Abstract
Secondary growth of stems is an important process for the radial increase of trees. To gain an insight into the molecular mechanisms underlying stem development from primary to secondary growth and to provide information for molecular research and breeding in Betula platyphylla (birch), the gene expression profiles of material from the first, third, and fifth internodes (IN) of 3-month-old seedlings were analyzed. Compared with the first IN, 177 genes were up-regulated and 157 genes down-regulated in the third IN; in the fifth IN, 180 genes were up-regulated and 275 genes were down-regulated. The expressions of 24 genes were up-regulated and 6 genes were down-regulated in the fifth IN relative to the third IN. The differentially expressed genes were annotated as having roles in cambium, xylem, and phloem development and formation; including cell wall expansion, cellulose biosynthesis, lignin biosynthesis and deposition, xylem extension, cell wall modification, and growth hormone responses. The expressions of genes related to cell wall expansion and cellulose biosynthesis in the primary cell wall were down-regulated in the third and fifth IN relative to the first IN. Genes involved in lignin biosynthesis, xylem extension, and cellulose synthesis in the secondary cell wall were up-regulated in the third and fifth IN relative to the first IN. These results described the patterns of gene expression during stem development in birch and provided candidate genes for further functional characterization.




Similar content being viewed by others
References
Adams DJ (2004) Fungal cell wall chitinases and glucanases. Microbiology 150:2029–2035
Almeida T, Menendez E, Capote T, Ribeiro T, Santos C, Goncalves S (2013) Molecular characterization of Quercus suber MYB1, a transcription factor up-regulated in cork tissues. J Plant Physiol 170:172–178
Andersson-Gunneras S, Mellerowicz EJ, Love J, Segerman B, Ohmiya Y, Coutinho PM, Nilsson P, Henrissat B, Moritz T, Sundberg B (2006) Biosynthesis of cellulose-enriched tension wood in Populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. Plant J: Cell Mol Biol 45:144–165
Audic S, Claverie JM (1997) The significance of digital gene expression profiles. Genome Res 7:986–995
Bai TT, Xie WB, Zhou PP, Wu ZL, Xiao WC, Zhou L, Sun J, Ruan XL, Li HP (2013) Transcriptome and expression profile analysis of highly resistant and susceptible banana roots challenged with Fusarium oxysporum f. sp. cubense tropical race 4. PLoS One 8:e73945
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546
Brown DM, Zeef LA, Ellis J, Goodacre R, Turner SR (2005) Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 17:2281–2295
Bryan AC, Obaidi A, Wierzba M, Tax FE (2012) XYLEM INTERMIXED WITH PHLOEM1, a leucine-rich repeat receptor-like kinase required for stem growth and vascular development in Arabidopsis thaliana. Planta 235:111–122
Burn JE, Hocart CH, Birch RJ, Cork AC, Williamson RE (2002) Functional analysis of the cellulose synthase genes CesA1, CesA2, and CesA3 in Arabidopsis. Plant Physiol 129:797–807
Cano-Delgado A, Yin Y, Yu C, Vafeados D, Mora-Garcia S, Cheng JC, Nam KH, Li J, Chory J (2004) BRL1 and BRL3 are novel brassinosteroid receptors that function in vascular differentiation in Arabidopsis. Development 131:5341–5351
Cato S, McMillan L, Donaldson L, Richardson T, Echt C, Gardner R (2006) Wood formation from the base to the crown in Pinus radiata: gradients of tracheid wall thickness, wood density, radial growth rate and gene expression. Plant Mol Biol 60:565–581
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Chen CY, Hsieh MH, Yang CC, Lin CS, Wang AY (2010) Analysis of the cellulose synthase genes associated with primary cell wall synthesis in Bambusa oldhamii. Phytochemistry 71:1270–1279
Chen HM, Pang Y, Zeng J, Ding Q, Yin SY, Liu C, Lu MZ, Cui KM, He XQ (2012) The Ca2+-dependent DNases are involved in secondary xylem development in Eucommia ulmoides. J Integr Plant Biol 54:456–470
Cominelli E, Sala T, Calvi D, Gusmaroli G, Tonelli C (2008) Over-expression of the Arabidopsis AtMYB41 gene alters cell expansion and leaf surface permeability. Plant J: Cell Mol Biol 53:53–64
Day A, Neutelings G, Nolin F, Grec S, Habrant A, Cronier D, Maher B, Rolando C, David H, Chabbert B, Hawkins S (2009) Caffeoyl coenzyme A O-methyltransferase down-regulation is associated with modifications in lignin and cell-wall architecture in flax secondary xylem. Plant Physiol Biochem: PPB/Societe Francaise de Physiologie Vegetale 47:9–19
Dharmawardhana P, Brunner AM, Strauss SH (2010) Genome-wide transcriptome analysis of the transition from primary to secondary stem development in Populus trichocarpa. BMC Genomics 11:150
Di Matteo A, Giovane A, Raiola A, Camardella L, Bonivento D, De Lorenzo G, Cervone F, Bellincampi D, Tsernoglou D (2005) Structural basis for the interaction between pectin methylesterase and a specific inhibitor protein. Plant Cell 17:849–858
Du J, Groover A (2010) Transcriptional regulation of secondary growth and wood formation. J Integr Plant Biol 52:17–27
Fagerstedt KV, Kukkola EM, Koistinen VV, Takahashi J, Marjamaa K (2010) Cell wall lignin is polymerised by class III secretable plant peroxidases in Norway spruce. J Integr Plant Biol 52:186–194
Fornale S, Sonbol FM, Maes T, Capellades M, Puigdomenech P, Rigau J, Caparros-Ruiz D (2006) Down-regulation of the maize and Arabidopsis thaliana caffeic acid O-methyl-transferase genes by two new maize R2R3-MYB transcription factors. Plant Mol Biol 62:809–823
Fry SC, Smith RC, Renwick KF, Martin DJ, Hodge SK, Matthews KJ (1992) Xyloglucan endotransglycosylase, a new wall-loosening enzyme activity from plants. Biochem J 282(Pt 3):821–828
Galvao FC, Rossi D, Silveira Wda S, Valentini SR, Zanelli CF (2013) The deoxyhypusine synthase mutant dys1-1 reveals the association of eIF5A and Asc1 with cell wall integrity. PLoS One 8:e60140
Ghosh JS, Chaudhuri S, Dey N, Pal A (2013) Functional characterization of a serine-threonine protein kinase from Bambusa balcooa that implicates in cellulose overproduction and superior quality fiber formation. BMC Plant Biol 13:128
Grell MN, Linde T, Nygaard S, Nielsen KL, Boomsma JJ, Lange L (2013) The fungal symbiont of Acromyrmex leaf-cutting ants expresses the full spectrum of genes to degrade cellulose and other plant cell wall polysaccharides. BMC Genomics 14:928
Herrero J, Fernandez-Perez F, Yebra T, Novo-Uzal E, Pomar F, Pedreno MA, Cuello J, Guera A, Esteban-Carrasco A, Zapata JM (2013) Bioinformatic and functional characterization of the basic peroxidase 72 from Arabidopsis thaliana involved in lignin biosynthesis. Planta 237:1599–1612
Hossain MA, Noh HN, Kim KI, Koh EJ, Wi SG, Bae HJ, Lee H, Hong SW (2010) Mutation of the chitinase-like protein-encoding AtCTL2 gene enhances lignin accumulation in dark-grown Arabidopsis seedlings. J Plant Physiol 167:650–658
Howles PA, Birch RJ, Collings DA, Gebbie LK, Hurley UA, Hocart CH, Arioli T, Williamson RE (2006) A mutation in an Arabidopsis ribose 5-phosphate isomerase reduces cellulose synthesis and is rescued by exogenous uridine. Plant J: Cell Mol Biol 48:606–618
Huang W, Zhang SB, Hu H (2015) Insusceptibility of oxygen-evolving complex to high light in Betula platyphylla. Journal of Plant Research
Inouhe M, Nevins DJ (1991) Inhibition of auxin-induced cell elongation of maize coleoptiles by antibodies specific for cell wall glucanases. Plant Physiol 96:426–431
Kamerewerd J, Zadra I, Kurnsteiner H, Kuck U (2011) PcchiB1, encoding a class V chitinase, is affected by PcVelA and PcLaeA, and is responsible for cell wall integrity in Penicillium chrysogenum. Microbiology 157:3036–3048
Karpinska B, Karlsson M, Srivastava M, Stenberg A, Schrader J, Sterky F, Bhalerao R, Wingsle G (2004) MYB transcription factors are differentially expressed and regulated during secondary vascular tissue development in hybrid aspen. Plant Mol Biol 56:255–270
Ko JH, Beers EP, Han KH (2006) Global comparative transcriptome analysis identifies gene network regulating secondary xylem development in Arabidopsis thaliana. Mol Gen Genomics 276:517–531
Kovi MR, Zhang Y, Yu S, Yang G, Yan W, Xing Y (2011) Candidacy of a chitin-inducible gibberellin-responsive gene for a major locus affecting plant height in rice that is closely linked to green revolution gene sd1. TAG Theoretical and Applied Genetics Theoretische und Angewandte Genetik 123:705–714
Liu Z, Duguay J, Ma F, Wang TW, Tshin R, Hopkins MT, McNamara L, Thompson JE (2008) Modulation of eIF5A1 expression alters xylem abundance in Arabidopsis thaliana. J Exp Bot 59:939–950
Liu X, Wang Q, Chen P, Song F, Guan M, Jin L, Wang Y, Yang C (2012) Four novel cellulose synthase (CESA) genes from birch (Betula platyphylla Suk.) involved in primary and secondary cell wall biosynthesis. Int J Mol Sci 13:12195–12212
Liu L, Shang-Guan K, Zhang B, Liu X, Yan M, Zhang L, Shi Y, Zhang M, Qian Q, Li J, Zhou Y (2013) Brittle Culm1, a COBRA-like protein, functions in cellulose assembly through binding cellulose microfibrils. PLoS Genet 9:e1003704
Lu S, Li L, Yi X, Joshi CP, Chiang VL (2008) Differential expression of three eucalyptus secondary cell wall-related cellulose synthase genes in response to tension stress. J Exp Bot 59:681–695
Matos DA, Whitney IP, Harrington MJ, Hazen SP (2013) Cell walls and the developmental anatomy of the Brachypodium distachyon stem internode. PLoS One 8:e80640
Mauriat M, Moritz T (2009) Analyses of GA20ox- and GID1-over-expressing aspen suggest that gibberellins play two distinct roles in wood formation. Plant J: Cell Mol Biol 58:989–1003
Milhinhos A, Miguel CM (2013) Hormone interactions in xylem development: a matter of signals. Plant Cell Rep 32:867–883
Mishima K, Fujiwara T, Iki T, Kuroda K, Yamashita K, Tamura M, Fujisawa Y, Watanabe A (2014) Transcriptome sequencing and profiling of expressed genes in cambial zone and differentiating xylem of Japanese cedar (Cryptomeria japonica). BMC Genomics 15:219
Morrissy AS, Morin RD, Delaney A, Zeng T, McDonald H, Jones S, Zhao Y, Hirst M, Marra MA (2009) Next-generation tag sequencing for cancer gene expression profiling. Genome Res 19:1825–1835
Nishikubo N, Takahashi J, Roos AA, Derba-Maceluch M, Piens K, Brumer H, Teeri TT, Stalbrand H, Mellerowicz EJ (2011) Xyloglucan endo-transglycosylase-mediated xyloglucan rearrangements in developing wood of hybrid aspen. Plant Physiol 155:399–413
Paiva JA, Garces M, Alves A, Garnier-Gere P, Rodrigues JC, Lalanne C, Porcon S, Le Provost G, Perez Dda S, Brach J, Frigerio JM, Claverol S, Barre A, Fevereiro P, Plomion C (2008) Molecular and phenotypic profiling from the base to the crown in maritime pine wood-forming tissue. New Phytol 178:283–301
Persson S, Wei H, Milne J, Page GP, Somerville CR (2005) Identification of genes required for cellulose synthesis by regression analysis of public microarray data sets. Proc Natl Acad Sci U S A 102:8633–8638
Pesquet E, Korolev AV, Calder G, Lloyd CW (2010) The microtubule-associated protein AtMAP70-5 regulates secondary wall patterning in Arabidopsis wood cells. Curr Biol: CB 20:744–749
Pfaffl MW, Horgan GW, Dempfle L (2002) Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 30:e36
Polle A, Janz D, Teichmann T, Lipka V (2013) Poplar genetic engineering: promoting desirable wood characteristics and pest resistance. Appl Microbiol Biotechnol 97:5669–5679
Prassinos C, Ko JH, Yang J, Han KH (2005) Transcriptome profiling of vertical stem segments provides insights into the genetic regulation of secondary growth in hybrid aspen trees. Plant Cell Physiol 46:1213–1225
Ragni L, Nieminen K, Pacheco-Villalobos D, Sibout R, Schwechheimer C, Hardtke CS (2011) Mobile gibberellin directly stimulates Arabidopsis hypocotyl xylem expansion. Plant Cell 23:1322–1336
Rajangam AS, Kumar M, Aspeborg H, Guerriero G, Arvestad L, Pansri P, Brown CJ, Hober S, Blomqvist K, Divne C, Ezcurra I, Mellerowicz E, Sundberg B, Bulone V, Teeri TT (2008) MAP20, a microtubule-associated protein in the secondary cell walls of hybrid aspen, is a target of the cellulose synthesis inhibitor 2,6-dichlorobenzonitrile. Plant Physiol 148:1283–1294
Ren B, Chen Q, Hong S, Zhao W, Feng J, Feng H, Zuo J (2013) The Arabidopsis eukaryotic translation initiation factor eIF5A-2 regulates root protoxylem development by modulating cytokinin signaling. Plant Cell 25:3841–3857
Sato T, Takabe K, Fujita M (2004) Immunolocalization of phenylalanine ammonia-lyase and cinnamate-4-hydroxylase in differentiating xylem of poplar. C R Biologies 327:827–836
Shani Z, Dekel M, Roiz L, Horowitz M, Kolosovski N, Lapidot S, Alkan S, Koltai H, Tsabary G, Goren R, Shoseyov O (2006) Expression of endo-1,4-beta-glucanase (cel1) in Arabidopsis thaliana is associated with plant growth, xylem development and cell wall thickening. Plant Cell Rep 25:1067–1074
Spicer R, Tisdale-Orr T, Talavera C (2013) Auxin-responsive DR5 promoter coupled with transport assays suggest separate but linked routes of auxin transport during woody stem development in Populus. PLoS One 8:e72499
Steinwand BJ, Kieber JJ (2010) The role of receptor-like kinases in regulating cell wall function. Plant Physiol 153:479–484
Tanaka K, Murata K, Yamazaki M, Onosato K, Miyao A, Hirochika H (2003) Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol 133:73–83
Thumma BR, Matheson BA, Zhang D, Meeske C, Meder R, Downes GM, Southerton SG (2009) Identification of a Cis-acting regulatory polymorphism in a eucalypt COBRA-like gene affecting cellulose content. Genetics 183:1153–1164
Tokunaga N, Kaneta T, Sato S, Sato Y (2009) Analysis of expression profiles of three peroxidase genes associated with lignification in Arabidopsis thaliana. Physiol Plant 136:237–249
van Raemdonck D, Pesquet E, Cloquet S, Beeckman H, Boerjan W, Goffner D, El Jaziri M, Baucher M (2005) Molecular changes associated with the setting up of secondary growth in aspen. J Exp Bot 56:2211–2227
Verma DP, Maclachlan GA, Byrne H, Ewings D (1975) Regulation and in vitro translation of messenger ribonucleic acid for cellulase from auxin-treated pea epicotyls. J Biol Chem 250:1019–1026
Wagner A, Donaldson L, Kim H, Phillips L, Flint H, Steward D, Torr K, Koch G, Schmitt U, Ralph J (2009) Suppression of 4-coumarate-CoA ligase in the coniferous gymnosperm Pinus radiata. Plant Physiol 149:370–383
Wakabayashi K, Soga K, Hoson T (2012) Phenylalanine ammonia-lyase and cell wall peroxidase are cooperatively involved in the extensive formation of ferulate network in cell walls of developing rice shoots. J Plant Physiol 169:262–267
Wang J, Kucukoglu M, Zhang L, Chen P, Decker D, Nilsson O, Jones B, Sandberg G, Zheng B (2013) The Arabidopsis LRR-RLK, PXC1, is a regulator of secondary wall formation correlated with the TDIF-PXY/TDR-WOX4 signaling pathway. BMC Plant Biol 13:94
Wang C, Zhang N, Gao C, Cui Z, Sun D, Yang C, Wang Y (2014) Comprehensive transcriptome analysis of developing xylem responding to artificial bending and gravitational stimuli in Betula platyphylla. PLoS One 9:e87566
Wilkins O, Nahal H, Foong J, Provart NJ, Campbell MM (2009) Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiol 149:981–993
Ye ZH, York WS, Darvill AG (2006) Important new players in secondary wall synthesis. Trends Plant Sci 11:162–164
Zabotina O, Malm E, Drakakaki G, Bulone V, Raikhel N (2008) Identification and preliminary characterization of a new chemical affecting glucosyltransferase activities involved in plant cell wall biosynthesis. Mol Plant 1:977–989
Zhang M, Henquet M, Chen Z, Zhang H, Zhang Y, Ren X, van der Krol S, Gonneau M, Bosch D, Gong Z (2009) LEW3, encoding a putative alpha-1,2-mannosyltransferase (ALG11) in N-linked glycoprotein, plays vital roles in cell-wall biosynthesis and the abiotic stress response in Arabidopsis thaliana. Plant J: Cell Mol Biol 60:983–999
Zhang X, Gou M, Liu CJ (2013) Arabidopsis Kelch repeat F-box proteins regulate phenylpropanoid biosynthesis via controlling the turnover of phenylalanine ammonia-lyase. Plant Cell 25:4994–5010
Zhong R, Lee C, Zhou J, McCarthy RL, Ye ZH (2008) A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 20:2763–2782
Zhou J, Lee C, Zhong R, Ye ZH (2009) MYB58 and MYB63 are transcriptional activators of the lignin biosynthetic pathway during secondary cell wall formation in Arabidopsis. Plant Cell 21:248–266
Acknowledgments
This work was supported by the National High Technology Research and Development Program (“863” program) of China (2013AA102704) and the National Natural Science Foundation of China (31470671).
Author contributions
HG, YW, CY, and CW conceived and designed the experiments. HG, PH, and YJ participated in the experiments. HG, YW, and CW performed the data analysis. HG, YW, CY, and CW drafted the manuscript. HG and CW provided reagents and analysis tools. All authors read and approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest, and the final manuscript is approved by all authors for publication.
Data archiving statement
The data is currently being submitted but should still be included with a note that the accession numbers will be supplied once available.
Additional information
Communicated by P. Ingvarsson
Electronic supplementary material
Table S1
Primer sequences and amplicon sizes used for real-time RT-PCR assays. (DOC 62 kb)
Table S2
Differentially expressed genes identified by digital gene expression (DGE) analysis (FDR ≤0.001, |log2 ratio| ≥1). (XLSX 139 kb)
Fig. S1
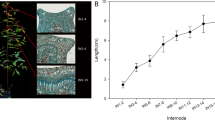
Sections analyzed by scanning electron microscopy (×500) in the first, third, and fifth stem internodes. co cortex, xy xylem, sx secondary xylem, pi pith. Bars = 25 μm. (GIF 118 kb)
Fig. S2
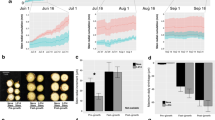
Tag expression distribution. The column indicates the distribution of total tags of the studied stem internodes. (GIF 45 kb)
Fig. S3
Saturation evaluation of the different gene expression levels. The solid line indicates genes that were mapped by all clean tags, and the dotted line indicates genes (GIF 321 kb)
Fig. S4
Statistics of the clean tags alignment. The column indicates mapping of the tags from the studied stem internodes. PM (Sense) perfect match to gene (Sense), 1 tag- >1 gene match to one gene; 1 tag- >n gene match to more than one gene, 1 MM (Sense) match to gene (Sense) with 1-bp mismatch, PM (Antisense) perfect match to an antisense gene sequence, 1 MM (Antisense) match to antisense gene with 1-bp mismatch, PM Genome 1 tag- >1 position perfect match to genome sequence with one best hit, PM Genome 1 tag- >n position perfect match to genome sequence with multiple best hits, 1 MM Genome match to genome sequence with 1 bp mismatch, Unknown Tag not matched to a gene (Sense and Antisense) or genome sequence. (GIF 58 kb)
Fig. S5
Gene ontology (GO) categories assigned to differentially expressed genes in the first, third, and fifth stem internodes. The x-axis represents the number of unigenes, and the y-axis indicates the GO subcategories. (GIF 51 kb)
Fig. S6
Hierarchical cluster analysis of differentially expressed genes in the studied stem internodes. All ratios are log2 transformed. Log ratios of 0 (ratios of 1) are indicated in black, and increasingly positive (induction) or negative (repression) log ratios are indicated in red or green, respectively, with increasing intensities. Red indicates induction, and green indicates repression in the arrays. (PNG 43 kb)
Rights and permissions
About this article
Cite this article
Guo, H., Wang, Y., Hu, P. et al. Gene expression profiles in different stem internodes reveal the genetic regulation of primary and secondary stem development in Betula platyphylla . Tree Genetics & Genomes 12, 113 (2016). https://doi.org/10.1007/s11295-016-1068-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-016-1068-x