Abstract
A new Pseudomonas putida strain (AQ8) was isolated from a decommissioned oil refinery’s soil in Italy and characterized for its ability to degrade BTEX. The draft genome of the new strain was sequenced and annotated for genes that encode enzymes putatively involved in BTEX degradation and quorum sensing. The strain was transformed with a plasmid expressing lactonase, which cleaves the autoinducer quorum sensing signal molecule, the acyl-homoserine lactone, to obtain a quorum sensing minus strain. P. putida AQ8 depleted the 40% on average of all the components of the initial BTEX concentration in 36 h. The quorum sensing minus strain, in the same time interval, depleted only the 10% of the initial BTEX concentration. The role of quorum sensing in regulating the expression of the annotated benzene/toluene dioxygenase gene (benzA) and biphenyl/toluene/benzene dioxygenase (bphA) genes, which are involved in BTEX degradation, was studied by quantitative RT-real-time quantitative (q)PCR analysis. The qPCR data showed decreased levels of expression of the benzA and bphA genes in the quorum sensing minus strain. Our results showed, for the first time, quorum sensing modulation of the level of transcription of dioxygenase genes in the upper BTEX oxidation pathway.
Similar content being viewed by others
Introduction
In Europe, soils and sediments are mainly polluted by organic pollutants, such as total petroleum hydrocarbons (TPH), polycyclic aromatic hydrocarbons (PAH), and monoaromatic hydrocarbons (benzene, toluene, ethylbenzene, xylenes) (European Environment Agency: http://www.eea.europa.eu 2012). Their presence in the environment is mainly due to activities of the chemical processing industry, leaks from underground hydrocarbon storage tanks, and accidental spills (Bowlen and Kosson 1995). Hydrocarbon compounds are recalcitrant to natural biodegradation processes in the environment, due to their aromatic structure, which conveys them with high electrochemical stability (Mackay and Callcott 1998; Marston et al. 2001). Hydrocarbons can bioaccumulate in adipose tissues and spread throughout whole food chains and ecosystems (Morehead et al. 1986; Xue and Warshawsky 2005).
BTEX, a mixture of benzene, toluene, ethylbenzene, and xylenes, which are present normally in diesel oil, readily volatizes into the atmosphere and may be distributed over large regions by wind (Durmusoglu et al. 2010; Mohammad et al. 2007). The monoaromatic hydrocarbon components of BTEX have been identified as potent carcinogens by the World Health Organization, but they can be widely found in the general environment (Horemans et al. 2008). In fact, they are the most abundant volatile organic compounds (VOC) species in the atmosphere and surface water (Liu et al. 2008a, b, c). This results in both direct and indirect contact with humans. Acute contacts in aquatic environments or inhalation are potentially toxic, while chronic contacts can lead to neurological, respiratory, genetic, and excretory system damage and to cancer (Irwin 1997).
Pollution of the environment by high concentrations of hydrocarbon compounds can select for microorganisms capable of utilizing them as sole carbon sources, eventually resulting in degradation of these compounds. The isolation of these hydrocarbonoclastic microorganisms from heavily polluted matrices ends up with the isolation of exploitable microbial strains for the depletion of contaminants in the matrices of origin and in different matrices (Becarelli et al. 2019; Di Gregorio et al. 2016; Ruffini Castiglione et al. 2016; Di Gregorio et al. 2015, 2014). To date, the best characterized are species of bacteria in the genus Pseudomonas, which demonstrate great abilities to use a variety of hydrocarbons, including BTEX, as sole carbon sources (Jahromi et al. 2014; Mukherjee et al. 2012). There are different metabolic pathways for BTEX degradation in bacteria. Monooxygenases and dioxygenases are involved in the key step of the first oxidative attack on the aromatic ring, described as the upper pathway for the oxidation of hydrocarbons (Jindrova et al. 2002). Monooxygenases oxidize the methyl or ethyl groups bound to aromatic rings. Dioxygenases oxidize the aromatic ring. Both the reactions lead to the formation of pyrocatechols, which are oxidized by catechol dioxygenases of the lower oxidation pathway, and the cleavage products from the oxidized aromatic ring are further metabolized through the tricarboxylic acid cycle (Farhadian et al. 2008; Andreoni and Gianfreda 2007).
It is reasonable to think that in complex matrices where the microbial community is well adapted to the presence of high concentrations of hydrocarbons, the communication among members of the community, or even between different communities, might play a role in the biodegradation process. Quorum sensing is a system that enables bacteria to communicate. In general, there are two types of quorum sensing signalling molecules, which are also referred as “autoinducers”: intraspecific signals, which are used by the same organism within a population, and interspecific signals used for communication among the entire microbial community (Mangwani et al. 2012). In Gram-positive and Gram-negative bacteria, there are two different types of autoinducers: Gram-positive bacteria utilize peptides, while Gram-negative bacteria utilize acyl-homoserine lactones (AHL) (Watson et al. 2002), varying in lengths and lateral side chains (Horinouchi 1999).
Quorum sensing autoinducers activate gene expression via signal transduction cascades. Some key bacterial behaviours regulated by quorum sensing include production of biosurfactants, formation of biofilms, horizontal gene transfer, and catabolic genes expression (Huang et al. 2003). These features are positively involved in hydrocarbon degradation. In fact, a strong correlation between the expression of genes involved in hydrocarbon depletion and quorum sensing has been proposed (Huang et al. 2003). As an example, a positive correlation between quorum sensing and the expression of the 2,3-cathechol dioxygenase of the meta-cleavage (lower) pathway for hydrocarbons degradation has been reported (Yong and Zhong 2013). On the other hand, there is a lack of evidence for quorum sensing control of dioxygenase and monooxygenase genes that encode enzymes involved in the upper pathway for BTEX oxidation.
In this work, a new strain of the Pseudomonas sp. was isolated from a decommissioned refinery’s soil in the north of Italy. The new strain showed the capacity to utilize several PAHs as sole carbon sources and the capacity to efficiently deplete the BTEX mixture, with a good percentage of depletion for all the composing aromatic moieties. The draft genome of the new strain was sequenced and annotated for genes putatively involved in the BTEX degradation and quorum sensing system. The strain was transformed with a plasmid expressing a lactonase, which cleaves the quorum sensing signal molecule, the acyl-homoserine lactone, to obtain a quorum sensing minus strain. The capacity of the wild type and the transformed strain to oxidize BTEX was studied by GC analysis. A quantitative RT-real-time PCR analysis was performed to study the role of quorum sensing in the regulation of the strain capacity to oxidize BTEX.
Materials and methods
Culture media
Basal salt medium (BSM) was used during the screening and the isolation of bacteria from hydrocarbon contaminated soil. On a per L basis, BSM contained 2.2 g Na2HPO4, 0.8 g KH2PO4, 3.0 g NH4NO3, 0.1 mM CaCl2, 2.0 mM MgCl2, 2.5 g NaCl, and 2.0 mL of Trace element solution, containing (on a per L basis) 0.03 g H3BO3, 0.2 g CoCl2.6 H2O, 0.1 g ZnSO4.7H2O, 30 mg MgCl2.4H2O, 30 mg NaMoO4.2H2O, 20 mg NiCl2.6H2O, and 10 mg CuSO4.5H2O. BSM-agar plates were made by autoclaving 1 L BSM containing 15 g/L Noble agar and pouring sufficiently cooled BSM-agar into sterile petri plates. Ramsay minimal medium (RMM) was used to grow P. putida AQ8 and its mutant in the presence of BTEX. On a per L basis, RMM contained 1.5 g KH2PO4, 6.7 g Na2H2PO4, 1.0 g NH4NO3, 0.2 g MgSO4.7H2O, 0.06 g FeNH4 citrate, 0.01 g CaCl2, and 1 mL of trace element solution. Luria Bertani (LB) medium used for growing the P. putida AQ8 and its mutant in the presence of readily available carbon sources contained 10 g of Tryptone, 5 g of yeast extract, 10 g of NaCl.
Isolation of bacteria
Bacteria were isolated from a petroleum hydrocarbon contaminated soil collected at the site of a decommissioned petroleum refinery in Trieste (Italy, 45° 36' 16.9" N; 13° 47' 56.4" E). Bacteria were isolated as follows: 2 g of soil were added to 200 mL of basal salt medium containing 2 mL of bitumen in toluene, 2 mL of diesel oil, 2 mL of pyrene, solubilized in diethyl ether at 500 ppm final concentration, and 2 mL of anthracene, solubilized in diethyl ether at 200 ppm final concentration. The cultures were maintained at room temperature on a rotary shaker at 150 rpm for 15 days in the dark. At the end of this incubation period, 20 mL was withdrawn and inoculated in a new flask with fresh culture medium in a total volume of 200 mL for a further incubation of 15 days in the dark. The passage was repeated two times. At the end of the third passage, serial dilutions of the liquid culture were prepared with sterile 0.9% NaCl, and 100 μl of each dilution was spread on LB-agar. The plates were inverted and incubated at 25 °C in the dark. Single colonies of bacteria were observed after 5–7 days of incubation. Single colonies were picked and transferred using a sterile loop to individual LB-agar plates. A total of 18 morphotypes were recovered.
Strain isolation
The different morphotypes were studied for their capabilities to grow on BTEX as sole carbon source. Bacterial strains were grown in BSM containing 2% BTEX v/v, in sealed tubes, and incubated on a rotary shaker at 150 rpm, in the dark, at 28 °C. The optical density at 600 nm (OD600) was measured daily. A single morphotype showed the capability to utilize BTEX as sole carbon source and was further analysed.
Strain identification
Bacterial DNA was extracted from the above-mentioned morphotype, using Gene Elute kit for bacterial cells (Sigma Aldrich) following the manufacturer’s protocol. The16S ribosomal rRNA genes were amplified by polymerase chain reaction (PCR) using 27 F (5'-AGAGTTTGATCCTGGCTCAG-3') and 1492R (5'-GGTTA CCTTGTTACGACTT-3') primers. The amplicons were purified using the Gel Elute kit (Sigma Aldrich) and then sequenced by GATC service (Ebersberg, Germany) and aligned to the sequence databases using BLASTN (Altschul et al. 1997). The morphotype was designated as Pseudomonas putida AQ8.
Genome sequencing and annotation
Pseudomonas putida AQ8 genomic DNA was extracted by Gen Elute™ Bacterial Genomic DNA (Sigma Aldrich), following the manufacturer’s instructions. DNA quantification was performed using Qubit® 3.0 (Invitrogen, Thermo Fisher Life Technologies) following manufacturer’s instructions. Pseudomonas putida AQ8 genome was sequenced by Admera Health (South Plainfield, New Jersey). MiSeq Illumina technology was applied to obtain a draft genome using 150-bp paired-end library. To test the quality of reads, Trimmomatic 0.36 and FastQC v0.11.5 tool were used. Abyss 2.0.1 and SPAdes 3.10.1 were used to assemble the genome using the shred reads approach. GapFiller v1.10 was used to close the gaps in the genome. The annotation was carried out using firstly Prodigal 2.50 and was used to find coding region (CDS) and translates them in to amino acidic sequences. The sequences then were aligned against different databases, such as UniProt, TrEMBL, Pfam, and Tigrfam, using Blast 2.5.0 and HMMer. Blast2go 4.1 was used to align our sequences against InterPro and Gene Ontology. BlastKOALA was used to annotate proteins in KEGG Orthology (KO) term. tRNAscaSE 1.3.1 was used to find tRNA. PROKKA 1.12 and IMG were used as automated pipelines to annotate the genome. The genome was submitted in IMG platform as Ga0299919 as GOLD Analysis project ID. BioLegato platform (Fristensky 2007) was used to annotate the pathways of BTEX degradation.
Quorum sensing binding site sequences were found in the genome using IMG platform (https://img.jgi.doe.gov). Genes involved in the oxidation of monoaromatic hydrocarbons were identified in the P. putida AQ8 genome using BioLegato (Fristensky 2007) and analysed for the presence of Lux box consensus sequences, 5’- NNCT(N12) AGNN -3’ (27), using KEGG pathways in the IMG platform, where the genome has been submitted (Ga0299919, IMG GOLD Analysis Project ID). The genes of interest were found, and the upstream sequences were retrieved and searched manually for lux box consensus sequences.
Transformation of Pseudomonas putida AQ8
The isolate Pseudomonas putida AQ8 was transformed with the plasmid pME6863 (Reimmann et al. 2002) containing aiiA gene from Bacillus sp. 240B1 encoding for a lactonase, which exerts an AHL-degrading activity. The plasmid contains tetracycline resistance Tcr gene. The transformation was performed by tri-parental mating (Smalla and Sobecky 2002), using Escherichia coli S17-1 containing the plasmid (pME6863), and the E. coli DH5α containing the pRK600 plasmid was used as helper (Finan et al. 1986) and P. putida AQ8 as the receiving strain. The three bacterial strains were grown separately in LB overnight. The cultures were then centrifuged, and the cell pellets were suspended in sterile saline phosphate buffered. P. putida AQ8, E. coli containing pME6863, and E. coli DH5α containing pRK600 were mixed in a ratio of 50:5:5. LB-agar plates with a sterile ring were used for the transformation experiment. Liquid cultures of each bacterial strain and the mixture of the three strains placed in the centre of the ring and incubated overnight at 28 °C.
Previously, a minimal inhibitory concentration (MIC) test was performed for each strain, using tetracycline and chloramphenicol at concentrations of 5–50 mg/L and 100–200 mg/L. Respectively E. coli strain (pME6863) was resistant to tetracycline at 30 mg/L but not resistant to chloramphenichol. E. coli helper DH5α (pRK600) was resistant to chloramphenichol at 25 mg/L but not to tetracycline. P. putida AQ8 was resistant to chloramphenichol at 100 mg/L and tolerant to tetracycline up to 5 mg/L. The optimum concentrations for the strain P. putida AQ8 receiving the pME6863 plasmid (P. putida AQ8 AHL−) were determined to be 30 mg/L for tetracycline and 100 mg/L for chloramphenicol. LB-agar plates containing 30 mg /L of tetracycline and 100 mg/L of chloramphenicol were used to select the mutant P. putida AQ8 AHL−.
Quorum sensing assay
The Quorum Sensing plate assay (Kawaguchi et al. 2008) was carried out using LB-agar plates containing 100 μL of an X-gal solution (8 mg/mL in DMSO), Pseudomonas aeruginosa PAO1 (Selin et al. 2012) as a positive control, and Agrobacterium tumefaciens ATCC2240 as the biosensor strain.
Growth of bacteria in LB and in RMM in the presence of BTEX as sole carbon sources
P. putida AQ8 and P. putida AQ8 AHL− were grown in LB and in RMM with BTEX as sole carbon source for 48 h in triplicate for each time of analyses. Each component of BTEX was present at a concentration of 50 mg/L. The medium was inoculated with a bacterial inoculum deriving from the overnight growth in LB of a single colony picked from an agarised plate and performed at an optical density at 600 nm (SPECTROstar Nano, BMG Labtech) of 0.01 units of absorbance and incubated at 28 ± 1 °C on an orbital shaker at 150 rpm overnight. LB growth curves were performed in 250-mL flasks containing 50 mL of LB. The BTEX mixture was added to 125-mL bottles containing 25 mL of RMM. The bottles were closed by rubber septa sealed with metallic caps. Plate counts were used to measure the growth of the isolate in the different growth conditions. Samples from 3 replicates bottles were withdrawn every 12 h. Serial dilutions of the bacterial cells were made in sterile saline solution (NaCl 0.9% w/v).
BTEX depletion
A BTEX depletion experiment was carried out using 25 mL of bacterial culture in 125-mL sealed bottles containing BTEX in RRM. The analyses were performed for 48 h in triplicate for each time of analysis. A control set of sealed bottles were prepared in the absence of the bacterial inoculum to measure the abiotic BTEX depletion. The quantification of BTEX was performed scarifying the entire volumes of the replicates. The initial inoculum was prepared as previously described and washed in sterile saline solution and pelleted by centrifugation. The pellets were suspended in RMM, and the OD600 of each was measured. A dilution in RMM with an OD600 of 0.01 was used to inoculate the 125-mL bottles for the experiment. Samples for plate counts and for the measurement of BTEX were taken each 12 h. At each sampling time, a small aliquot (1 mL) was withdrawn for plate counts, and then the leftover was used to measure the concentration of BTEX with liquid extraction as described in the following section.
BTEX measurement
A total volume of 5 mL of pentane and styrene (at a concentration of 0.5 mg/mL in pentane) was added to each sample bottle (25 mL), and then they were stored overnight at 4 °C. The entire volume was then centrifuged at 4500 x g for 10 min, and aliquots of 500 μL were taken. In GC vials, 500 uL of sample and 1 mL of pentane were added. Samples (1 μL) were injected into an Agilent 7890A GC coupled with a split–splitless injector and a flame ionization detector with a split ratio at 1:10. Separation of target chemicals was achieved using a DB-23 capillary column (30 m × 320 m × 0.25 m; Agilent, California, USA). Helium was used as carrier gas at a flow rate of 1.78 mL/min. The oven ramping program was set as follows: initial temperature at 36 °C for 3 min, then from 36 to 106 °C with incremental increases of 5 °C/min, and a final hold at 186 °C, with incremental increases of 20 °C/min for 1 min.; the detector’s heater was 250 °C, 35 mL/min for H2 flow, 400 mL/min for air flow, and 25 mL/min for He. Before the measurement, a calibration curve was performed both for each compounds separately and in the mixture. The analysis was performed in 5 mL of pentane. Each point and each compound/mixture was treated the same as the actual samples. Depletion percentages were calculated with reference to T0 as follows: [B/T/E/X (T0)] – [B/T/E/X (Ti)]/[B/T/E/X (T0)] × 100, subtracting the BTEX abiotic depletion.
RNA extraction
RNA extraction was performed using the RNeasy Mini Kit (Qiagen). Some modifications were made to the manufacturer’s protocol. The extraction of RNA in samples containing BTEX showed very low yields of RNA. The samples were treated with RNAprotect® Bacteria Reagent (Qiagen) (2:1 RNAprotect® Bacteria Reagent: cell culture) and centrifuged for 10 min at 5000 x g. In order to achieve higher yields, buffer TEN (100 mM Tris-HCl pH 8.0, 50 mM EDTA, 500 mM NaCl) (Canul-Chan et al. 2017) was used to wash the cells. Cells were resuspended and then pelleted three times at 5000 x g for 10 min. Cells were lysed using lysozyme (100 μL, 15 mg/mL in 300 mM Tris-EDTA pH 8) and Proteinase K (20 μl, 20 mg/mL) shaking at RT for 10 min. The manufacturer’s protocol was then followed. DNAse treatment and cDNA synthesis were performed using Maxima First strand cDNA Synthesis Kit (Thermo Fischer Scientific) following the manufacturer’s protocols.
Quantitative real-time PCR
Quantitative real-time PCR was performed using SsoFast EvaGreen Supermix (BioRad) (Wei et al. 2011). Primers for the benzene/toluene dioxygenase gene (benzA), the biphenyl dioxygenase (bphA), and rpoB gene encoding the DNA-dependent RNA-polymerase beta subunit were designed on the single copy of the genes annotated in the genome of P. putida AQ8. Benzene/toluene dioxygenase gene (benzA) encodes the enzyme that carries out the initial ring cleavage oxidation reaction of monoaromatic hydrocarbons; biphenyl dioxygenase gene (bphA) is involved in the upper pathway of monoaromatic hydrocarbon oxidation. The primers were designed using the Pick primers function of BlastN (Altschul et al. 1997). The rpoB gene encoding for the RNA polymerase beta subunit was adopted as housekeeping gene. The list and sequences for primers are shown in Table 1.
The expression levels described were calculated on the basis of fold expression with 2^-ΔΔCt formula (Livak and Schmittgen 2001) comparing P. putida AQ8AHL with P. putida AQ8.
Statistical analysis of data
Relative normalized gene expression, plate counts, and BTEX depletion data were analysed by RM 2 ways ANOVA using Ŝidák as post hoc test to compare two sets of means (AHL− vs AQ8). The statistical analysis concerning relative normalized gene expression was performed on the ΔCt of the two strains for both bphA and benzA genes. All the statistical analyses were performed in GraphPad Prism (version 8.2.1).
Results
Isolation, selective screening, and characterization
Bacterial strains, isolated from a sample of soil collected in a decommissioned oil refinery, were initially selected for their capabilities to grow on bitumen and diesel oil as sole carbon sources. Isolated strains were then cultured with BTEX as the sole carbon source, and the unique strain that demonstrated the best growth was selected to be used in further experiments. The isolate was identified as a strain of Pseudomonas putida by sequencing of gene coding for the 16S rRNA and alignment in BlastN obtaining a 99% of sequence identity with Pseudomonas putida (Accession No. CP022561). It was designated Pseudomonas putida AQ8.
Transformation of P. putida AQ8 and quorum sensing assays
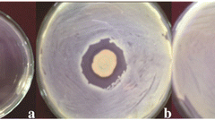
The P. putida AQ8 AHL− (quorum sensing minus), harbouring the pME6863 plasmid for the constitutive expression of the lactonase, was obtained by triparental mating (Fig. 1). A plate assay utilizing Agrobacterium tumefaciens ATCC 2240 as reporter strain was adopted to verify the capability of the P. putida AQ8 to produce the long-chain AHL and of the AQ8 AHL− strain to express the lactonase for the long-chain AHL degradation. In fact, P. putida AQ8 was found to be positive for the production of long-chain AHL (Fig. 1, panel b), resulting in inducing the expression of the β-galactosidase in the reporter strain, which metabolize the plate added X-Gal reagent, producing a blue-coloured indigo derivative, indicating a positive bioassay. The lack of blue colour of the reporter strain in the presence of the P. putida AQ8 AHL− (Fig. 1, panel c) indicated the autoinducer; the acyl-homoserine lactone (AHL) was degraded in P. putida AQ8 AHL− by the constitutive expression of the lactonase harboured by the pME6863 plasmid.
Quorum sensing assays with a the positive control strain for AHL production, Pseudomonas aeruginosa PA01 (1) and the bioreporter stain, Agrobacterium tumefaciens ATCC 2240 (2); b the test strain, Pseudomonas putida AQ8 (3) and bioreporter stain, Agrobacterium tumefaciens ATCC 2240 (2); and c the test strain, Pseudomonas putida AQ8 AHL− (4) and the bioreporter stain, Agrobacterium tumefaciens ATCC 2240 (2)
Growth of P. putida AQ8 and P. putida AQ8 AHL− on BTEX
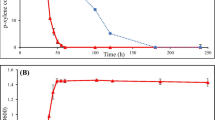
In order to compare the growth kinetics of the P. putida AQ8 and its transformed AHL− strain, both in the absence and presence of a BTEX, growth curves in rich medium (LB) and minimal medium (RMM) with BTEX mixture as sole carbon sources were performed (Fig. 2). In rich medium, the two strains showed very similar kinetics of growth, reaching the stationary phase at the same time points. In minimal medium with BTEX as sole carbon sources, the two strains reached the same cell density, but P. putida AQ8 grew more slowly than P. putida AQ8 AHL−. P. putida AQ8 AHL− reached the highest cell density after 12 h post-inoculation, decreasing in cell density in the following hours. On the other hand, P. putida AQ8 reached the highest cell density after 24 h post-inoculation. Then, even though less dramatic than for AHL−, a progressive decrease in AQ8 cell density was observed.
Growth of Pseudomonas putida AQ8 and Pseudomonas putida AQ8 AHL− in LB (panel a) and RMM with BTEX as sole carbon sources (panel b) determined by optical density and plate count assays, respectively. The error bars represent standard deviations from the means of three biological replicates. Values referred to AQ8 and AHL− at different sampling times in LB (panel a) showed a P ≥ 0.05. Values referred to AQ8 and AHL− at different sampling times in RMM (panel b) showed a P ≤ 0.05
BTEX depletion
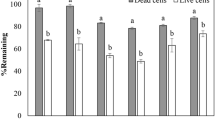
The components of BTEX have very high vapour pressures and evaporate rapidly. Uninoculated RRM BTEX medium was incubated for 48 h at 30 °C with shaking on a rotary shaker (150 rpm) to determine the extent of evaporation under the experimental conditions. The percent (%) degradation of each BTEX components in cultures inoculated with P. putida AQ8 and P. putida AQ8 AHL− was calculated taking into account the depletion of each BTEX components due to evaporation. The depletion of BTEX due to evaporation was subtracted from the total depletion in order to obtain the actual depletion due to the bacterial degradation process. Comparison of the percentages of depletion of each BTEX components in the absence and in presence of the bacterial inocula, the AQ8 and the AQ8 AHL− strains (Fig. 3a–e), revealed that although there was a considerable amount of evaporation in the control tubes, there was greater depletion by P. putida AQ8, over and above that resulting from evaporation, starting from 12 h of incubation and lasting up to 36 h. Depletion of BTEX components by P. putida AQ8 AHL−, over and above evaporation in the control tubes was observable only after 12 h of incubation with lower percentages compared with the ones observed for AQ8 at 24 and 36 h p.i.
Depletion of BTEX components over 48 h of incubation in P. putida AQ8 and P. putida AQ8 AHL−: panel a, benzene; panel b, toluene; panel c, ethylbenzene; panel d, para/meta-xylene; panel e, ortho-xylene; and panel f, abiotic BTEX (control conditions). The error bars represent the standard deviations from the means of three biological replicates. Values at different sampling times showed a P ≤ 0.05
More in details, in tubes inoculated with P. putida AQ8, the % depletion of all BTEX components clearly increased between 12 and 24 h. p.i.: from 15 to 40% for benzene; from 4 to 35% for toluene; from 0 to 37% for ethyl-benzene; from 0 to almost 37% for meta/para-xylene; and from 0 to 34% for ortho-xylene. By 36 h. p.i., benzene and ortho-xylene were depleted by 45% and 40%, respectively, and toluene, ethyl-benzene, and para/meta-xylene were depleted 37–38%. In tubes inoculated with P. putida AQ8 AHL−, the percentages of BTEX depletion over and above depletion resulting from evaporation was approximately 10% for all BTEX components, after 12 h. p.i. In panel f, the depletion of the BTEX compounds is showed and it is related to T0.
Genome annotation
Statistics of AQ8 genome assembly and annotation are reported in Table 2. The draft genome of the isolate P. putida AQ8 consists of 53 contigs greater than 500 kbp, contigs which were less than 500 bp in length were discarded. The total length of contigs was 5,894,357 bp. The G + C percentage determined from the genome sequence is 62.02%. A total of 5425 coding DNA sequences (CDS) were predicted, of which 5294 were annotated as protein-coding sequences, while 78 were annotated as RNA genes. One operon of rRNA genes was retrieved, including seven copies of 5S and one copy of 16S and 23S. More than 73% of CDS were assigned to cluster of orthologous-group classification and functional categories (COG).
The annotation of pathways involved in BTEX degradation was performed specifically with BioLegato (Fristensky 2007). BioLegato aligned the CDS protein sequences of the genome to a small database made by the user based on selected sequences, e.g. for specific enzymes and organisms, retrieved from NCBI databases. The annotation was performed using also other tools; e.g., BlastN/P, COG, KEGG, IMG, and results were compared together in order to improve the genome annotation.
A detail of the results of the genomic annotation of the draft genome of P. putida AQ8, referred to the codifying for proteins and enzymes involved in hydrocarbon degradation and quorum sensing, are listed in Table 3. Quorum sensing was annotated using tools as IMG, Prokka, Prodigal, and Blast N/P. Las I/R system, containing both the genes for the synthase and the receptor of the autoinducer molecule, was found in a single copy. Three more copies of the gene for the receptor of the autoinducer, putative lux solos, were retrieved.
Two putative ring hydroxylating dioxygenases of the upper pathway of BTEX degradation were retrieved. One, putatively coding for the bphA dioxygenases, ascribable to the biphenyl/toluene/benzene class, was annotated as part of the ethylbenzene degradation pathway. A second dioxygenase, the benzA, was ascribable to the benzoate class and annotated as part of the benzoate degradation pathway, organized in the ben operon. BioLegato annotated the benzA as part of the complete degradation pathway of benzene and toluene. The benR gene, encoding the transcriptional regulator of the benzene/toluene degradation pathway, was retrieved at the 5′ end of the ben operon (Fig. 4).
The ben operon: benR is a XylS regulator, an AraC family regulator; benA encodes for benzoate dioxygenase α-subunit; benB encodes for benzoate dioxygenase β-subunit; benC encodes for oxydase NAD-binding site; and benD a cis-diol dehydrogenase, whose product is cathecol, which is cleaved by catechol dioxygenase (CatA), producing cic, cis-muconate. benK, benE, and benF are transporters: benK and benE are H+/aromatic acid symporters, while benF is outer membrane porine. The ben operon is located on Scaffold 5, from position 5_162-5_170
Ring-hydroxylating monooxygenases were not retrieved. On the other hand, the genes coding for the rest of the pathway for xylene isomers degradation were retrieved in single copies. Two copies of the catechol 1, 2-dioxygenase of the lower pathway of BTEX degradation by the ortho-cleavage of the aromatic rings were also annotated. DNA sequences upstream of genes that encode for both bphA and benzA dioxygenases were characterized by the presence of regulatory lux boxes. In particular, it was found the Lux box close to regulator protein at the ben operon (Fig. 5a) and directly upstream to bphA gene (Fig. 5b).
Identification of lux boxes in the P. putida AQ8 genome. a Upstream and partial coding sequence of the benR gene (in italics), which encodes the transcriptional regulator of the benzene/toluene degradation pathway. b Upstream and partial coding sequence of the bphA gene (in italics), which encodes for dioxygenase for benzoate/ethylbenzene degradation pathway. The Lux boxes are highlighted in bold letters and the coding sequence is in italic font, the translation start site of the coding sequence is underlined
Analysis of gene expression by real time PCR
The expression levels of the benzA and bphA dioxygenase during the growth of AQ8 and AQ8 AHL− strains were studied by RT-real-time qPCR. The levels of expression of the genes of interest were measured in P. putida AQ8 and P. putida AQ8 AHL− during the growth of the strains in presence of BTEX as sole carbon sources. Results are expressed as relative normalized expression of the two dioxygenases in AQ8 AHL− compared with AQ8. The expression levels described were calculated on the basis of fold expression with 2−ΔΔCt formula (Livak and Schmittgen 2001) comparing P. putida AQ8AHL− with P. putida AQ8 by calculating the expression with ΔΔCt method. Values below 0.5 indicate downregulation, greater than 2 indicate upregulation, and between 0.5 and 2 indicate no differences in expression levels. Results obtained are shown in Table 4. The table indicates that in P. putida AQ8 AHL− with reference to P. putida AQ8, at 12-h post-inoculum (p.i.), the expression of benzA and bphA was upregulated of 5.4- and 13.8-fold, respectively. As the incubation progresses, the two genes benzA and bphA were downregulated with the exception of the bphA at 36 h of incubation and the benzA at 48 h of incubation, when the levels of expression of the two genes were comparable in the two strains, despite the suffering metabolic state of the AQ8 AHL− with reference to AQ8. A comparison between the levels of expression of the two genes in the two strains when they reached the same cell density (12 h for AQ8 AHL− and 24 h for AQ8) showed that the levels of expression of the two genes in P. putida AQ8 AHL− were lower with reference to P. putida AQ8 (Table 4).
Discussion
A new P. putida strain, AQ8, was isolated from a polluted soil located in a decommissioned oil refinery in Italy. The strain was selected for its ability to grow on BTEX mixture as sole carbon sources. Several Pseudomonas species have been found to be both tolerant and capable to degrade BTEX, even though the AQ8 strain actually resulted to be more efficient in terms of concentration of abatement and kinetics of depletions of other candidates (Imperato et al. 2019, You et al. 2013). Moreover, P. putida AQ8 was able to degrade all the xylene isomers to the same extent and to the same extent of the other different monoaromatics in the BTEX mixture. Many studies showed that xylene isomers are associated to lower degradation percentages or are degraded more slowly than the other BTEX components (Mangwani et al. 2015; El-Naas et al. 2014; You et al. 2013). Xylenes are more resistant to oxidation because of methyl groups on the aromatic rings, which must be oxidized by the intervention of dedicated monooxygenases (El-Naas et al. 2014). Consortia of BTEX transforming bacteria are reported as more efficient in BTEX depletion when compared with specialized strains in axenic culture. An almost complete degradation of BTEX at higher concentration than here tested was obtained in a similar time interval, by a consortium of Pseudomonas sp., Yarronia sp., Acinetobacter sp., Corynebacterium sp., and Sphingomonas sp. (Jo et al. 2008). An efficient degradation of BTEX mixture concentrations identical to the here tested was obtained in only 8 h by a gasoline acclimatized consortium of P. putida and P. stutzeri (Chavez et al. 2017). The higher efficiency of consortia instead of axenic cultures might be associated to the exploitation of different pathways involved in BTEX degradation. In this context, AQ8 strain might be considered as a valid candidate for the development of consortium-based approaches for BTEX depletion.
The pathways reported in literature to be involved in BTEX depletion are two, distinguished on the base of the type of first oxidative attack on the aromatic rings, catalysed by dioxygenase or monooxygenase (Jindrova et al. 2002). The annotation of the draft genome of AQ8 indicated that both pathways are present, conferring to the strain the capability to deplete all the components of the BTEX mixture. In fact, the pathways for BTE transformation were annotated as complete pathways, encoded in single copy. On the other hand, only a truncated pathway for xylene isomer oxidation, which was missing genes for monooxygenases, was identified in the AQ8 genome. Nonetheless, AQ8 was able to degrade xylenes. This apparent contradiction may be explained by the fact that the draft genome sequence of P. putida AQ8 was not complete, and thus sequences encoding monooxygenases may have been absent from the sequence data set. On the other hand, the recalcitrance to biodegradation of the xylenes isomer is reported as generally attenuated when the mixture all the BTEX are depleted. In consumption experiments with monoaromatic compounds by P. putida BTEX 30, the meta-xylene isomer was not utilized when present as a sole carbon source. However, in experiments where the full BTEX mixture was used as carbon source, all the aromatic compounds, including the meta-xylene isomer, were depleted almost at the same level. This may be due to the capability of the strain to degrade the entire mixture of BTEX, using some components as intermediates for the degradation of the others (Mangwani et al. 2015). This mechanism may also be a metabolic feature of P. putida AQ8. The lack of the complete sequence of the genome might hide some functional features of the strain.
In relation to quorum sensing, the lack of the complete AQ8 genome sequence leads to a number of possible interpretations of the data obtained. Although the Las I/R system encoded in the P. putida AQ8 genome contains both the gene coding for the synthase and the for the receptor of the autoinducer molecule, was found in a single copy, more than one copy for the gene encoding for the autoinducer receptor, putative LuxR-solos, were retrieved. These multiple copies might be consistent with the presence of LuxR-solos receptors, which permit to bacterial strains to exploit autoinducers produced outside of the cell and to receive chemical signals from other species present in the local environment (Fuqua et al. 2006). However, the lack of the complete sequence of the AQ8 genome cannot exclude the presence of other copies of genes for the synthesis of the acyl-homoserine lactone autoinducer, which might be sensed by the putative LuxR-solos receptor. In this context, the strategy to eliminate quorum sensing in AQ8 to study its possible involvement in BTEX degradation was the approached by the insertion of a plasmid expressing lactonase, which inactivated autoinducer molecules after they were produced. The resulting transformation of the strain was associated with the loss of its capacity to reach the same levels of BTEX depletion, suggesting an involvement of quorum sensing in the regulation of the dedicated metabolism.
The annotation of the AQ8 genome identified two genes encoding for the benzA and bphA dioxygenases, clustered in pathways annotated for the BTEX oxidation, and preceded by regulatory sequences responding to the quorum sensing system. In order to understand if quorum sensing has a role in BTEX depletion, the comparison of gene expression levels was performed at an equal cell density of AQ8 and the AQ8 AHL− cultures. In fact at 12 h for AHL− and 24 h for AQ8), the two strains showed equal cell density, and the transcription levels of both the benzA and bphA dioxygenase genes were lower in the absence of an intact quorum sensing system. At the maximum cell density of the P. putida AQ8 AHL− culture, 12 h p.i., the percentage of BTEX depletion was lower than the percentage depleted by the P. putida AQ8 culture at 24 h p.i.. The AQ8 AHL− culture reached a steady state of growth in a shorter time than the AQ8 culture. At the same time, the decreased ability of the AQ8 AHL− culture to deplete BTEX was correlated with a faster decrease in cell viability, when compared with the AQ8 culture. However, although the viability of AQ8 AHL− cells decreased, AQ8 AHL− cells continued to express genes for BTEX oxidation. The normalized level of bphA transcription at 36 h of incubation and benzA transcription at 48 h were both at the same level of expression of the healthier AQ8 cells at the same time of incubation, even though AQ8 AHL− was at a significant lower cell density when compared with AQ8. These data confirmed the key role of the dioxygenases of the upper BTEX oxidative pathway, encoded by the bphA and benzA genes, in the ability of P. putida AQ8 to eventually cope with the toxicity of monoaromatic hydrocarbons. In this context, it is reasonable to assess that, even though the lack of the complete sequence of the genome might hide some regulatory features of the strain, results obtained are actually the first evidences of the involvement of the quorum system in the modulation of the level of expression of the dioxygenases of the upper pathway for the BTEX depletion.
Conclusions
The ability of P. putida AQ8 to degrade BTEX as a single strain and in a relatively short interval time suggests that the strain could be exploited, in axenic culture or in a consortium, as a candidate for bio-based treatment for liquid matrices and/or in biofilters contaminated with BTEX. This study suggests, for the first time, that the quorum sensing system might be involved in the modulation of the dioxygenases of the upper BTEX oxidative pathways.
Abbreviations
- rRNA:
-
Ribosomal DNA
- X-gal:
-
5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside
- DMSO:
-
Dimethyl sulfoxide
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman, DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids Res 25(17):3389–3402
Andreoni V, Gianfreda L (2007) Bioremediation and monitoring of aromatic-polluted habitats. Appl Microbiol Biotechnol 76:287–308
Becarelli S, Chicca I, Siracusa G, La China S, Gentini A, Lorenzi R, Munz G, Petroni G, Levin DB, Di Gregorio S (2019) Hydrocarbonoclastic Ascomycetes to enhance co-composting of total petroleum hydrocarbon (TPH) contaminated dredged sediments and lignocellulosic matrices. New Biotechnol 50:27–36
Bowlen GF, Kosson DS (1995) In situ processes for bioremediation of BTEX and petroleum fuel products. In: Young LY, Cerniglia CE (eds) Microbial transformations and degradation of toxic organic chemicals. Wiley-Liss, Inc., New York, pp 515–542
Canul-Chan M, Chable-Naal J, Rojas-Herrera R, Zepeda A (2017) Hydrocarbon degradation capacity and population dynamics of a microbial consortium obtained using a sequencing batch reactor in the presence of molasses. Biotechnol Bioprocess Eng 22(2):170–177
Chavez JAM, Martinez JAA, Haskins WE, Askar KA, Saldana HAB (2017) Gene expression during BTEX biodegradation by a microbial consortium acclimatized to unleaded gasoline and a Pseudomonas putida strain (HM346961) isolated from it. Pol J Microbiol 66:189–199
Di Gregorio S, Gentini A, Siracusa G, Becarelli S, Azaizeh H, Lorenzi R (2014) Phytomediated biostimulation of the autochthonous acceleration of the depletionof polycyclic aromatic hydrcarbons in contaminated sediments. BioMed Res Int:891630
Di Gregorio S, Giorgetti L, Ruffini Castiglione M, Mariotti L, Lorenzi R (2015) Phytoremediation for improving the quality of effluents from a conventional tannery wastewater treatment plant. Int J Environ Sci Technol 12:1387–1400
Di Gregorio S, Siracusa G, Becarelli S, Mariotti L, Gentini A, Lorenzi R (2016) Isolation and characterization of a hydrocarbonoclastic bacterial enrichment from total petroleum hydrocarbon contaminated sediments: potential candidates for bioaugmentation in bio-based processes. Environ Sci Pollut Res Int 23(11):10587–10594
Durmusoglu E, Taspinar F, Karademir A (2010) Health risk assessment of BTEX emissions in the landfill environment. J Hazard Mater 176:870–877
El-Naas MH, Acio JA, El Telib AE (2014) Aerobic biodegradation of BTEX: Progresses and Prospects. J Environ Chem Eng 2(2):1104–1122
European Environment Agency: www.eea.europa.eu (2012)
Farhadian M, Vachelard C, Duchez D, Larroche C (2008) In situ bioremediation of monoaromatic pollutants in groundwater: a review. Bioresour Technol 99:5296–5308
Finan TM, Kunkel B, De Vos GF, Signer ER (1986) Second symbiotic megaplasmid in Rhizobium meliloti carrying exopolysaccharide and thiamine synthesis genes. J Bacteriol 167:66–72
Fristensky B (2007) BIRCH: a user-oriented, locally-customizable, bioinformatics system. BMC Bioinformatics 8(1):54
Fuqua C (2006) The QscR quorum-sensing regulon of Pseudomonas aeruginosa: an orphan claims its identity. J Bacteriol Res 188(9):3169–3171
Horemans B, Worobiec A, Buczynska A, Van Meel K, Van Grieken R (2008) Airborne particulate matter and BTEX in office environments[J]. J Environ Monit 10:867–876
Horinouchi S (1999) Butyrolactones that control secondary metabolism and cell differentiation in Streptomyces. In: Dunny GM, Winans SC (eds) Cell-cell signaling in Bacteria. ASM, Washington, pp 193–207
Huang Y, Zeng Y, Yu Z, Zhang J, Feng H, Lin X (2003) In silico and experimental methods revealed highly diverse bacteria with quorum sensing and aromatics biodegradation systems – a potential broad application on bioremediation. Bioresour Technol 148:311–316
Imperato V, Portillo-Estrada M, McAmmond BM, Douwen Y, Van Hamme JD, Gawronski SW, Thijs S (2019) Genomic diversity of two hydrocarbon-degrading and plant growth-promoting pseudomonas species isolated from the oil field of Bóbrka (Poland). Genes 10(6):443
Irwin REA (1997) Fuel oil number 2 – heating oil entry. In: Irwin RJ (ed) Environmental contaminants encyclopedia. Fort Collins, National Park Service, pp 805–825
Jahromi H, Fazaelipoor MH, Ayatollahi Sh, Niazi A (2014) Asphaltenes biodegradation under shaking and static conditions. Fuel 117:230–235
Jindrova E, Chocova M, Demnerova K, Brenner V (2002) Bacterial aerobic degradation of benzene, toluene, ethylbenzene and xylene. Folia Microbiol 47:83–93
Jo MS, Rene ER, Kim SH, Park HS (2008) An analysis of synergistic and antagonistic behavior during BTEX removal in batch system using response surface methodology. J Hazard Mater 152(3):1276–1284
Kawaguchi T, Chen YP, Norman RS, Decho AW (2008) Rapid screening of quorum-sensing signal N-acyl homoserine lactones by an in vitro cell-free assay. Appl Environ Microbiol 74(12):3667–3671
Liu Y, Shao M, Fu L, Lu S, Zeng L, Tang D (2008a) Source profiles of volatile organic compounds (VOCs) measured in China: part I[J]. Atmos Environ 42:6247–6260
Liu Y, Shao M, Lu S, Lu S, Zeng L, Tang D (2008b) Source apportionment of ambient volatile organic compounds in the Pearl River Delta, China: part II[J]. Atmos Environ 42:6261–6274
Liu Y, Shao M, Lu S, Lu S, Zeng L, Tang D (2008c) Volatile organic compound (VOC) measurements in the Pearl River Delta (PRD) region, China[J]. Atmos Chem Phys 8:1531–1545
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25(4):402–408
Mackay D, Callcott D (1998) Partitioning and physical chemical properties of PAHs. In: Neilson AH (ed) The handbook of environmental chemistry. Springer-Verlag, Berlin, pp 325–346
Mangwani N, Dash HR, Chauhan A, Das S (2012) Bacterial quorum sensing: functional features and potential applications in biotechnology. J Mol Microb Biotech 22(4):215–227
Mangwani N, Kumari S, Das S (2015) Involvement of quorum sensing genes in biofilm development and degradation of polycyclic aromatic hydrocarbons by a marine bacterium Pseudomonas aeruginosa N6P6. Appl Microbiol Biotechnol 99(23):10283–10297
Marston CP, Pereira C, Ferguson J, Fischer K, Hedstrom O, Dashwood WM (2001) Effect of a complex environmental mixture from coal tar containing polycyclic aromatic hydrocarbons (PAH) on the tumor initiation, PAH-DNA binding and metabolic activation of carcinogenic PAH in mouse epidermis. Carcinogenesis 22:1077–1086
Mohammad BT, Veiga MC, Kennes C (2007) Mesophilic and thermophilic biotreatment of BTEX-polluted air in reactors. Biotechnol Bioeng 97:1423–1438
Morehead NR, Eadie BJ, Lake B, Landrum PD, Berner D (1986) The sorption of PAH onto dissolved organic matter in Lake Michigan waters. Chemosphere 15:403–412
Mukherjee AK, Bordoloi NK (2012) Biodegradation of benzene, toluene, and xylene (BTX) in liquid culture and in soil by Bacillus subtilis and Pseudomonas aeruginosa strains and a formulated bacterial consortium. Environ Sci Pollut Res 19:3380–3388
Reimmann C, Ginet N, Michel L, Keel C, Michaux P, Krishnapillai V, Zala M, Heurlier K, Triandafillu K, Harms H, Défago G, Haas D (2002) Genetically programmed autoinducer destruction reduces virulence gene expression and swarming motility in Pseudomonas aeruginosa PAO1 The GenBank accession number for the aiiA nucleotide sequence is AF397400. The GenBank accession numbers for the nucleotide sequences of the 16S rRNA genes of strains A23 and A24 are AF397398 and AF397399. Microbiology 148(4):923–932
Ruffini Castiglione M, Giorgetti L, Becarelli S, Siracusa G, Lorenzi R (2016) Polycyclic aromatic hydrocarbon-contaminated soils: bioaugmentation of autochthonous bacteria and toxicological assessment of the bioremediation process by means of Vicia faba L. Environ Sci Pollut Res 23:7930–7941
Selin C, Fernando WGD, de Kievit T (2012) The PhzI/PhzR quorum-sensing system is required for pyrrolnitrin and phenazine production, and exhibits cross-regulation with RpoS in Pseudomonas chlororaphis PA23. Microbiology 158:896–907
Smalla K, Sobecky PA (2002) The prevalence and diversity of mobile genetic elements in bacterial communities of different environmental habitats: insights gained from different methodological approaches. FEMS Microbiol Ecol 42:165–175
Watson WT, Minogue TD, Val DL, Von Bodman SB, Churchill MEA (2002) Structural basis and specificity of acyl-homoserine lactone signal production in bacterial quorum sensing. Mol Cell 9:685–694
Wei ZH, Bai L, Deng Z, Zhong JJ (2011) Enhanced production of validamycin a by H2O2-induced reactive oxygen species in fermentation of Streptomyces hygroscopicus 5008. Bioresour Technol 102(2):1783–1787
Xue W, Warshawsky D (2005) Metabolic activation of polycyclic and heterocyclic aromatic hydrocarbons and DNA damage:a review. Toxicol Appl Pharmacol 206:73–93
Yong YC, Zhong JJ (2013) Regulation of aromatics biodegradation by rhl quorum sensing system through induction of catechol meta-cleavage pathway. Bioresour Technol 136:761–765
You Y, Shim J, Cho CH, Ryu MH, Shea P, Kamala-Kannan S, Chae JC, Oh BT (2013) Biodegradation of BTEX mixture by Pseudomonas putida YNS1 isolated from oil-contaminated soil. J Basic Microbiol 53:469–475
Funding
This manuscript presents research results derived from the Bioresnova project 135/11, co-financed by Fondazione Pisa and the Department of Biology, University of Pisa. The research was supported, in part, by a Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery grant (RGPIN-04945-2017) held by DBL. SB was supported by the Department of Biology and BD Biodigressioni srl Pisa, Italy. IC was a visiting PhD student in the Department of Biosystems Engineering at the University of Manitoba, Canada, supported by Discovery grant (RGPIN-04945-2017) held by DBL.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Gerald Thouand
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chicca, I., Becarelli, S., Dartiahl, C. et al. Degradation of BTEX mixture by a new Pseudomonas putida strain: role of the quorum sensing in the modulation of the upper BTEX oxidative pathway. Environ Sci Pollut Res 27, 36203–36214 (2020). https://doi.org/10.1007/s11356-020-09650-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-020-09650-y