Abstract
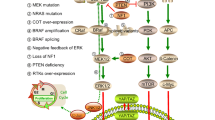
Malignant melanoma is the most aggressive and notorious skin cancer, and metastatic disease is associated with very poor long-term survival outcomes. Although metastatic melanoma patients with oncogenic mutations in the BRAF gene initially respond well to the treatment with specific BRAF inhibitors, most of them will eventually develop resistance to this targeted therapy. As a highly conserved catabolic process, autophagy is responsible for the maintenance of cellular homeostasis and cell survival, and is involved in multiple diseases, including cancer. Recent study results have indicated that autophagy might play a decisive role in the resistance to BRAF inhibitors in BRAF-mutated melanomas. In this review, we will discuss how autophagy is up-regulated by BRAF inhibitors, and how autophagy induces the resistance to these agents.

Similar content being viewed by others
References
Meng XX, Xu HX, Yao M, et al. Implication of unfolded protein response and autophagy in the treatment of BRAF inhibitor resistant melanoma. Anti Cancer Agents Med Chem. 2016;16(3):291–8.
Spagnolo F, Ghiorzo P, Orgiano L, et al. BRAF-mutant melanoma: treatment approaches, resistance mechanisms, and diagnostic strategies. Onco Targets Ther. 2015;8:157–68.
Rastrelli M, Tropea S, Rossi CR, et al. Melanoma: epidemiology, risk factors, pathogenesis, diagnosis and classification. In Vivo. 2014;28(6):1005–11.
Luke JJ, Flaherty KT, Ribas A, et al. Targeted agents and immunotherapies: optimizing outcomes in melanoma. Nat Rev Clin Oncol. 2017;14(8):463–82.
Atkins MB, Lotze MT, Dutcher JP, et al. High-dose recombinant interleukin 2 therapy for patients with metastatic melanoma: analysis of 270 patients treated between 1985 and 1993. J Clin Oncol. 1999;17(7):2105–16.
Schiller JH, Pugh M, Kirkwood JM, et al. Eastern cooperative group trial of interferon gamma in metastatic melanoma: an innovative study design. Clin Cancer Res. 1996;2(1):29–36.
Parker BS, Rautela J, Hertzog PJ. Antitumour actions of interferons: implications for cancer therapy. Nat Rev Cancer. 2016;16(3):131–44.
Rozeman EA, Dekker T, Haanen J, et al. Advanced melanoma: current treatment options, biomarkers, and future perspectives. Am J Clin Dermatol. 2017; https://doi.org/10.1007/s40257-017-0325-6.
Olszanski AJ, Hoffner BW. Evolving paradigms in melanoma therapy. J Adv Pract Oncol. 2016;7(3):291–4.
Bradish JR, Montironi R, Lopez-Beltran A, et al. Towards personalized therapy for patients with malignant melanoma: molecular insights into the biology of BRAF mutations. Future Oncol. 2013;9(2):245–53.
Ossio R, Roldan-Marin R, Martinez-Said H, et al. Melanoma: a global perspective. Nat Rev Cancer. 2017;17(7):393–4.
Schadendorf D, Fisher DE, Garbe C, et al. Melanoma. Nat Rev Dis Primers. 2015;1:15003.
Su MY, Fisher DE. Immunotherapy in the precision medicine era: melanoma and beyond. PLoS Med. 2016;13(12):e1002196.
Merlino G, Herlyn M, Fisher DE, et al. The state of melanoma: challenges and opportunities. Pigment Cell Melanoma Res. 2016;29(4):404–16.
Lawrence MS, Stojanov P, Polak P, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499(7457):214–8.
Chen SH, Gong X, Zhang Y, et al. RAF inhibitor LY3009120 sensitizes RAS or BRAF mutant cancer to CDK4/6 inhibition by abemaciclib via superior inhibition of phospho-RB and suppression of cyclin D1. Oncogene. 2018;37(6):821–32.
Colombino M, Capone M, Lissia A, et al. BRAF/NRAS mutation frequencies among primary tumors and metastases in patients with melanoma. J Clin Oncol. 2012;30(20):2522–9.
Curtin JA, Fridlyand J, Kageshita T, et al. Distinct sets of genetic alterations in melanoma. N Engl J Med. 2005;353(20):2135–47.
Hayward NK, Wilmott JS, Waddell N, et al. Whole-genome landscapes of major melanoma subtypes. Nature. 2017;545(7653):175–80.
Bradish JR, Cheng L. Molecular pathology of malignant melanoma: changing the clinical practice paradigm toward a personalized approach. Hum Pathol. 2014;45(7):1315–26.
Cheng L, Lopez-Beltran A, Massari F, et al. Molecular testing for BRAF mutations to inform melanoma treatment decisions: a move toward precision medicine. Mod Pathol. 2018;31(1):24–38.
Burotto M, Chiou VL, Lee JM, et al. The MAPK pathway across different malignancies: a new perspective. Cancer. 2014;120(22):3446–56.
Arozarena I, Wellbrock C. Overcoming resistance to BRAF inhibitors. Ann Transl Med. 2017;5(19):387.
Bollag G, Hirth P, Tsai J, et al. Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature. 2010;467(7315):596–9.
Waizenegger IC, Baum A, Steurer S, et al. A novel RAF kinase inhibitor with DFG-out-binding mode: high efficacy in BRAF-mutant tumor xenograft models in the absence of normal tissue Hyperproliferation. Mol Cancer Ther. 2016;15(3):354–65.
Dean L. Vemurafenib therapy and BRAF and NRAS genotype. In: Pratt V, McLeod H, Dean L, et al., editors. Medical genetics summaries [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2012.
Flaherty KT, Puzanov I, Kim KB, et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med. 2010;363(9):809–19.
Sosman JA, Kim KB, Schuchter L, et al. Survival in BRAF V600-mutant advanced melanoma treated with vemurafenib. N Engl J Med. 2012;366(8):707–14.
McArthur GA, Chapman PB, Robert C, et al. Safety and efficacy of vemurafenib in BRAF(V600E) and BRAF(V600K) mutation-positive melanoma (BRIM-3): extended follow-up of a phase 3, randomised, open-label study. Lancet Oncol. 2014;15(3):323–32.
Banzi M, De Blasio S, Lallas A, et al. Dabrafenib: a new opportunity for the treatment of BRAF V600-positive melanoma. Onco Targets Ther. 2016;9:2725–33.
Hauschild A, Grob JJ, Demidov LV, et al. Dabrafenib in BRAF-mutated metastatic melanoma: a multicentre, open-label, phase 3 randomised controlled trial. Lancet. 2012;380(9839):358–65.
Long GV, Weber JS, Infante JR, et al. Overall survival and durable responses in patients with BRAF V600-mutant metastatic melanoma receiving Dabrafenib combined with Trametinib. J Clin Oncol. 2016;34(8):871–8.
Turajlic S, Furney SJ, Stamp G, et al. Whole-genome sequencing reveals complex mechanisms of intrinsic resistance to BRAF inhibition. Ann Oncol. 2014;25(5):959–67.
Shi H, Hugo W, Kong X, et al. Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy. Cancer Discov. 2014;4(1):80–93.
Van Allen EM, Wagle N, Sucker A, et al. The genetic landscape of clinical resistance to RAF inhibition in metastatic melanoma. Cancer Discov. 2014;4(1):94–109.
Nazarian R, Shi H, Wang Q, et al. Melanomas acquire resistance to B-RAF(V600E) inhibition by RTK or N-RAS upregulation. Nature. 2010;468(7326):973–7.
Whittaker SR, Theurillat JP, Van Allen E, et al. A genome-scale RNA interference screen implicates NF1 loss in resistance to RAF inhibition. Cancer Discov. 2013;3(3):350–62.
Girotti MR, Pedersen M, Sanchez-Laorden B, et al. Inhibiting EGF receptor or SRC family kinase signaling overcomes BRAF inhibitor resistance in melanoma. Cancer Discov. 2013;3(2):158–67.
Villanueva J, Vultur A, Lee JT, et al. Acquired resistance to BRAF inhibitors mediated by a RAF kinase switch in melanoma can be overcome by cotargeting MEK and IGF-1R/PI3K. Cancer Cell. 2010;18(6):683–95.
Ascierto PA, McArthur GA, Dreno B, et al. Cobimetinib combined with vemurafenib in advanced BRAF(V600)-mutant melanoma (coBRIM): updated efficacy results from a randomised, double-blind, phase 3 trial. Lancet Oncol. 2016;17(9):1248–60.
Long GV, Flaherty KT, Stroyakovskiy D, et al. Dabrafenib plus trametinib versus dabrafenib monotherapy in patients with metastatic BRAF V600E/K-mutant melanoma: long-term survival and safety analysis of a phase 3 study. Ann Oncol. 2017;28(7):1631–9.
Brighton HE, Angus SP, Bo T, et al. New mechanisms of resistance to MEK inhibitors in melanoma revealed by intravital imaging. Cancer Res. 2018;78(2):542–57.
Shi H, Hong A, Kong X, et al. A novel AKT1 mutant amplifies an adaptive melanoma response to BRAF inhibition. Cancer Discov. 2014;4(1):69–79.
Muller J, Krijgsman O, Tsoi J, et al. Low MITF/AXL ratio predicts early resistance to multiple targeted drugs in melanoma. Nat Commun. 2014;5:5712.
Smith MP, Brunton H, Rowling EJ, et al. Inhibiting drivers of non-mutational drug tolerance is a salvage strategy for targeted melanoma therapy. Cancer Cell. 2016;29(3):270–84.
Straussman R, Morikawa T, Shee K, et al. Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature. 2012;487(7408):500–4.
Hirata E, Girotti MR, Viros A, et al. Intravital imaging reveals how BRAF inhibition generates drug-tolerant microenvironments with high integrin beta1/FAK signaling. Cancer Cell. 2015;27(4):574–88.
Smith MP, Sanchez-Laorden B, O'Brien K, et al. The immune microenvironment confers resistance to MAPK pathway inhibitors through macrophage-derived TNFalpha. Cancer Discov. 2014;4(10):1214–29.
Wang T, Xiao M, Ge Y, et al. BRAF inhibition stimulates melanoma-associated macrophages to drive tumor growth. Clin Cancer Res. 2015;21(7):1652–64.
Klionsky DJ. Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol. 2007;8(11):931–7.
Mizushima N, Klionsky DJ. Protein turnover via autophagy: implications for metabolism. Annu Rev Nutr. 2007;27:19–40.
Yu L, Chen Y, Tooze SA. Autophagy pathway: cellular and molecular mechanisms. Autophagy 2017:0.
Gallagher LE, Chan EY. Early signalling events of autophagy. Essays Biochem. 2013;55:1–15.
Axe EL, Walker SA, Manifava M, et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J Cell Biol. 2008;182(4):685–701.
Ganley IG, Lam DH, Wang J, et al. ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy. J Biol Chem. 2009;284(18):12297–305.
Itakura E, Kishi C, Inoue K, et al. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol Biol Cell. 2008;19(12):5360–72.
Zhong Y, Wang QJ, Li X, et al. Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1-phosphatidylinositol-3-kinase complex. Nat Cell Biol. 2009;11(4):468–76.
Sun Q, Fan W, Chen K, et al. Identification of Barkor as a mammalian autophagy-specific factor for Beclin 1 and class III phosphatidylinositol 3-kinase. Proc Natl Acad Sci U S A. 2008;105(49):19211–6.
Chan EY, Longatti A, McKnight NC, et al. Kinase-inactivated ULK proteins inhibit autophagy via their conserved C-terminal domains using an Atg13-independent mechanism. Mol Cell Biol. 2009;29(1):157–71.
Suzuki SW, Yamamoto H, Oikawa Y, et al. Atg13 HORMA domain recruits Atg9 vesicles during autophagosome formation. Proc Natl Acad Sci U S A. 2015;112(11):3350–5.
Lamb CA, Nuhlen S, Judith D, et al. TBC1D14 regulates autophagy via the TRAPP complex and ATG9 traffic. EMBO J. 2016;35(3):281–301.
Webster CP, Smith EF, Bauer CS, et al. The C9orf72 protein interacts with Rab1a and the ULK1 complex to regulate initiation of autophagy. EMBO J. 2016;35(15):1656–76.
Lin MG, Hurley JH. Structure and function of the ULK1 complex in autophagy. Curr Opin Cell Biol. 2016;39:61–8.
Mao K, Klionsky DJ. AMPK activates autophagy by phosphorylating ULK1. Circ Res. 2011;108(7):787–8.
Kim J, Kundu M, Viollet B, et al. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13(2):132–41.
Jung CH, Jun CB, Ro SH, et al. ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell. 2009;20(7):1992–2003.
Backer JM. The intricate regulation and complex functions of the class III phosphoinositide 3-kinase Vps34. Biochem J. 2016;473(15):2251–71.
Lamb CA, Yoshimori T, Tooze SA. The autophagosome: origins unknown, biogenesis complex. Nat Rev Mol Cell Biol. 2013;14(12):759–74.
Fimia GM, Di Bartolomeo S, Piacentini M, et al. Unleashing the Ambra1-Beclin 1 complex from dynein chains: Ulk1 sets Ambra1 free to induce autophagy. Autophagy. 2011;7(1):115–7.
Hurley JH, Young LN. Mechanisms of autophagy initiation. Annu Rev Biochem. 2017;86:225–44.
Lang T, Reiche S, Straub M, et al. Autophagy and the cvt pathway both depend on AUT9. J Bacteriol. 2000;182(8):2125–33.
Yamamoto H, Kakuta S, Watanabe TM, et al. Atg9 vesicles are an important membrane source during early steps of autophagosome formation. J Cell Biol. 2012;198(2):219–33.
Jin M, Klionsky DJ. Transcriptional regulation of ATG9 by the Pho23-Rpd3 complex modulates the frequency of autophagosome formation. Autophagy. 2014;10(9):1681–2.
Hanada T, Noda NN, Satomi Y, et al. The Atg12-Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J Biol Chem. 2007;282(52):37298–302.
Fujita N, Itoh T, Omori H, et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell. 2008;19(5):2092–100.
Pierdominici M, Vomero M, Barbati C, et al. Role of autophagy in immunity and autoimmunity, with a special focus on systemic lupus erythematosus. FASEB J. 2012;26(4):1400–12.
Kim SE, Park HJ, Jeong HK, et al. Autophagy sustains the survival of human pancreatic cancer PANC-1 cells under extreme nutrient deprivation conditions. Biochem Biophys Res Commun. 2015;463(3):205–10.
Noman MZ, Berchem G, Janji B. Targeting autophagy blocks melanoma growth by bringing natural killer cells to the tumor battlefield. Autophagy. 2018:1–3.
Rangwala R, Chang YC, Hu J, et al. Combined MTOR and autophagy inhibition: phase I trial of hydroxychloroquine and temsirolimus in patients with advanced solid tumors and melanoma. Autophagy. 2014;10(8):1391–402.
Saglar E, Unlu S, Babalioglu I, et al. Assessment of ER stress and autophagy induced by ionizing radiation in both radiotherapy patients and ex vivo irradiated samples. J Biochem Mol Toxicol. 2014;28(9):413–7.
Mukubou H, Tsujimura T, Sasaki R, et al. The role of autophagy in the treatment of pancreatic cancer with gemcitabine and ionizing radiation. Int J Oncol. 2010;37(4):821–8.
Galluzzi L, Pietrocola F, Bravo-San PJ, et al. Autophagy in malignant transformation and cancer progression. EMBO J. 2015;34(7):856–80.
Menzies FM, Fleming A, Rubinsztein DC. Compromised autophagy and neurodegenerative diseases. Nat Rev Neurosci. 2015;16(6):345–57.
Mizushima N, Komatsu M. Autophagy: renovation of cells and tissues. Cell. 2011;147(4):728–41.
Gomes LC, Dikic I. Autophagy in antimicrobial immunity. Mol Cell. 2014;54(2):224–33.
Ji C, Zhang Z, Chen L, et al. Endoplasmic reticulum stress-induced autophagy determines the susceptibility of melanoma cells to dabrafenib. Drug Des Devel Ther. 2016;10:2491–8.
Martin S, Dudek-Peric AM, Garg AD, et al. An autophagy-driven pathway of ATP secretion supports the aggressive phenotype of BRAF(V600E) inhibitor-resistant metastatic melanoma cells. Autophagy. 2017;13(9):1512–27.
Ma XH, Piao SF, Dey S, et al. Targeting ER stress-induced autophagy overcomes BRAF inhibitor resistance in melanoma. J Clin Invest. 2014;124(3):1406–17.
Martin S, Dudek-Peric AM, Maes H, et al. Concurrent MEK and autophagy inhibition is required to restore cell death associated danger-signalling in Vemurafenib-resistant melanoma cells. Biochem Pharmacol. 2015;93(3):290–304.
Li Z, Jiang K, Zhu X, et al. Encorafenib (LGX818), a potent BRAF inhibitor, induces senescence accompanied by autophagy in BRAFV600E melanoma cells. Cancer Lett. 2016;370(2):332–44.
Goodall ML, Wang T, Martin KR, et al. Development of potent autophagy inhibitors that sensitize oncogenic BRAF V600E mutant melanoma tumor cells to vemurafenib. Autophagy. 2014;10(6):1120–36.
Wang W, Kang H, Zhao Y, et al. Targeting autophagy sensitizes BRAF-mutant thyroid Cancer to Vemurafenib. J Clin Endocrinol Metab. 2017;102(2):634–43.
Goulielmaki M, Koustas E, Moysidou E, et al. BRAF associated autophagy exploitation: BRAF and autophagy inhibitors synergise to efficiently overcome resistance of BRAF mutant colorectal cancer cells. Oncotarget. 2016;7(8):9188–221.
Mulcahy LJ, Zahedi S, Griesinger AM, et al. Autophagy inhibition overcomes multiple mechanisms of resistance to BRAF inhibition in brain tumors. eLife. 2017;6:e19671.
Ge L, Zhang M, Schekman R. Phosphatidylinositol 3-kinase and COPII generate LC3 lipidation vesicles from the ER-Golgi intermediate compartment. eLife. 2014;3:e4135.
Luo M, Wu L, Zhang K, et al. miR-216b enhances the efficacy of vemurafenib by targeting Beclin-1, UVRAG and ATG5 in melanoma. Cell Signal. 2017;42:30–43.
Chen N, Karantza V. Autophagy as a therapeutic target in cancer. Cancer Biol Ther. 2011;11(2):157–68.
Kaufman RJ. Stress signaling from the lumen of the endoplasmic reticulum: coordination of gene transcriptional and translational controls. Genes Dev. 1999;13(10):1211–33.
Jhaveri KD, Sakhiya V, Fishbane S. Nephrotoxicity of the BRAF inhibitors Vemurafenib and Dabrafenib. JAMA Oncol. 2015;1(8):1133–4.
B'Chir W, Maurin AC, Carraro V, et al. The eIF2alpha/ATF4 pathway is essential for stress-induced autophagy gene expression. Nucleic Acids Res. 2013;41(16):7683–99.
Yang IV, Lozupone CA, Schwartz DA. The environment, epigenome, and asthma. J Allergy Clin Immunol. 2017;140(1):14–23.
Grunt TW. Interacting Cancer machineries: cell signaling, lipid metabolism, and epigenetics. Trends Endocrinol Metab. 2017;
Kim JH, Ahn JH, Lee M. Upregulation of MicroRNA-1246 is associated with BRAF inhibitor resistance in melanoma cells with mutant BRAF. Cancer Res Treat. 2017;49(4):947–59.
Flockhart RJ, Webster DE, Qu K, et al. BRAFV600E remodels the melanocyte transcriptome and induces BANCR to regulate melanoma cell migration. Genome Res. 2012;22(6):1006–14.
Davies H, Bignell GR, Cox C, et al. Mutations of the BRAF gene in human cancer. Nature. 2002;417(6892):949–54.
Wang Y, Guo Q, Zhao Y, et al. BRAF-activated long non-coding RNA contributes to cell proliferation and activates autophagy in papillary thyroid carcinoma. Oncol Lett. 2014;8(5):1947–52.
Michaud M, Martins I, Sukkurwala AQ, et al. Autophagy-dependent anticancer immune responses induced by chemotherapeutic agents in mice. Science. 2011;334(6062):1573–7.
Martins I, Wang Y, Michaud M, et al. Molecular mechanisms of ATP secretion during immunogenic cell death. Cell Death Differ. 2014;21(1):79–91.
Robertson GP. Functional and therapeutic significance of Akt deregulation in malignant melanoma. Cancer Metastasis Rev. 2005;24(2):273–85.
Zhao Y, Wang W, Min I, et al. BRAF V600E-dependent role of autophagy in uveal melanoma. J Cancer Res Clin Oncol. 2017;143(3):447–55.
Garg AD, Galluzzi L, Apetoh L, et al. Molecular and translational classifications of DAMPs in immunogenic cell death. Front Immunol. 2015;6:588.
Stagg J, Smyth MJ. Extracellular adenosine triphosphate and adenosine in cancer. Oncogene. 2010;29(39):5346–58.
Qiu Y, Li WH, Zhang HQ, et al. P2X7 mediates ATP-driven invasiveness in prostate cancer cells. PLoS ONE. 2014;9(12):e114371.
Zimmerer RM, Korn P, Demougin P, et al. Functional features of cancer stem cells in melanoma cell lines. Cancer Cell Int. 2013;13(1):78.
Wei Q, Zhang Y, Sun L, et al. High dose of extracellular ATP switched autophagy to apoptosis in anchorage-dependent and anchorage-independent hepatoma cells. Purinergic Signal. 2013;9(4):585–98.
Young CN, Sinadinos A, Lefebvre A, et al. A novel mechanism of autophagic cell death in dystrophic muscle regulated by P2RX7 receptor large-pore formation and HSP90. Autophagy. 2015;11(1):113–30.
Orioli E, De Marchi E, Giuliani AL, et al. P2X7 receptor orchestrates multiple Signalling pathways triggering inflammation, autophagy and metabolic/trophic responses. Curr Med Chem. 2017;24(21):2261–75.
Kepp O, Senovilla L, Vitale I, et al. Consensus guidelines for the detection of immunogenic cell death. Oncoimmunology. 2014;3(9):e955691.
Buzzi N, Bilbao PS, Boland R, et al. Extracellular ATP activates MAP kinase cascades through a P2Y purinergic receptor in the human intestinal Caco-2 cell line. Biochim Biophys Acta. 2009;1790(12):1651–9.
Chang SJ, Tzeng CR, Lee YH, et al. Extracellular ATP activates the PLC/PKC/ERK signaling pathway through the P2Y2 purinergic receptor leading to the induction of early growth response 1 expression and the inhibition of viability in human endometrial stromal cells. Cell Signal. 2008;20(7):1248–55.
Hill LM, Gavala ML, Lenertz LY, et al. Extracellular ATP may contribute to tissue repair by rapidly stimulating purinergic receptor X7-dependent vascular endothelial growth factor release from primary human monocytes. J Immunol. 2010;185(5):3028–34.
Xu Y, Li N, Xiang R, et al. Emerging roles of the p38 MAPK and PI3K/AKT/mTOR pathways in oncogene-induced senescence. Trends Biochem Sci. 2014;39(6):268–76.
Courtois-Cox S, Jones SL, Cichowski K. Many roads lead to oncogene-induced senescence. Oncogene. 2008;27(20):2801–9.
Lin AW, Barradas M, Stone JC, et al. Premature senescence involving p53 and p16 is activated in response to constitutive MEK/MAPK mitogenic signaling. Genes Dev. 1998;12(19):3008–19.
Michaloglou C, Vredeveld LC, Soengas MS, et al. BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature. 2005;436(7051):720–4.
Ryder M, Gild M, Hohl TM, et al. Genetic and pharmacological targeting of CSF-1/CSF-1R inhibits tumor-associated macrophages and impairs BRAF-induced thyroid cancer progression. PLoS ONE. 2013;8(1):e54302.
Zou M, Baitei EY, Al-Rijjal RA, et al. TSH overcomes Braf(V600E)-induced senescence to promote tumor progression via downregulation of p53 expression in papillary thyroid cancer. Oncogene. 2016;35(15):1909–18.
Vredeveld LC, Possik PA, Smit MA, et al. Abrogation of BRAFV600E-induced senescence by PI3K pathway activation contributes to melanomagenesis. Genes Dev. 2012;26(10):1055–69.
Liu H, He Z, Simon HU. Autophagy suppresses melanoma tumorigenesis by inducing senescence. Autophagy. 2014;10(2):372–3.
Liu H, He Z, von Rutte T, et al. Down-regulation of autophagy-related protein 5 (ATG5) contributes to the pathogenesis of early-stage cutaneous melanoma. Sci Transl Med. 2013;5(202):123r–202r.
Xie X, Koh JY, Price S, et al. Atg7 overcomes senescence and promotes growth of BrafV600E-driven melanoma. Cancer Discov. 2015;5(4):410–23.
Schlegel J, Sambade MJ, Sather S, et al. MERTK receptor tyrosine kinase is a therapeutic target in melanoma. J Clin Invest. 2013;123(5):2257–67.
Xue G, Kohler R, Tang F, et al. mTORC1/autophagy-regulated MerTK in mutant BRAFV600 melanoma with acquired resistance to BRAF inhibition. Oncotarget. 2017;8(41):69204–18.
Han J, Bae J, Choi CY, et al. Autophagy induced by AXL receptor tyrosine kinase alleviates acute liver injury via inhibition of NLRP3 inflammasome activation in mice. Autophagy. 2016;12(12):2326–43.
Brosius LA, Chung WS, Sloan SA, et al. Schwann cells use TAM receptor-mediated phagocytosis in addition to autophagy to clear myelin in a mouse model of nerve injury. Proc Natl Acad Sci U S A. 2017;114(38):E8072–80.
Solomon VR, Lee H. Chloroquine and its analogs: a new promise of an old drug for effective and safe cancer therapies. Eur J Pharmacol. 2009;625(1–3):220–33.
Thome R, Lopes SC, Costa FT, et al. Chloroquine: modes of action of an undervalued drug. Immunol Lett. 2013;153(1–2):50–7.
Wolpin BM, Rubinson DA, Wang X, et al. Phase II and pharmacodynamic study of autophagy inhibition using hydroxychloroquine in patients with metastatic pancreatic adenocarcinoma. Oncologist. 2014;19(6):637–8.
Boone BA, Bahary N, Zureikat AH, et al. Safety and biologic response of pre-operative autophagy inhibition in combination with gemcitabine in patients with pancreatic adenocarcinoma. Ann Surg Oncol. 2015;22(13):4402–10.
Monma H, Iida Y, Moritani T, et al. Chloroquine augments TRAIL-induced apoptosis and induces G2/M phase arrest in human pancreatic cancer cells. PLoS ONE. 2018;13(3):e193990.
Li ML, Xu YZ, Lu WJ, et al. Chloroquine potentiates the anticancer effect of sunitinib on renal cell carcinoma by inhibiting autophagy and inducing apoptosis. Oncol Lett. 2018;15(3):2839–46.
Cai Y, Cai J, Ma Q, et al. Chloroquine affects autophagy to achieve an anticancer effect in EC109 esophageal carcinoma cells in vitro. Oncol Lett. 2018;15(1):1143–8.
Funding
No external funding was used in the preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Xiao Liu, Jinfeng Wu, Haihong Qin, and Jinhua Xu declare that they have no conflicts of interest that might be relevant to the contents of this manuscript.
Rights and permissions
About this article
Cite this article
Liu, X., Wu, J., Qin, H. et al. The Role of Autophagy in the Resistance to BRAF Inhibition in BRAF-Mutated Melanoma. Targ Oncol 13, 437–446 (2018). https://doi.org/10.1007/s11523-018-0565-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-018-0565-2