Abstract
Objectives
To evaluate potential relationship between qualitative CT features, quantitative texture analysis (QTA), histology, WHO staging, Masaoka classification and myasthenic syndrome in patients with thymic tumors.
Materials and methods
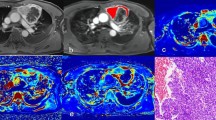
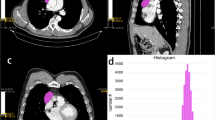
Sixteen patients affected by histologically proven thymic tumors were retrospectively included in the study population. Clinical information, with special regard to myasthenic syndrome and serological positivity of anti-AchR antibodies, were recorded. Qualitative CT evaluation included the following parameters: (a) location; (b) tumor edges; (c) necrosis; (d) pleural effusion; (e) metastases; (f) chest wall infiltration; (g) tumor margins. QTA included evaluation of “Mean” (M), “Standard Deviation” (SD), “Kurtosis” (K), “Skewness” (S), “Entropy” (E), “Shape from Texture” (TX_sigma) and “average of positive pixels” (MPP). Pearson–Rho test was used to evaluate the relationship of continuous non-dichotomic parameters, whereas Mann–Whitney test was used for dichotomic parameters.
Results
Histological evaluation demonstrated thymoma in 12 cases and thymic carcinoma in 4 cases. Tumor necrosis was significantly correlated with QTA Mean (p = 0.0253), MPP (p = 0.0417), S (p = 0.0488) and K (p = 0.0178). WHO staging was correlated with Mean (p = 0.0193), SD (p = 0.0191) and MPP (p = 0.0195). Masaoka classification was correlated with Mean (p = 0.0322), MPP (p = 0.0315), skewness (p = 0.0433) and Kurtosis (p = 0.0083). Myasthenic syndrome was significantly associated with Mean (p = 0.0211) and MPP (p = 0.0261), whereas tumor size was correlated with Mean (p = 0.0241), entropy (p = 0.0177), MPP (p = 0.0468), skewness (p = 0.009) and Kurtosis (p = 0.006).
Conclusion
Our study demonstrates significant relationship between radiomics parameters, histology, grading and clinical manifestations of thymic tumors.




Similar content being viewed by others
References
Colen R, Foster I, Gatenby R, Giger ME, Gillies R, Gutman D, Heller M, Jain R, Madabhushi A, Madhavan S, Napel S, Rao A, Saltz J, Tatum J, Verhaak R, Whitman G (2014) NCI workshop report: clinical and computational requirements for correlating imaging phenotypes with genomics signatures. Transl Oncol 7(5):556–569
Jain R, Poisson LM, Gutman D, Scarpace L, Hwang SN, Holder CA, Wintermark M, Rao A, Colen RR, Kirby J, Freymann J, Jaffe CC, Mikkelsen T, Flanders A (2014) Outcome prediction in patients with glioblastoma by using imaging, clinical, and genomic biomarkers: focus on the nonenhancing component of the tumor. Radiology 272(2):484–493
Prasanna P, Patel J, Partovi S (2016) Radiomic features from the peritumoral brain parenchyma on treatment-naïve multi-parametric MR imaging predict long versus short-term survival in glioblastoma multiforme: preliminary findings. Eur Radiol 27(10):4188–4197
Xi YB, Guo F, Xu ZL (2017) Radiomics signature: a potential biomarker for the prediction of MGMT promoter methylation in glioblastoma. J Magn Reson Imaging. https://doi.org/10.1002/jmri.25860
Karlo CA, Di Paolo PL, Chaim J, Hakimi AA, Ostrovnaya I, Russo P, Hricak H, Motzer R, Hsieh JJ, Akin O (2014) Radiogenomics of clear cell renal cell carcinoma: associations between CT imaging features and mutations. Radiology 270(2):464–471
Farber NJ, Kim CJ, Modi PK (2017) Renal cell carcinoma: the search for a reliable biomarker. ransl. Cancer Res 6(3):620–632
Rizzo S, Petrella F, Buscarino V, De Maria F, Raimondi S, Barberis M, Fumagalli C, Spitaleri G, Rampinelli C, De Marinis F, Spaggiari L, Bellomi M (2015) CT radiogenomic characterization of EGFR, K-RAS, and ALK mutations in non-small cell lung cancer. Eur Radiol 26(1):32–42
Buettner R, Wolf J, Thomas RK (2013) Lessons learned from lung cancer genomics: the emerging concept of individualized diagnostics and treatment. J Clin Oncol 31(15):1858–1865
Dehmeshki J, Amin H, Valdivieso M, Ye X (2008) Segmentation of pulmonary nodules in thoracic CT scans: a region growing approach. IEEE Trans Med Imaging 27(4):467–480. https://doi.org/10.1109/TMI.2007.907555
De Cecco CN, Ganeshan B, Ciolina M, Rengo M, Meinel FG, Musio D, De Felice F, Raffetto N, Tombolini V, Laghi A (2015) Texture analysis as imaging biomarker of tumoral response to neoadjuvant chemoradiotherapy in rectal cancer patients studied with 3-T magnetic resonance. Invest Radiol 50(4):239–245
Sacconi B, Anzidei M, Leonardi A (2017) Analysis of CT features and quantitative texture analysis in patients with lung adenocarcinoma: a correlation with EGFR mutations and survival rates. Clin Radiol 72(6):443–450
Lambin P, Rios-Velazquez E, Leijenaar R, Carvalho S, Stiphout RV, Granton P, Zeghers CM, Gillies R, Boellard R, Dekker A, Aerts HJ (2011) Radiomics: extracting more information from medical images using advance feature analysis. Eur J Cancer 48(4):441–446
Jaffe CC (2006) Measures of response: RECIST, WHO, and new alternatives. J Clin Oncol 24(20):3245–3251
Janssen MH, Ollers MC, van Stiphout RG, Riedl RG, van den Bogaard J, Buijsen J, Lambin P, Lammering G (2010) Blood glucose level normalization and accurate timing improves the accuracy of PET-based treatment response predictions in rectal cancer. Radiother Oncol 95(2):203–208
Rubin DL (2008) Creating and curating a terminology for radiology: ontology modeling and analysis. J Digit Imaging 21(4):355–362
Gonzales RC, Woods RE (2008) Digital image processing. Prentice Hall, Upper Saddle River
Chen CH, Pau LF, Wang PSP (eds) (1998) The handbook of pattern recognition and computer vision, 2nd edn. World Scientific Publishing Co., Singapore, pp 207–248
Venuta F, Anile M et al (2010) Thymoma and thymic carcinoma. Review. Eur J Cardio-Thorac Surg 37:13–25
Krishnan K, Ibanez L, Turner WD, Jomier J (2010) Avila RS. An open-source toolkit for the volumetric measurement of CT lung lesions. Opt Express 18(14):15256–15266
Boykov Y, Veksler O, Zabih R (2001) Fast approximate energy minimization via graph cuts. IEEE Trans Pattern Anal Mach Intell 23(11):1222–1239
Funding
No funding was received.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Iannarelli, A., Sacconi, B., Tomei, F. et al. Analysis of CT features and quantitative texture analysis in patients with thymic tumors: correlation with grading and staging. Radiol med 123, 345–350 (2018). https://doi.org/10.1007/s11547-017-0845-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11547-017-0845-4