(1):13–20
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D (2017) Grad-cam: visual explanations from deep networks via gradient-based localization. In: Proceedings of the IEEE international conference on computer vision, pp 618–626
Fan M, Wu G, Cheng H, Zhang J, Shao G, Li L (2017) Radiomic analysis of DCE-MRI for prediction of response to neoadjuvant chemotherapy in breast cancer patients. Eur J Radiol 94:140–147
Chaplin DD (2010) Overview of the immune response. J Allergy Clin Immunol 125(2):S3–S23
Abraham DC, Jones RC, Jones SE, HaroldCheek J, Peters GN, Knox SM, Grant MD, Hampe DW, Savino DA, Harms SE (1996) Evaluation of neoadjuvant chemotherapeutic response of locally advanced breast cancer by magnetic resonance imaging. Cancer 78(1):91–100
Mahmoudi SA, El Adoui M, Belarbi MA, Larhmam MA, Lecron F (2017) Cloud-based platform for computer vision applications. In: Proceedings of the 2017 international conference on smart digital environment (ICSDE). ACM, pp 195–200
Acknowledgements
We would like to address our sincere acknowledgments to Dr. Marc Lemort, the head of the Radiology Department at the Jules Bordet Institute—Brussels, for offering the dataset used to evaluate our implemented models.
Funding
This research was financially supported by the University of Mons in Belgium.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
All procedures performed in studies involving human participants were following the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. For this type of study, formal consent is not required.
Informed consent
Informed consent was obtained from all individuals included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
El Adoui, M., Drisis, S. & Benjelloun, M. Multi-input deep learning architecture for predicting breast tumor response to chemotherapy using quantitative MR images. Int J CARS 15, 1491–1500 (2020). https://doi.org/10.1007/s11548-020-02209-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-020-02209-9