Abstract
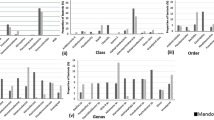
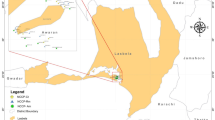
Mangroves are unique but endangered coastal ecosystems that play a vital role in the tropical and subtropical environments. Mauritius has two species of mangroves, Bruguiera gymnorrhiza (L.) Lam. and Rhizophora mucronata Lam., growing along its coast. The mangrove rhizosphere harbours a diverse microbial community and the use of RNA-sequencing can reveal both the taxonomic composition and active biochemical functions of the complex microbial community. Metatranscriptomic study was carried out by comparing the microbial community of rhizosphere microbiomes sediments from the two mangroves species. The study also included a comparison between a natural and a man grown mangrove microbiome. Overall, samples showed predominance by Proteobacteria, Bacteroidetes and Firmicutes, with high abundance of sulphate reducers, nitrogen reducers and methanogens. Significant difference was, however, noted at both taxonomic and functional levels among the mangroves species. The data also indicate that the microbial core involved in methane, nitrogen, and sulphur metabolism consisted mainly of Burkholderiaceae, Planctomycetaceae, Rhodobacteraceae, and Desulfobacteraceae. Also, genes encoding enzymes involved in carbon cycling, the metabolism of nitrogen, methane and sulphur were dominant in the rhizosphere of the natural mangrove ecosystem. To our knowledge, this is a first metatranscriptomic study on the microbiome of mangroves in the Mauritius, and our results provide the first insights in the range of functions and microbial diversity of the local mangrove species.







Similar content being viewed by others
References
Acosta-Gonzalez A, Marques S (2016) Bacterial diversity in oil-polluted marine coastal sediments. Curr Opin Biotechnol 38:24–32. https://doi.org/10.1016/j.copbio.2015.12.010
Altschul SF, Gish W, Miller W, Myers EW, Lipman D (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Alzubaidy H, Essack M, Malas TB, Bokhari A, Motwalli O, Kamanu FK, Jamhor SA, Mokhtar NA, Antunes A, Simões MF, Alam I, Bougouffa S, Lafi FF, Bajic VB, Archer JA (2016) Rhizosphere microbiome metagenomics of gray mangroves (Avicennia marina) in the Red Sea. Gene 576:626–636. https://doi.org/10.1016/j.gene.2015.10.032
Andreote FD, Jiménez DJ, Chaves D, Dias ACF, Luvizotto DM, Dini-Andreote F, Fasanella CC, Lopez MV, Baena S, Taketani RG, Soares de Melo I (2012) The microbiome of Brazilian mangrove sediments as revealed by metagenomics. PLoS ONE 7:e38600. https://doi.org/10.1371/journal.pone.0038600
Appadoo C (2003) Status of mangroves in Mauritius. J Coastal Dev 7:1–4
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:1
Beck DA, Kalyuzhnaya MG, Malfatti S, Tringe SG, Glavina Del Rio T, Ivanova N, Lidstrom ME, Chistoserdova L (2013) A metagenomic insight into freshwater methane-utilizing communities and evidence for cooperation between the Methylococcaceae and the Methylophilaceae. PeerJ 1:e23 https://doi.org/10.7717/peerj.23
Braker G, Tiedje JM (2003) Nitric oxide reductase (norB) genes from pure cultures and environmental samples. Appl Environ Microbiol 69:3476–3483. https://doi.org/10.1128/aem.69.6.3476-3483.2003
Cabral L, Júnior GV, Pereira de Sousa ST, Dias AC, Lira Cadete L, Andreote FD, Hess M, de Oliveira VM (2016) Anthropogenic impact on mangrove sediments triggers differential responses in the heavy metals and antibiotic resistomes of microbial communities. Environ Pollut 216:460–469. https://doi.org/10.1016/j.envpol.2016.05.078
Dias AC, Pereira e Silva MC, Cotta SR, Dini-Andreote F, Soares FL Jr, Salles JF, Azevedo JL, van Elsas JD, Andreote FD (2012) Abundance and genetic diversity of nifH gene sequences in anthropogenically affected Brazilian mangrove sediments. Appl Environ Microbiol 78:7960–7967. https://doi.org/10.1128/AEM.02273-12
dos Santos HF, Cury JC, do Carmo FL, dos Santos AL, Tiedje J, van Elsas JD, Rosado AS, Peixoto RS (2011) Mangrove bacterial diversity and the impact of oil contamination revealed by pyrosequencing: bacterial proxies for oil pollution. PLoS ONE 6:e16943. https://doi.org/10.1371/journal.pone.0016943
Feller IC, Whigham DF, McKee KL, Lovelock CE (2003) Nitrogen limitation of growth and nutrient dynamics in a disturbed mangrove forest, Indian River Lagoon, Florida. Oecologia 134:405–414. https://doi.org/10.1007/s00442-002-1117-z
Glass EM, Wilkening J, Wilke A, Antonopoulos D, Meyer F (2010) Using the metagenomics RAST server (MG-RAST) for analyzing shotgun metagenomes. https://doi.org/10.1101/pdb.prot5368
Gomes NC, Flocco CG, Costa R, Junca H, Vilchez R, Pieper DH, Krögerrecklenfort E, Paranhos R, Mendonça-Hagler LC, Smalla K (2010) Mangrove microniches determine the structural and functional diversity of enriched petroleum hydrocarbon-degrading consortia. FEMS Microbiol Ecol 74:276–290. https://doi.org/10.1111/j.1574-6941.2010.00962.x
Goncalves AC, dos Santos AC, dos Santos TF, Pessoa TB, Dias JC, Rezende RP (2015) High yield of functional metagenomic library from mangroves constructed in fosmid vector. Genet Mol Res 14:11841–11847. https://doi.org/10.4238/2015.October.2.17
Jean MR, Gonzalez-Rizzo S, Gauffre-Autelin P, Lengger SK, Schouten S, Gros O (2015) Two new Beggiatoa species inhabiting marine mangrove sediments in the Caribbean. PLoS ONE 10:e0117832. https://doi.org/10.1371/journal.pone.0117832
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280
Kopylova E, Noe L, Touzet H (2012) SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28:3211–3217. https://doi.org/10.1093/bioinformatics/bts611
Liang JB, Chen YQ, Lan CY, Tam NFY, Zan QJ, Huang LN (2006) Recovery of novel bacterial diversity from mangrove sediment. Mar Biol 150:739–747. https://doi.org/10.1007/s00227-006-0377-2
Nogueira VLR, Rocha LL, Colares GB, Angelima AL, Normandoa LRO, Cantão ME, Agnez-Lima LF, Andreote FD, Meloa VMM (2015) Microbiomes and potential metabolic pathways of pristine and anthropized Brazilian mangroves. Reg Stud Marine Sci 2:56–64. https://doi.org/10.1016/j.rsma.2015.08.008
Overbeek R. Begley T, Butler RM, Choudhuri JV, Chuang HY, Cohoon M, de Crécy-Lagard V, Diaz N, Disz T, Edwards R, Fonstein M, Frank ED, Gerdes S, Glass EM, Goesmann A, Hanson A, Iwata-Reuyl D, Jensen R, Jamshidi N, Krause L, Kubal M, Larsen N, Linke B, McHardy AC, Meyer F, Neuweger H, Olsen G, Olson R, Osterman A, Portnoy V, Pusch GD, Rodionov DA, Rückert C, Steiner J, Stevens R, Thiele I, Vassieva O, Ye Y, Zagnitko O, Vonstein V (2005) The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33:5691–5702. https://doi.org/10.1093/nar/gki866
Tatusova T, Ciufo S, Fedorov B, O’Neill K, Tolstoy I (2014) RefSeq microbial genomes database: new representation and annotation strategy. Nucleic Acids Res 42:D553–D559. https://doi.org/10.1093/nar/gkt1274
Tripathy S, Padhi SK, Mohanty S, Samanta M, Maiti NK (2016) Analysis of the metatranscriptome of microbial communities of an alkaline hot sulfur spring revealed different gene encoding pathway enzymes associated with energy metabolism. Extremophiles 20:525–536. https://doi.org/10.1007/s00792-016-0846-6
Acknowledgements
The authors wish to thank Chowdhoory Pravin for the collection of samples. This work was supported by the University of Mauritius (Grant: 0085) and LANDRACE (Surya Seeds Research).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Sequences data
Sequence submitted to NCBI (SRA accession: SRP133367) and is accessible at: https://www.ncbi.nlm.nih.gov/sra/SRP133367.
Rights and permissions
About this article
Cite this article
Rampadarath, S., Bandhoa, K., Puchooa, D. et al. Metatranscriptomics analysis of mangroves habitats around Mauritius. World J Microbiol Biotechnol 34, 59 (2018). https://doi.org/10.1007/s11274-018-2442-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-018-2442-7