Abstract
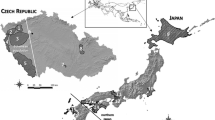
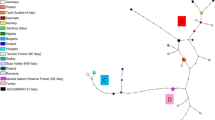
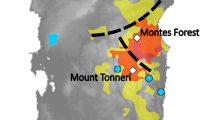
In southern Kantoh, Japanese sika deer (Cervus nippon) are distributed discontinuously due to large urban areas and developed road networks. To assess the impact of habitat fragmentation on sika deer subpopulations, we examined mitochondrial D-loop sequences from 435 individuals throughout southern Kantoh. About 13 haplotypes were detected, and their distributions revealed spatial genetic structure. Significant genetic differentiation was observed among seven of eight subpopulations. We found no significant correlation between pairwise F ST and geographical distance among subpopulations. Genetic diversity indices suggested that seven of eight subpopulations had probably experienced population bottlenecks in the recent past. Therefore, and in the light of the results of a nested clade analysis of these haplotypes, we conclude that recent fluctuations in population size and the interruption of gene flow due to past and present habitat fragmentation have played major roles influencing the spatial genetic structure of the sika deer population. This is the first evidence of spatial genetic population structure in the highly fragmented sika deer population in Honshu, Japan.






Similar content being viewed by others
References
Alpers DL, Van Vuuren BJ, Arctander P, Robinson TJ (2004) Population genetics of the roan antelope (Hippotragus equinus) with suggestions for conservation. Mol Ecol 13:1771–1784
Avise JC (2004) Molecular markers, natural history and evolution, 2nd edn. Sinauer Associates, Sunderland, Massachusetts
Bideau E, Gerard JF, Vincent JP, Maublanc ML (1993) Effects of age and sex on space occupation by European roe deer. J Mammal 74:745–751
Biodiversity Center of Japan (2004) The national survey on the natural environment—report of the distributional survey of Japanese animal (Mammals) (in Japanese). pp 34–39
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659
Clutton-Brock TH, Guiness FE, Albon SD (1982) Red deer: behavior and ecology of two sexes. The University of Chicago Press, Chicago
Coulon A, Cosson JF, Angibault JM, Cargnelutti B, Galan M, Morellet N, Petit E, Aulagnier S, Hewison JM (2004) Landscape connectivity influences gene flow in a roe deer population inhabiting a fragmented landscape: an individual-based approach. Mol Ecol 13:2841–2850
Cronin MA, Bleich VC (1995) Mitochondrial DNA variation among populations and subspecies of mule deer in California. Calif Fish Game 81:45–54
Cronin MA, Nelson ME, Pac DF (1991) Spatial heterogeneity of mitochondrial DNA and allozymes among populations of white-tailed deer and mule deer. J Hered 82:118–127
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Furubayashi K (1996) The study on protection of sika deer in the Tanzawa Mountains (in Japanese with English summary). Ph.D. thesis, University of Kyoto, Kyoto
Furubayashi K, Shinoda Y (2001) Distribution of sika deer (Cervus nippon) around Edo in Edo era (in Japanese with English summary). Wildl Conserv Jpn 7:1–24
Gerlach G, Musolf K (2000) Fragmentation of landscape as a cause for genetic subdivision in bank voles. Conserv Biol 14:1066–1074
Grant WS, Bowen BW (1998) Shallow population histories in deep evolutionary lineage of marine fishes: insight from sardines and anchovies and lessons for conservation. J Hered 89:415–426
Hirota T, Hirohata T, Mashima H, Satoh T, Obara Y (2004) Population structure of the large Japanese field mouse, Apodemus speciosus (rodentia: muridae), in suburban landscape, based on mitochondrial D-loop sequences. Mol Ecol 13:3275–3282
Hutchison DW, Templeton AR (1999) Correlation of pair wise genetic and geographic distance measure: inferring the relative influences of gene flow and drift on the distribution of genetic variability. Evolution 53:1898–1914
Ishibashi Y, Saitoh T (2004) Phylogenetic relationships among fragmented Asian black bear (Ursus thibetanus) populations in western Japan. Conserv Genet 5:311–323
Kanagawa Prefectural Government (1984) Kanagawa no rinsei-shi (history of silvicultural policy in Kanagawa Prefecture) (in Japanese). Kanagawa Prefecture Department of Agriculture Forestry Division, Yokohama
Kanagawa Prefectural Government (2003) Management plan of the Japanese sika deer in Kanagawa Prefecture (in Japanese). Kanagawa Prefecture Department of Environment and Agriculture, Yokohama
Kanagawa Prefectural Government (2004) Choujyu hogo-ku tou ichi-zu (wildlife protection area map) (in Japanese). Syobunsya Publications Inc., Tokyo
Keller I, Nentwig W, Largiadèr CR (2004) Recent habitat fragmentation due to roads can lead to significant genetic differentiation in an abundant flightless ground beetle. Mol Ecol 13:2983–2994
Kimura M, Weiss GH (1964) The stepping stone model of population structure and the decrease of genetic correlation with distance. Genetics 49:561–576
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Koganezawa M (1989) The 19th century distribution of sika deer (Cervus nippon) and wild boar (Sus scrofa) in Tochigi Prefecture, estimated by the local documents – preliminary report (in Japanese). Bull Tochigi Prefectural Museum 6:65–80
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Kyle CJ, Strobeck C (2001) Genetic structure of North American wolverine (Gulo gulo) populations. Mol Ecol 10:337–347
Matsushita M (1992) Lucidophyllous forest development along the Pacific coast of the Japanese archipelago during the Holocene (in Japanese with English summary). Quaternary Res 31:375–387
McCullough DR (ed) (1996) Metapopulations and wildlife conservation. Island Press, Washington, DC
Moritz C (1994) Applications of mitochondrial DNA analysis in conservation: a critical review. Mol Ecol 3:401–411
Nagata J, Masuda R, Kaji K, Kaneko M, Yoshida MC (1998) Genetic variation and population structure of the Japanese sika deer (Cervus nippon) in Hokkaido Island, based on mitochondrial D-loop sequences. Mol Ecol 7:871–877
Nagata J, Masuda R, Tamate HB, Hamasaki S, Ochiai K, Asada M, Tatsuzawa S, Suda K, Tado H, Yoshida MC (1999) Two genetically distinct lineages of the sika deer, Cervus nippon, in Japanese Islands: comparison of mitochondrial D-loop region sequences. Mol Phylogenet Evol 13:511–519
Nelson ME, Mech LD (1987) Demes within a northeastern Minnesota deer population. In: Chepko-Sade BD, Halpin Z (eds) Mammalian dispersal patterns. University of Chicago Press, Chicago, pp 27–40
Onorato DP, Hellgren EC, Van Den Bussche RA, Doan-Crider DL (2004) Phylogeographic patterns within a metapopulation of black bears (Ursus americanus) in the American southwest. J Mammal 85:140–147
Ota Y, Omura A (1991) Late quaternary shoreline in the Japanese Islands. Quaternary Res 30:175–186
Patton JL, Da Silva MNF, Malcolm JR (1996) Hierarchical genetic structure and gene flow in three sympatric species of Amazonian rodents. Mol Ecol 5:229–238
Pitra C, Hansen AJ, Lieckfeldt D, Arctander P (2002) An exceptional case of historical out breeding in African sable antelope populations. Mol Ecol 11:1197–1208
Posada D, Crandall KA, Templeton AR (2000) GeoDis: a program for the cladistic nested analysis of the geographical distribution of genetic haplotypes. Mol Ecol 9:487–488
Purdue JR, Smith MH, Patton JC (2000) Female philopatry and extreme spatial genetic heterogeneity in white-tailed deer. J Mammal 81:179–185
Raymond M, Rousset F (1995) GENEPOP Version 1.2: population genetic software for exact tests and ecumenicism. J Hered 86:248–249
Saccheri I, Kuussaari M, Kankare M, Vikman P, Fortelius W, Hanski I (1998) Inbreeding and extinction in a butterfly metapopulation. Nature 392:491–494
Saitama Prefectural Government (2000) Choujyu hogo-ku tou ichi-zu (a map of game hunting in Saitama Prefecture) (in Japanese). Naigai-chizu Inc., Tokyo
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Satoh T, Obara Y (1995) DNA fingerprints in the honey bess, Apis mellifera, using ant-derived DNA probe. Appl Entomol Zool 31:148–151
Schaschl H, Kaulfus D, Hammer S, Suchentrunk F (2003) Spatial patterns of mitochondrial and nuclear gene pools in chamois (Rupicapra r. rupicapra) from the Eastern Alps. Heredity 91:125–135
Schneider S, Roessli D, Excoffier L (2000) ARLEQUIN, Version 2: software for population genetics data analysis. Genetics and Biometry Laboratory, University of Geneva, Switzerland
Shizuoka Prefectural Government (2000) Choujyu hogo-ku tou ichi-zu (wildlife protection area map) (in Japanese). Chuo-chizu Inc., Tokyo
Taguchi H (1997) Historical relationship between cultivation and exclusion of wildlife in vicinity of Tanzawa mountains. In: Environment Department of Kanagawa Prefecture (eds) Report on the natural environmental study in Tanzawa-Ooyama (in Japanese). Kanagawa Prefecture, Yokohama, pp 422–452
Tamate HB, Tatsuzawa S, Suda K, Izawa M, Doi T, Sunagawa K, Miyahira F, Tado H (1998) Mitochondrial DNA variations in local populations of the Japanese sika deer, Cervus nippon. J Mammal 79:1396–1403
Templeton AR (2004) Statistical phylogeography: methods of evaluating and minimizing inference errors. Mol Ecol 13:789–809
Templeton AR, Routman E, Phillips C (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132:619–633
Templeton AR, Sing CF (1993) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping. IV. Nested analyses with cladogram uncertainty and recombination. Genetics 134:659–669
Templeton AR, Routman E, Phillips CA (1995) Separating population structure from population history: a cladistic analysis of the geographical distribution of mitochondrial DNA haplotypes in the tiger salamander, Ambystoma tigrinum. Genetics 140:767–782
Templeton AR (1998) Nested clade analyses of phylogeographic data: testing hypotheses about gene flow and population history. Mol Ecol 7:381–397
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL-Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res (Computer File) 25:4876–4882
Tierson WC, Mattfeld GF, Sage RW, Behrend DF (1985) Seasonal movements and home ranges of white-tailed deer in the Adirondacks. J Wildl Manage 49:760–769
Tokyo Metropolitan Government (2002) Choujyu hogo-ku tou ichi-zu (wildlife protection area map) (in Japanese). Naigai-chizu Inc., Tokyo
Tomasik E, Cook JA (2005) Mitochondrial phylogeography and conservation genetics of wolverine (Gulo gulo) of northwestern North America. J Mammal 86:386–396
Tsukada M (1982) Cryptomeria japonica: glacial refugia and late-glacial and postglacial migration. Ecology 63:1091–1105
Umitsu M (1991) Holocene sea-level changes and coastal evolution in Japan. Quaternary Res 30:187–196
Wang M, Schreiber A (2001) The impact of habitat fragmentation and social structure on the population genetics of roe deer (Capreolus capreolus L.) in central Europe. Heredity 86:703–715
Werle E, Schneider C, Renner M, Volker M, Fiehen W (1994) Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res (Computer File) 22:4354–4355
Yamanashi Prefectural Government (2004) Choujyu hogo-ku tou ichi-zu (wildlife protection area map) (in Japanese). Midorikawa-chizu-Insatu Inc., Tokyo
Yasuda Y (1985) Climatic fluctuations in the last glacial period: influences on the evolution of Japanese paleolithic culture. J Geogr 94:586–594
Zenger KR, Eldridge MDB, Cooper DW (2003) Intraspecific variation, sex-biased dispersal and phylogeography of the eastern gray kangaroo (Macropus giganteus). Heredity 91:153–162
Acknowledgements
We thank Kanagawa Natural Environment Conservation Center and Yamanashi Forest and Forestry Products Research Institute, Mr. Shinji Oda, Mr. Takahiro Ohba, Mr. Yukitoshi Totake, Dr. Shin-ichi Hayama and the hunting association members in Kanagawa, Yamanashi, Shizuoka, Saitama Prefectures, and Tokyo Metropolitan for collecting sika deer samples. We express our thanks to all members of the Wildlife Ecology Laboratory of the Forestry and Forest Products Research Institute for the use of facilities and helpful support. We are indebted to Mr. Tadamasa Okumura, Mr. Misao Okano, and other members of the Wildlife Management Office Inc., for assistance in making figures with GIS and other helpful support. This study was supported in part by a grant from the Japan Forest Technology Association and the Nature Conservation Society of Tanzawa.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Yuasa, T., Nagata, J., Hamasaki, S. et al. The impact of habitat fragmentation on genetic structure of the Japanese sika deer (Cervus nippon) in southern Kantoh, revealed by mitochondrial D-loop sequences. Ecol Res 22, 97–106 (2007). https://doi.org/10.1007/s11284-006-0190-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11284-006-0190-x